Abstract
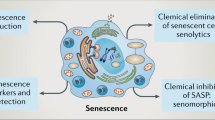
Cellular senescence is a stable type of cell cycle arrest triggered by different stresses. As such, senescence drives age-related diseases and curbs cellular replicative potential. Here, we show that 3-deazaadenosine (3DA), an S-adenosyl homocysteinase inhibitor, alleviates replicative and oncogene-induced senescence. 3DA-treated senescent cells showed reduced global histone H3 lysine 36 trimethylation, an epigenetic modification that marks the bodies of actively transcribed genes. By integrating transcriptome and epigenome data, we demonstrate that 3DA treatment affects key factors of the senescence transcriptional program. Notably, 3DA treatment alleviated senescence and increased the proliferative and regenerative potential of muscle stem cells from very old mice in vitro and in vivo. Moreover, ex vivo 3DA treatment was sufficient to enhance the engraftment of human umbilical cord blood cells in immunocompromised mice. Together, our results identify 3DA as a promising drug enhancing the efficiency of cellular therapies by restraining senescence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
All software and packages used are listed in the Reporting Summary and are publicly available. The code relevant to the ChIP-seq analysis is hosted on Zenodo (https://zenodo.org, https://doi.org/10.5281/zenodo.6865749).
References
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Salama, R., Sadaie, M., Hoare, M. & Narita, M. Cellular senescence and its effector programs. Genes Dev. 28, 99–114 (2014).
Collado, M. et al. Tumour biology: senescence in premalignant tumours. Nature 436, 642 (2005).
Krizhanovsky, V. et al. Senescence of activated stellate cells limits liver fibrosis. Cell 134, 657–667 (2008).
Munoz-Espin, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014).
Baker, D. J. et al. Naturally occurring p16(Ink4a)-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Ovadya, Y. & Krizhanovsky, V. Strategies targeting cellular senescence. J. Clin. Invest. 128, 1247–1254 (2018).
Lapasset, L. et al. Rejuvenating senescent and centenarian human cells by reprogramming through the pluripotent state. Genes Dev. 25, 2248–2253 (2011).
Latorre, E. et al. Small molecule modulation of splicing factor expression is associated with rescue from cellular senescence. BMC Cell Biol. 18, 31 (2017).
Georgilis, A. et al. PTBP1-mediated alternative splicing regulates the inflammatory secretome and the pro-tumorigenic effects of senescent cells. Cancer Cell 34, 85–102 (2018).
Xu, M. et al. JAK inhibition alleviates the cellular senescence-associated secretory phenotype and frailty in old age. Proc. Natl Acad. Sci. USA 112, E6301–E6310 (2015).
Krishnamurthy, J. et al. p16INK4a induces an age-dependent decline in islet regenerative potential. Nature 443, 453–457 (2006).
Molofsky, A. V. et al. Increasing p16INK4a expression decreases forebrain progenitors and neurogenesis during ageing. Nature 443, 448–452 (2006).
Sousa-Victor, P. et al. Geriatric muscle stem cells switch reversible quiescence into senescence. Nature 506, 316–321 (2014).
Guerrero, A. et al. Cardiac glycosides are broad-spectrum senolytics. Nat Metab. 1, 1074–1088 (2019).
Chiang, P. K. Biological effects of inhibitors of S-adenosylhomocysteine hydrolase. Pharmacol. Ther. 77, 115–134 (1998).
De La Haba, G. & Cantoni, G. L. The enzymatic synthesis of S-adenosyl-l-homocysteine from adenosine and homocysteine. J. Biol. Chem. 234, 603–608 (1959).
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–990 (2013).
Jacobs, J. J. & de Lange, T. Significant role for p16INK4a in p53-independent telomere-directed senescence. Curr. Biol. 14, 2302–2308 (2004).
Innes, A. J. et al. XPO7 is a tumor suppressor regulating p21(CIP1)-dependent senescence. Genes Dev. 35, 379–391 (2021).
d’Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Chiang, P. K. & Cantoni, G. L. Perturbation of biochemical transmethylations by 3-deazaadenosine in vivo. Biochem. Pharmacol. 28, 1897–1902 (1979).
Tan, J. et al. Pharmacologic disruption of Polycomb-repressive complex 2-mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev. 21, 1050–1063 (2007).
Wagner, E. J. & Carpenter, P. B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 13, 115–126 (2012).
Hou, P. et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 341, 651–654 (2013).
Banito, A. et al. Senescence impairs successful reprogramming to pluripotent stem cells. Genes Dev. 23, 2134–2139 (2009).
Garcia-Prat, L. et al. Autophagy maintains stemness by preventing senescence. Nature 529, 37–42 (2016).
Scarpa, S. et al. Differentiation of myoblast cell lines and biological methylation: 3-deazaadenosine stimulates formation of multinucleated myofibers. Proc. Natl Acad. Sci. USA 81, 3064–3068 (1984).
Sousa-Victor, P., Garcia-Prat, L. & Munoz-Canoves, P. Control of satellite cell function in muscle regeneration and its disruption in ageing. Nat. Rev. Mol. Cell Biol. 23, 204–226 (2022).
Kwon, S. M., Hong, S. M., Lee, Y. K., Min, S. & Yoon, G. Metabolic features and regulation in cell senescence. BMB Rep. 52, 5–12 (2019).
Yang, S. et al. Oval cells compensate for damage and replicative senescence of mature hepatocytes in mice with fatty liver disease. Hepatology 39, 403–411 (2004).
Fausto, N. & Campbell, J. S. The role of hepatocytes and oval cells in liver regeneration and repopulation. Mech. Dev. 120, 117–130 (2003).
Broxmeyer, H. E. Enhancing the efficacy of engraftment of cord blood for hematopoietic cell transplantation. Transfus. Apher. Sci. 54, 364–372 (2016).
Brunstein, C. G. & Wagner, J. E. Cord blood transplