Abstract
Epidemiological modelling has assisted in identifying interventions that reduce the impact of COVID-19. The UK government relied, in part, on the CovidSim model to guide its policy to contain the rapid spread of the COVID-19 pandemic during March and April 2020; however, CovidSim contains several sources of uncertainty that affect the quality of its predictions: parametric uncertainty, model structure uncertainty and scenario uncertainty. Here we report on parametric sensitivity analysis and uncertainty quantification of the code. From the 940 parameters used as input into CovidSim, we find a subset of 19 to which the code output is most sensitive—imperfect knowledge of these inputs is magnified in the outputs by up to 300%. The model displays substantial bias with respect to observed data, failing to describe validation data well. Quantifying parametric input uncertainty is therefore not sufficient: the effect of model structure and scenario uncertainty must also be properly understood.
Similar content being viewed by others
Main
CovidSim is an individual-based simulation code developed by the MRC Centre for Global Infectious Disease Analysis at Imperial College London. It is a modified version of an earlier model designed to support pandemic influenza planning1 and has now been used to explore various non-pharmaceutical interventions (NPIs) with the aim of reducing the transmission of the coronavirus, as documented in the key paper2 denoted Report 9. CovidSim played an important role in the United Kingdom in reorienting UK Government policy from herd immunity to a strategy focused on suppression of the viral infection. It should be noted, however, that many competitor models exist. Notable examples include the work performed at the London School of Hygiene and Tropical Medicine (see, for instance, refs. 3,4 and especially ref. 5), in which the effects of different NPIs in the UK are modelled. Another noteworthy model is CovaSim6, which is similar in structure to CovidSim in the sense that it models a population of individuals via discrete agents.
Likewise, CovidSim creates a network of individuals located in areas defined by high-resolution population density data. In the model, contacts with other individuals can be made in four different types of place, namely, within households, at schools, universities and work places. It is possible to model a combination of different NPIs, namely, general social distancing (SD), social distancing for those over 70 years of age (SDOL70), home isolation of suspected cases (CI), voluntary home quarantine (HQ) and place closure of universities and schools (PC) (see Table 2 of ref. 2). CovidSim contains over 900 input parameters, which are mainly located in two input files. Furthermore, a small number of parameters that define certain characteristics of the intervention scenario one wishes to study are supplied via the command line.
We investigated the reproducibility of the code, as has been done in past work7,8. That said, we especially focus on CovidSim’s robustness to uncertainty in the input parameters. By robustness in this context, we mean the extent to which the code amplifies uncertainties from the input to the output. Our main aim is to thus take the model as given and examine the uncertainty in its predictions when its parameters are treated as random variables instead of deterministic inputs. We will use a dimension-adaptive sampling method for this purpose9 to be able to handle the high-dimensional input space. This type of anisotropic sampling method adaptively exploits a possible low effective dimension, where only a subset of all inputs have a substantial impact on the model output. A wide range of domains have seen the application of such dimension-adaptive samplers, for example, computational electromagnetism10, finance11,12 and natural convection problems13, to name just a few. Here we perform a validation study to examine the ability of the predicted output distribution to envelop the observed COVID-19 death count, conditional on a predefined intervention scenario.
Due to the large number of inputs, one cannot hope to obtain an accurate, data-informed value of all parameters in contention. Moreover, considering CovidSim’s influential status and its likely use in future COVID-19 predictions, it is important to assess the impact of parametric uncertainty on the model output. We will argue the case for the prediction of uncertainty in high-impact decision-making, after we first describe our results.
Results
We have performed an analysis on the original closed-source version of the code; however, the majority of our sensitivity analysis and uncertainty quantification efforts lie with the current updated open-source release of CovidSim.
With respect to the original version, we have been able to achieve exact reproducibility of the results2 in Report 9, although only when running within an Azure cloud environment. Attempts to run the code on a Linux-based machine failed; we could not reproduce the same results here and as this version is no longer supported it was not investigated further.
Uncertainty in CovidSim
The predictions of most computational models are affected by uncertainty from a variety of different sources. We identify the following three sources of uncertainty in CovidSim; namely, parametric uncertainty, model structure uncertainty and scenario uncertainty. This breakdown is not uncommon (see, for example, refs. 14,15,16).
Parametric uncertainty arises due to imperfect knowledge of the model input parameters \({\boldsymbol{\xi }}\in {{\mathbb{R}}}^{d}\), described in the ‘CovidSim parameters’ section. Model structure uncertainty is more fundamental, as it relates to uncertainty about the appropriate mathematical structure of the model, denoted by \({\mathcal{M}}\); one can think of missing epidemiological processes that are not implemented in CovidSim (see the discussion in Supplementary Section 6). Finally, a scenario \({\mathcal{S}}\) is the set of conditions under which a model \({\mathcal{M}}\left({\boldsymbol{\xi }}\right)\) is applied. In the case of CovidSim, \({\mathcal{S}}\) includes the choice of NPI scenarios, the initialization of the model and the well-known reproduction number R0. Note that the actual implementation of \({\mathcal{S}}\) will be parameterized as well, and that we could technically lump these parameters in with ξ; however, the scenario parameters are of a different nature than the internal inputs ξ, and treating \({\mathcal{S}}\) as a separate category mirrors the way in which the results were presented in Report 9, which showed results for different NPIs and R0 values.
If we denote q as the predicted output quantity of interest, we therefore have \(q=q\left({\boldsymbol{\xi }},{\mathcal{M}},{\mathcal{S}}\right)\), where all three arguments are uncertain. As noted, our main goal is to quantify the impact of parametric uncertainty. By treating the inputs as random variables with probability density function (PDF) \(p\left({\boldsymbol{\xi }}\right)\), our mean prediction is given by
where Ωξ is the support of p(ξ). The uncertainty in the prediction in equation (1) can be represented by either the corresponding variance or confidence intervals. It is important to note that our results are conditional on \({\mathcal{M}}\) and \({\mathcal{S}}\). We are not in a position to change the former and we illustrate the importance of scenario uncertainty by repeating the parametric uncertainty analysis for two different scenarios.
Uncertainty propagation
We use EasyVVUQ17,18 from the Verified Exascale Computing for Multiscale Applications (VECMA) toolkit19 to propagate the input uncertainties through CovidSim. Templates from the CovidSim input files are generated to interface CovidSim with EasyVVUQ. In the process, a single file is generated, which contains all inputs, with their types and default values specified. Simply counting the number of entries in this file allows us to exactly determine the number of parameters present in the code, which is how we arrived at a number of 940 inputs.
We will not vary all 940 parameters (see the ‘CovidSim parameters’ section). We will instead assign to d (d ≪ 940) input parameters ξi an independent PDF, that is, ξi ≈ p(ξi); a d-dimensional sampling plan from the joint PDF, ∏ip(ξi), is then created, after which CovidSim is evaluated at each input point. We refine the sampling plan in a dimension-adaptive manner to handle the high-dimensional input space, the details of which can be found in the ‘Statistics’ section in the Methods.
CovidSim parameters
In this section we describe how we arrived at our selection of input parameters that we vary as part of our uncertainty quantification study. We have divided the parameters present in the input files into three groups:
-
(1)
Group 1, intervention parameters—these are parameters meant to slow down the viral infection, which can still be varied for a fixed \({\mathcal{S}}\) (for example, the length of time households are quarantined) when HQ is part of the selected NPI set.
-
(2)
Group 2, disease parameters—related to the characteristics of COVID-19 (for example, the latent period).
-
(3)
Group 3, spatial/geographic parameters—parameters that apply to the properties of the network (for example, the relative transmission rate for place types).
The purpose of this classification was to direct initial, exploratory uncertainty quantification (UQ) and sensitivity analysis (SA) campaigns on a coherent subset of parameters. The final UQ campaign contains parameters from all three groups. By campaign, we mean a single forward propagation step of uncertainty from the input to the output.
Before starting the UQ analysis we first performed a parameter study using, in part, expert domain knowledge from the CovidSim team at Imperial College London to reduce the number of inputs. We focus on a scenario based on the suppression release in the Report 9 folder on GitHub20, using the intervention setting that combines PC, CI, HQ and SD, as this class is the closest to actual NPIs that were implemented in the UK. In this case we have the aforementioned total of 940 parameters. Note that some input parameters are vectors, in which case we counted each entry as a separate input parameter. On top of our own initial selection, we received feedback over the course of our analysis from the developers of CovidSim as to the inclusion of given parameters in the UQ study.
Many of the parameters are (currently) not used in the case of COVID-19 simulation, such as numerous vaccination parameters. See the Supplementary Data for the full list with all input parameters, their default values, and the reasons for their inclusion or exclusion by the Imperial College London CovidSim team. This list also contains a short description of the parameters.
Although we made our own considerations and decisions as to which parameters to include in the UQ study, the large number of parameters in play requires expert knowledge to make a suitable initial selection. A total of 60 of these parameters were included in a UQ campaign at some point (these are displayed separately in the Supplementary Data). We choose uninformative uniform distributions to reflect our lack of knowledge in the most likely values of these inputs, with bounds either based on data or expert knowledge.
Any input that was selected at least once for refinement during the dimension-adaptive sampling in one of the three exploratory UQ campaigns of the ‘CovidSim parameters’ section was included in the final, large-scale UQ campaign. This led to a total of 19 final parameter distributions (see Supplementary Section 3).
Important scenario parameters are R0 and two trigger parameters, which are specified via the command line. In the case of modelling a suppression strategy, the SD and PC interventions are triggered when the weekly number of new intensive care unit (ICU) cases exceeds the value supplied by the first trigger. Likewise, they are suspended when this metric drops below the second specified trigger2. The results below are conditional on the selected NPI measures, as well as fixed values for R0 and the ICU triggers.
Confidence intervals
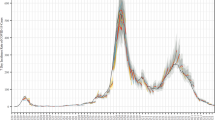
Here we consider two different PC_CI_HQ_SD suppression scenarios. The results that follow were obtained using a computational budget of 3,000 CovidSim evaluations per scenario. Figure 1a shows the 68% and 95% confidence intervals of the cumulative death prediction for \({{\mathcal{S}}}_{1}\), with R0 = 2.4 and on/off ICU triggers of 60/15. Remember that the PC and SD interventions are turned on and off based on a specified number of new weekly ICU cases (60 and 15 new cases here, which is one of the scenarios considered in Report 9). The PDF of the total death count after 800 days is also plotted. The latter shows clear non-Gaussian behaviour, with a heavy tail towards a higher death count. The corresponding Report 9 total death count2 is 8,700, whereas the current version, which now supports averaging over stochastic realizations, predicts21 9,500. Our mean prediction from equation (1) is almost double this amount. The Report 9 predictions are still captured by the distribution (at approximately the lower boundary of the 68% confidence interval), but the distribution also supports low-probability events which are about five to six times higher than those given in Report 9.
a,b, The mean cumulative death prediction for \({{\mathcal{S}}}_{1}\) (R0 = 2.4, ICU on/off triggers 60/15) (a) and \({{\mathcal{S}}}_{2}\) (R0 = 2.6, ICU on/off triggers 400/300) (b) plus confidence intervals. The PDFs of the total death count after 800 days are shown to to the right. Day zero corresponds to 1 January 2020. We also plot the observed cumulative death count data for the UK (green squares) in both figures, which were obtained from ref. 22. The striped line is a single sample from CovidSim (current release), run with the baseline parameter values of Report 9.
Note that the output distribution conditioned on \({{\mathcal{S}}}_{1}\) clearly underpredicts the observed death count in the UK, which is also plotted in Fig. 1a. We therefore selected \({{\mathcal{S}}}_{2}\) using the parameters from Report 9 that gave the highest predicted mortality (R0 = 2.6 and ICU on/off triggers of 400/300 cases; Fig. 1b). The deterministic Report 9 prediction is still located at the 68% confidence interval lower border; however, the total death count PDF is notably less skewed, although still not exactly symmetric.
Figure 1 clearly indicates that the results are very sensitive to \({\mathcal{S}}\). As noted, we also plot the observed death count validation data from ref. 22 in both subfigures, which are evidently not captured well by the output distributions, although scenario \({{\mathcal{S}}}_{2}\) does perform better than \({{\mathcal{S}}}_{1}\). It is also plain from Fig. 1 that the rate of infection starts too slowly in both cases; it must be assumed that the epidemic started earlier than suggested in Report 9, which is in line with the findings of ref. 7. Hence, if one aims to validate CovidSim in a probabilistic sense (that is, obtaining a distribution that captures validation data with high probability), it is crucial to either tune the scenario parameters or to quantify the scenario uncertainty.
CovidSim also has a number of random seeds, whose influence on the death count is examined in Supplementary Section 5. See Supplementary Section 7 for confidence intervals on quantities of interest other than the cumulative death count.
Finally, we emphasize that the authors of Report 9 did not claim that their parameterization at the time would be able to match the death count data of the coming months. The main message was that it would “…be necessary to layer multiple interventions, regardless of whether suppression or mitigation is the overarching policy goal”2, and it also showed that doing nothing at all would have disastrous consequences.
Sensitivity analysis
With sensitivity analysis, the aim is to apportion the uncertainty of the model output to specific (combinations of) input parameter uncertainties. To this end, Sobol indices measure the fraction of the output variance that each combination of input parameters is responsible for when given a distribution on the inputs23. They can be computed in a post-processing step once the input uncertainties are propagated through the computational model24 (see the ‘Sobol index calculation’ section in the Methods).
The first-order Sobol indices (Si) are defined as \({S}_{i}:= {\mathbb{V}}\left[{q}_{i}\right]/{\mathbb{V}}\left[q\right]\in [0,1]\), for i = 1, ⋯ , d. Here \({\mathbb{V}}\left[q\right]\) is the total output variance and \({\mathbb{V}}\left[{q}_{i}\right]\) is the partial variance attributed to one particular input parameter23. Figure 2 displays the three Si with the highest values for \({{\mathcal{S}}}_{1}\) and \({{\mathcal{S}}}_{2}\) (see Supplementary Section 8 for more results). The Sobol indices are plotted against time, showing that the latent period (the period in which a patient is infected but not yet infectious) is the most influential at the beginning, although only for a short amount of time. A longer latent period therefore means that the rate of disease spread is slower in this early exponential growth stage, when there are still relatively few cases present.
a,b First-order Sobol indices of the same three most dominant parameters for \({{\mathcal{S}}}_{1}\) (a) and \({{\mathcal{S}}}_{2}\) (b), plotted against time at one month intervals. The plot shows the fraction of the variance that each parameter is responsible for, over time. Blue circles, relative spatial contact rate given social distancing; orange, delay the start case isolation; green, latent period; blue stars, the sum of all 19 first-order indices; purple diamonds, the sum of the three most dominant parameters.
The second important parameter is the relative spatial contact rate given social distancing parameter, which indicates the assumed effectiveness of social distancing. Finally, the third parameter (in both scenarios) to dominate the variance is the delay to start case isolation. The latent period originally belonged to the disease parameter group of the ‘CovidSim parameters’ section, whereas the other two inputs are intervention parameters. Overall, it can be said that the intervention parameters, which influence control measures and human behaviour, are most influential. The inputs from the spatial/geographic group have a comparatively small effect.
In Fig. 2 we also plot the sum of all 19 first-order Sobol indices. This shows that first-order effects (that is, the fraction of the variance obtained by varying individual parameters) account for a little under 80% in the case of \({{\mathcal{S}}}_{1}\), and roughly 90% of the variance for \({{\mathcal{S}}}_{2}\). Conversely, interaction effects between parameters therefore account for no more than 10–20% in our chosen scenarios. We also show the sum of the first-order indices for just the three most important parameters (that is, those actually plotted in Fig. 2), which already accounts for roughly 50% and 67% of the observed variance in cumulative deaths for \({{\mathcal{S}}}_{1}\) and \({{\mathcal{S}}}_{2}\), respectively.
Uncertainty amplification
Although we based our input distributions (see Supplementary Table 1) on a combination of available data and expert knowledge, (in general) a certain level of ambiguity remains with respect to the choice of input distribution. We therefore devise a measure that examines the amplification of uncertainty in the outputs with respect to a given set of input distributions (as explained below). This relative measure of output-to-input variability is based on the coefficients of variation ratio (CVR), which serves as our robustness score and is given by
A coefficient of variation (CV) is a dimensionless quantity that measures the variability of a random variable with respect to its mean, and is defined as the standard deviation over the mean (σ/μ). In equation (2), \({\mathrm{CV}}(\bar{q})\) and \({\mathrm{CV}}(\bar{{\boldsymbol{\xi }}})\) are the mean CV of the output \(q\in {{\mathbb{R}}}^{N}\) and input \({\boldsymbol{\xi }}\in {{\mathbb{R}}}^{d}\), respectively. The results for CovidSim using equation (2) are displayed in Table 1, which shows that the uncertainty in the input is amplified by a factor of three for scenario 1. By contrast, CovidSim is more robust under \({{\mathcal{S}}}_{2}\), in which case the same input uncertainty is still amplified to the output, although now by a factor of two.
Note that \({{\mathcal{S}}}_{1}\) has a higher CVR while imposing stronger control than \({{\mathcal{S}}}_{2}\). Although the stronger control results in a much lower absolute number of predicted deaths, the output is more uncertain in a relative sense due to the long tail (see Fig. 1a), which results in a higher output CV and therefore a higher CVR.
Discussion
Conditional on a given \({\mathcal{S}}\), we found that the Report 9 predictions are captured by the parametric uncertainty at the lower bound of the 68% confidence interval. The PDF of the total death count is skewed and can support low-probability events with a predicted death count that is about five to six times higher.
We find that CovidSim amplifies the input uncertainty by 300% (that is, roughly by a factor of three; see Table 1) depending on the chosen NPI scenario. Despite this amplification of uncertainty, the distribution of the output does not envelope available validation data well for the two scenarios we considered. We do note, however, that the predictions will be very sensitive to the chosen \({\mathcal{S}}\), which therefore must be tuned if one wishes to validate CovidSim against available data (see, for example, ref. 7). Tuning the ICU triggers alone is insufficient. In Supplementary Section 4 we show the results of an additional UQ campaign where we sought to extract the best-guess ICU trigger values from data. These results are similar to those presented in the main manuscript.
Predicting the uncertainty in computational models is already considered as vitally important in weather and climate models. For instance, the author of ref. 25 claims that “…no weather or climate prediction can be considered complete without a forecast of the associated flow-dependent predictability”. We also argue that, in the case of COVID-19 predictions, a single deterministic prediction paints an incomplete picture, as we showed that such a prediction is better viewed as only one member of a much wider distribution. Hence, some measure of uncertainty is required for a correct interpretation of the results, so that those tasked with policy-making are presented with a more complete picture of the outcomes that the model is capable of predicting.
For instance, if the policymaker is presented with just the deterministic model outcome of Fig. 1b, they may draw the conclusion that the UK will suffer 50,000 deaths after approximately 600 days by adopting scenario \({{\mathcal{S}}}_{2}\). However, by taking some reasonable input uncertainty into account, we see that the same model can also predict that number in less than 200 days with the same NPI settings. Another example concerns predictions with hard thresholds (such as the maximum number of available ICU beds). A single prediction might lie on the safe side of the threshold, yet the model may exhibit a considerable non-zero probability that this threshold can be exceeded, if it were admitted that the models are uncertain. We expect that such kinds of information pertaining to uncertainty would influence the decision-making process in an important way.
Let us briefly discuss applying the proposed method to models other than CovidSim, which may well be beneficial for the same reasons mentioned above. The dimension-adaptive sampling scheme has a black-box assumption, and can therefore be applied without modification to other models; however, note that EasyVVUQ requires that a template for the input file must be created17. We used the FabSim3 automation toolkit to execute the ensembles on a supercomputer (in our case the PSNC Eagle machine26; see the Code Availability section for the relevant links to our software). In summary, (dimension-adaptive) parametric uncertainty propagation is general enough to be applied to other models and it is important to do so moving forward; however, although the dimension-adaptive approach is efficient, it is ultimately still limited by the dimension of the input space. We could not have applied our method to all inputs of CovidSim, for example.
Conclusion
To conclude, to retrofit the model’s outputs with the observed data requires additional post-hoc tuning of certain parameters that control the scenario in which the model is applied. These issues need to be addressed in seeking to provide a more quantitative albeit strongly probabilistic version of the code that might be suitable for its future application in healthcare and governmental decision-making. Our findings exemplify how sensitivity analysis and uncertainty quantification can help improve model development efforts, and in this case support the creation of epidemiological forecasting with quantified uncertainty.
As an alternative to retrofitting the scenario parameters, one could attempt to quantify the uncertainty related to the scenario the model is applied in. One such potential route for future research could involve creating cheap surrogate models for CovidSim, for example, in the stochastic space of the most influential parameters identified, which opens up the possibility of Bayesian inference27. This would allow us to update our assumptions on the input distributions and obtain posterior input distributions conditioned on observed data instead. Furthermore, such a statistical calibration can eliminate a bias between the mean prediction and real-world observations. Repeating the procedure for a discrete set of scenario parameters then allows for the combined estimation of the parametric and the scenario uncertainty using Bayesian ensemble methods (see, for example, refs. 15,16).
Methods
In this section we first describe our method for computing the statistical results and subsequently describe the uncertainty amplification factor.
Statistics
Here we describe how we compute the probability distribution of the code output, the corresponding ensemble execution and how the Sobol indices are calculated.
Dimension-adaptive uncertainty propagation
The traditional forward uncertainty quantification methods present in EasyVVUQ (for example, stochastic collocation and polynomial chaos), are subject to the curse of dimensionality. To illustrate the problem, consider first the standard stochastic collocation method, which creates a polynomial approximation of the code output q, as a function of the uncertain inputs \({\boldsymbol{\xi }}=({\xi }_{d},\cdots \ ,{\xi }_{d})\in {{\mathbb{R}}}^{d}\):
Here, \(\tilde{q}\) denotes the polynomial approximation of q, and \(q({\xi }_{{j}_{1}},\cdots \ ,{\xi }_{{j}_{d}})\) is the actual code output, evaluated at some location inside of the stochastic domain of \({\boldsymbol{\xi }}\in {{\mathbb{R}}}^{d}\). Each input ξi &isi n; ξ is assigned an independent PDF p(ξi), and the goal is to propagate these through CovidSim to examine the corresponding distribution of the output q. The basic building blocks for the SC method are one-dimensional quadrature and interpolation rules, which are extended to higher dimension through a tensor-product construction. In equation (3), \({a}_{{j}_{1}}({\xi }_{1})\otimes \cdots \otimes {a}_{{j}_{d}}({\xi }_{d})\) is the tensor product of one-dimensional Lagrange interpolation polynomials, used to interpolate the code outputs \(q({\xi }_{{j}_{1}},\cdots \ ,{\xi }_{{j}_{d}})\) to a (potentially) unsampled location ξ. For instance, unlike the Monte Carlo method, the sample locations \(({\xi }_{{j}_{1}},\cdots \ ,{\xi }_{{j}_{d}})\) are not random. Instead, each \({\xi }_{{j}_{i}}\) is a point drawn from a one-dimensional quadrature rule, used to approximate integrals weighted by the chosen input distribution p(ξi). The order of the quadrature rule for the ith input determines the number of points mi, and due to the tensor product construction the total number of code evaluations for d inputs equals M = m1 ⋅ m2 ⋯ md, or M = md if all inputs receive the same quadrature order (see Supplementary Fig. 1 for an example). The exponential increase with d, known as the curse of dimensionality, renders the SC method intractable beyond d ≈ 10. Hence, although our parameter analysis in the main article indicates that only roughly 6% of the inputs will be varied at some point, due to the large number of inputs this is far too much for such brute-force UQ methods.
A dimension-adaptive version of the stochastic collocation method (based on the work of refs. 9,28) has therefore been implemented in EasyVVUQ. It is reasonable to expect that the output q will not be equally sensitive to each input ξi. Hence, although our input space is d-dimensional, a dimension-adaptive approach banks on the existence of a lower effective dimension. The basic idea is to start with a zeroth-order quadrature rule for all inputs, and to adaptively rank order the inputs, keeping all ineffective inputs at a low (possible zeroth) order, while increasing the order of those that are effective (see Supplementary Fig. 1 for an example in two dimensions).
The dimension-adaptive approach is explained in detail in ref. 9, here we only provide a general outline. Let Λ be the set containing all selected quadrature-order multi-indices (the grey squares of Supplementary Fig. 1), which is initialized as Λ ≔ {(0, , ⋯ , 0)}. Let the forward neighbours of any multi index l be defined by the set {l + ei∣1 ≤ i ≤ d}, where ei is the elementary basis vector in the ith direction, for example, e2 = (0, 1, 0, ⋯ , 0). The forward neighbours of the set Λ are then the forward neighbours for all l ∈ Λ, which are not already in Λ. Similarly, the backward neighbours of l are given by {l − ei∣li > 0, 1 ≤ i ≤ d}. An index set Λ is said to be admissible if all backward neighbours of Λ are in Λ.
To adaptively refine the sampling plan, a look-ahead step29 is executed, where the computational model is evaluated at the new unique sample locations generated by those forward neighbours l where Λ ∪ {l} remains an admissible set, corresponding to the × symbols of Supplementary Fig. 1. For each admissible forward neighbour l, a local error measure is computed. As proposed in ref. 10, we will base our error measure on the so-called hierarchical surplus, defined as the difference between the code output q and the surrogate prediction \(\tilde{q}\), evaluated at new sample locations of an admissible forward neighbour l,
Here, XΛ is the sampling plan generated by the one-dimensional quadrature rules in Λ, and Xl is the sampling plan generated by Λ ∪ {l}. Futhermore, \({\tilde{q}}_{{{\Lambda }}}\) is the polynomial surrogate constructed from points in XΛ alone. A local error measure can now be defined as
Note that other error measures, based on quadrature errors9,30, or Sobol sensitivity indices29 can also be defined. The admissible forward neighbour with the highest error measure η(l) is added to Λ, which can cause new forward neighbours to become admissible, and the algorithm repeats.
Note that every index l = (l1, ⋯ , ld) ∈ Λ constitutes a separate tensor product of one-dimensional quadrature rules with orders given by l. Unlike the standard approach in equation (3), the SC expansion in the adaptive case is therefore constructed as a linear combination of tensor products, that is
where \(q({{\boldsymbol{\xi }}}_{{\bf{j}}}^{({\bf{l}})})=q({\xi }_{{j}_{1}}^{({l}_{1})},\cdots \ ,{\xi }_{{j}_{d}}^{({l}_{d})})\), and \({m}_{{l}_{i}}\) is the number of points generated by a one-dimensional rule of order li. The coefficients cl are computed as
see ref. 28 for details.
As equation (6) consists of a linear combination of tensor products, the choice of the quadrature rule chosen to generate the one-dimensional points substantially affects the total number of code evaluations. It is common practice to select a nested rule, which has the property that a rule of a given order contains all points generated by that same rule at lower orders. When taking linear combinations of tensor products built from nested one-dimensional rules of different order, may points will overlap. This leads to a more efficient sparse sampling plan, especially in higher dimensions. For our calculations, we employ the well-known Clenshaw–Curtis quadrature rule (see, for example, ref. 10).
Ensemble execution
Consequently, through the use of adaptive methods we make the uncertainty analysis of CovidSim tractable, but our analysis nevertheless required us to perform thousands of runs, each with its own unique set of input parameters. Specifically, we used the Eagle supercomputer at the Posnan Supercomputing and Network Centre31, which has a track record of reliably supporting large ensemble calculations. The workflows associated with these UQ/SA procedures are large, multifaceted and iterative, and to handle and curate them efficiently, we rely on the FabSim3 automation toolkit26. FabSim3 allows us to capture commonly used workflow patterns in single-line bash commands, and it automatically captures all of the relevant input parameters, output data and variables of both the job submission environment and the local machine environment in which each simulation has been executed.
Sobol index calculation
Sobol indices are variance-based sensitivity measures of a function q(ξ) with respect to its inputs \({\boldsymbol{\xi }}\in {{\mathbb{R}}}^{d}\) (refs. 23,32). Let \({\mathbb{V}}\left[{q}_{{\bf{u}}}\right]\) be a so-called partial variance, where the multi-index u can be any subset of \({\mathcal{U}}:= \{1,2,\cdots \ ,d\}\). Each partial variance measures the fraction of the total variance in the output q that can be attributed to the input parameter combination indexed by u. The Sobol indices are defined as the normalized partial variances, that is
where \({\mathbb{V}}[q]={\sum }_{u\subseteq {\mathcal{U}}}{\mathbb{V}}[{q}_{{\bf{u}}}]\) is the is the total variance of q (ref. 32). As all partial variances are positive, the sum of all possible Su equals 1.
To perform the Sobol sensitivity analysis, we employ the method described in ref. 24, which is an adaptation of a method originally proposed in ref. 33. The general idea is to transform the adaptive SC expansion into a polynomials chaos expansion (PCE) to facilitate the computation of the Sobol indices. The PCE equivalent of equation (3) reads
Here, the basis functions ϕk are usually constructed to be orthonormal to the input density, and the response coefficients ηk are normally computed via a spectral projection technique or via a regression method. Unlike equation (3), summation does not take place over the collocation points ξj. It instead takes place over multi indices \({\bf{k}}=({k}_{1},\cdots \ ,{k}_{d})\in {\mathcal{K}}\), determined by a selected truncation scheme (see below). The PCE method is a well-know technique; please refer to refs. 34,35 for more details.
The PCE method is particularly suited for sensitivity analysis, as the Sobol indices can be calculated from the response coefficients ηk in a post-processing procedure36. The PCE mean and variance (when the ϕk are orthonormal), are given by34
Similarly, the partial variances can be computed with
The multi index set \({{\mathcal{K}}}_{{\bf{u}}}\) can be interpreted as the set of all multi indices corresponding to varying only the inputs indexed by u. That is, if, for instance, u = (1, 3), \({{\mathcal{K}}}_{{\bf{u}}}\) is the subset of \({\mathcal{K}}\), with all indices k where k1 > 0 and k3 > 0, with all other kj = 0. Note that with equations (10) and (11), the Sobol indices in equation (8) are readily available, provided we have the PCE coefficients ηk.
To compute the PCE coefficients from our anisotropic sparse grid, we can transform the Lagrange basis to a PCE basis on the level of the one-dimensional basis functions24. Applying this transformation \({\mathcal{T}}\) to equation (6) yields
and so we have to apply the transformation separately to each tensor product. Equating a tensor product of equation (12) to a corresponding PCE expansion in equation (9) yields
where the PCE truncation is Λl ≔ {k∣k ≤ l}24. By using the orthogonality property of the PCE basis functions (and the independence of the input distributions), we can find an expression for each coefficient \({\eta }_{{\bf{k}}}^{({\bf{l}})}\) as
where each univariate transformation coefficient \({\nu }_{{k}_{i}}^{({l}_{i},{j}_{i})}\) is given by
This is integrated over the support of p(ξi) using Gaussian quadrature. To generate the orthonormal \({\phi }_{{k}_{i}}\) we use the Chaospy package37.
Once in possession of the \({\eta }_{{\bf{k}}}^{{\bf{l}}}\), we can compute the statistics and the Sobol indices corresponding to an adaptive sparse grid. The first two moments are given by
where \({{\mathcal{K}}}_{{\bf{l}}}:= \{{\bf{k}}| {\bf{l}}\in {{{\Lambda }}}_{{\bf{k}}},\forall {\bf{k}}\in {{\Lambda }}\}\). The expression for the variance is obtained by (1) inserting equation (6) into \({\mathbb{E}}\left[{\tilde{q}}^{2}\right]-{\mathbb{E}}{\left[\tilde{q}\right]}^{2}\); (2) grouping all terms with like k in \({\mathbb{E}}\left[{\tilde{q}}^{2}\right]\), which is what \({{\mathcal{K}}}_{{\bf{l}}}\) indicates; and (3) using the orthogonality of the ϕk to remove all cross terms ϕkϕj, j ≠ k. The statistics in equation (16) represent a more general version than those given in equation (10), and will revert to these equations when given the combination coefficients cl corresponding to a standard, non-adaptive SC grid. The partial variances \({\mathbb{V}}\left[{\tilde{q}}_{{\bf{u}}}\right]\), and by extension the Sobol indices, are computed in the same way as before, namely by summing individual variance contributions indexed by the set \({{\mathcal{K}}}_{{\bf{u}}}\) shown in equation (11).
Uncertainty amplification factor
The aim here is to find a robustness score of a computational model, under uncertainty in the input parameters. A simple (dimensionless) measure for variability in some random variable X is the CV, which is defined as the standard deviation over the mean, that is
Any forward uncertainty propagation method approximates the first two moments of the output \(q\in {{\mathbb{R}}}^{N}\), and so \(CV(q)\in {{\mathbb{R}}}^{N}\) is readily available. Assuming we can (analytically) compute the first two moments of each input ξi ∈ ξ, i = 1, ⋯ , d, \(CV({\xi }_{i})\in {\mathbb{R}}\) is also easily computed. Although ξ may contain inputs defined on vastly different scales, as the CV is a dimensionless quantity, this will not pose a problem. We propose to use the ratio of CV(Q) and CV(ξ) as a relative measure of variability between the input and the output. To do so we first have to account for the fact that in general, d ≠ N. Here we choose to average over all points:
The basic idea of equation (18) is to say something about the robustness of the code to input uncertainty, given the fact that in all likelihood the choice of input distributions can be at least partly ambiguous. We have, for instance, prescribed an input distribution for the relative household contact rate after closure, with end points located at 20% of the default value (see Supplementary Table 1). Although this was within the range suggested by expert opinion, the number of 20% is still just a user-specified choice, and it might as well have been for instance 15%. It therefore makes sense to look at the relative input-to-output uncertainty; thus, when given a user-specified average input perturbation of say 20% (\({\mathrm{CV}}(\bar{{\boldsymbol{\xi }}})=0.2\)), equation (18) tells us to what extent the code (which is a nonlinear mapping from the input to the output) amplifies this assumed uncertainty. Relative damping of uncertainty is also possible, corresponding to CVR < 1.
Data availability
Figure 1a,b displays publicly available cumulative death count data for the UK, which were obtained from ref. 22. Source Data are available with this paper. Furthermore, the parameter list—with all input parameters, a description, their default values and reasons for inclusion or exclusion from the Imperial College London CovidSim team—is available as Supplementary Data.
Code availability
The version of EasyVVUQ that was used to generate our results has been pushed to a separate, publicly available GitHub branch, see refs. 38,39 for a Zenodo link. Likewise, the FabSim3 interface between EasyVVUQ and CovidSim that was used to execute the ensembles on the PNSC Eagle supercomputer can be found in refs. 40,41.
References
Ferguson, N. M. et al. Strategies for mitigating an influenza pandemic. Nature 442, 448–452 (2006).
Ferguson, N. et al. Report 9: Impact of Non-pharmaceutical Interventions (NPIs) to Reduce COVID19 Mortality and Healthcare Demand (Imperial College London, 2020); https://www.imperial.ac.uk/mrc-global-infectious-disease-analysis/covid-19/report-9-impact-of-npis-on-covid-19/
Kucharski, A. J. et al. Early dynamics of transmission and control of COVID-19: a mathematical modelling study. Lancet Infect. Dis. 20, 553 (2020).
Clark, A et al. Global, regional, and national estimates of the population at increased risk of severe COVID-19 due to underlying health conditions in 2020: a modelling study. Lancet Glob. Health 8, E1003–E1017 (2020).
Davies, N. G Effects of non-pharmaceutical interventions on COVID-19 cases, deaths, and demand for hospital services in the UK: a modelling study. Lancet Public Health 5, E375–E385 (2020).
Kerr, C. C. et al. CovaSim: an agent-based model of COVID-19 dynamics and interventions. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.10.20097469v1 (2020).
Rice, K., Wynne, B., Martin, V. & Ackland, G. J. Effect of school closures on mortality from coronavirus disease 2019: old and new predictions. Br. Med. J. https://doi.org/10.1136/bmj.m3588 (2020).
Eglen, S. J. CODECHECK 2020-010 (Zenodo, 2020); https://zenodo.org/record/3865491#.XuPW_y-ZPGI
Gerstner, T. & Griebel, M. Dimension-adaptive tensor-product quadrature. Computing 71, 65–87 (2003).
Loukrezis, D., Römer, U. & De Gersem, H. Assessing the performance of Leja and Clenshaw–Curtis collocation for computational electromagnetics with random input data. Int. J. Uncertainty Quant. 9, 33–57 (2019).
Judd, K. L., Maliar, L., Maliar, S. & Valero, R. Smolyak method for solving dynamic economic models: lagrange interpolation, anisotropic grid and adaptive domain. J. Econ. Dyn. Control 44, 92–123 (2014).
Griebel, M. & Holtz, M. Dimension-wise integration of high-dimensional functions with applications to finance. J. Complexity 26, 455–489 (2010).
Ganapathysubramanian, B. & Zabaras, N. Sparse grid collocation schemes for stochastic natural convection problems. J. Comput. Phys. 225, 652–685 (2007).
Draper, D. Assessment and propagation of model uncertainty. J. Royal Stat. Soc. B 57, 45–70 (1995).
Meyer, P. D., Ye, M., Rockhold, M. L., Neuman, S. P. & Cantrell, K. J. Combined Estimation of Hydrogeologic Conceptual Model, Parameter, and Scenario Uncertainty with Application to Uranium Transport at the Hanford Site 300 Area Technical report (US Nuclear Regulatory Commission Office of Nuclear Regulatory Research, 2007).
Edeling, W. N., Cinnella, P. & Dwight, R. P. Predictive RANS simulations via bayesian model-scenario averaging. J. Comput. Phys. 275, 65–91 (2014).
Richardson, R. A. et al. EasyVVUQ: a library for verification, validation and uncertainty quantification in high performance computing. J. Open Res. Softw. 8, 11 (2020).
Wright, D. W. et al. Building confidence in simulation: applications of EasyVVUQ. Adv. Theory Simul. 3, 1900246 (2020).
Groen, D. et al. Introducing VECMAtk—verification, validation and uncertainty quantification for multiscale and HPC simulations. In International Conference on Computational Science 479–492 (Springer, 2019).
COVID-19 CovidSim Model—Report 9 Folder (MRC Centre for Global Infectious Disease Analysis, 2020); https://github.com/mrc-ide/covid-sim/tree/master/report9
Deaths Involving COVID-19 in the Care Sector, England and Wales: Deaths Occurring up to 12 June 2020 and Registered up to 20 June 2020 (Provisional) (Office of National Statistics, 2020); https://www.ons.gov.uk/peoplepopulationandcommunity/birthsdeathsandmarriages/deaths/articles/deathsinvolvingcovid19inthecaresectorenglandandwales/latest
Coronavirus (COVID19) in the UK. Deaths in United Kingdom (GOV.UK, 2020); https://coronavirus-staging.data.gov.uk/deaths
Sobol, I. M. On sensitivity estimation for nonlinear mathematical models. Matematich. Model. 2, 112–118 (1990).
Jakeman, J. D., Eldred, M. S., Geraci, G. & Gorodetsky, A. Adaptive multi-index collocation for uncertainty quantification and sensitivity analysis. Int. J. Numer. Methods Eng. 121, 1314–1343 (2020).
Palmer, T. N. Predicting uncertainty in forecasts of weather and climate. Rep. Prog. Phys. 63, 71 (2000).
Groen, D. et al. FabSim: facilitating computational research through automation on large-scale and distributed e-infrastructures. Comput. Phys. Commun. 207, 375–385 (2016).
Marzouk, Y. & Xiu, D. A stochastic collocation approach to Bayesian inference in inverse problems. Commun. Comput. Phys. 6, 826–847 (2009).
Gerstner, T. & Griebel, M. Numerical integration using sparse grids. Numer. Algorithms 3, 209–232 (1998).
Dwight, R. P., Desmedt, S. G. L. & Omrani, P. S. Sobol indices for dimension adaptivity in sparse grids. In Simulation-Driven Modeling and Optimization 371–395 (Springer, 2016).
Narayan, A. & Jakeman, J. D. Adaptive Leja sparse grid constructions for stochastic collocation and high-dimensional approximation. SIAM J. Sci. Comput. 36, A2952–A2983 (2014).
The Eagle Supercomputer (Poznan Supercomputing and Networking Center, 2020); https://wiki.man.poznan.pl/hpc/index.php?title=Eagle
Sobol, I. M. Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Math. Comput. Simul. 55, 271–280 (2001).
Buzzard, G. T. Global sensitivity analysis using sparse grid interpolation and polynomial chaos. Reliability Engineering & System Safety 107, 82–89 (2012).
Sudret, B. in Risk and Reliability in Geotechnical Engineering (Phoon, K.-K. & Ching, J.) 265–300 (CRC, 2015).
Xiu, D. & Karniadakis, G. E. The Wiener–Askey polynomial chaos for stochastic differential equations. SIAM J. Sci. Comput. 24, 619–644 (2002).
Sudret, B. Global sensitivity analysis using polynomial chaos expansions. Reliability Eng. Syst. Safety 93, 964–979 (2008).
Feinberg, J. & Langtangen, H. P. Chaospy: an open source tool for designing methods of uncertainty quantification. J. Comput. Sci. 11, 46–57 (2015).
VECMA Consortium. EasyVVUQ Software—CovidSim Branch (VECMA, 2020); https://github.com/UCL-CCS/EasyVVUQ/tree/CovidSim
Jancauskas, V. et al. Wedeling/EasyVVUQ: Covidsim Version (Zenodo, 2021); https://doi.org/10.5281/zenodo.4445140
VECMA Consortium. FabSim3—EasyVVUQ Interface for CovidSim (2021); https://github.com/arabnejad/FabCovidsim/tree/dev
Edeling, W. et al. FabCovidSim (Zenodo, 2021); https://doi.org/10.5281/zenodo.4445290
Acknowledgements
This project was undertaken on behalf of the Royal Society’s RAMP initiative. The work was funded as part of the European Union Horizon 2020 research and innovation programme under grant agreement nos. 800925 (VECMA project; www.vecma.eu) and 823712 (CompBioMed2 Centre of Excellence; www.compbiomed.eu), as well as the UK EPSRC for the UK High-End Computing Consortium (grant no. EP/R029598/1). We are grateful to M. Cates, G. Ackland and K. Rice for helpful discussions while this work was in progress. Finally, we would like to thank N. Ferguson for his support of our investigation and his response to our questions. The calculations were performed in part at the Poznan Supercomputing and Networking Center.
Author information
Authors and Affiliations
Contributions
W.E. performed much of the analysis, carried out many of the computational campaigns and co-wrote the paper. H.A., R.S. and D.S. performed sensitivity analysis on subgroups of the parameters within the model. B.B. and K.G. provided technical support for the computational aspects of the work. D.G., I.M. and D.C. provided scientific and technical advice. P.V.C. initiated and directed the research, and co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Computational Science thanks Kathy Leung and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ananya Rastogi was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary discussion, Figs. 1–7 and Table 1.
Supplementary Data 1
All input parameters, a description, their default values and reasons for inclusion or exclusion from the Imperial College London CovidSim team.
Source data
Source Data Fig. 1
The computed cumulative death results for Fig. 1 and the observed death count data.
Source Data Fig. 2
The Sobol indices of Fig. 2.
Rights and permissions
About this article
Cite this article
Edeling, W., Arabnejad, H., Sinclair, R. et al. The impact of uncertainty on predictions of the CovidSim epidemiological code. Nat Comput Sci 1, 128–135 (2021). https://doi.org/10.1038/s43588-021-00028-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-021-00028-9
This article is cited by
-
An epidemiological modeling framework to inform institutional-level response to infectious disease outbreaks: a Covid-19 case study
Scientific Reports (2024)
-
Confidence in Covid-19 models
Synthese (2024)
-
Dynamic calibration with approximate Bayesian computation for a microsimulation of disease spread
Scientific Reports (2023)
-
Inaccuracies of deterministic finite-element models of human middle ear revealed by stochastic modelling
Scientific Reports (2023)
-
Die voreingenommene Deutung des Unbekannten. Das Nichtwissensregime der Pandemieberatung und der Ausschluss der Sozialwissenschaften
Berliner Journal für Soziologie (2023)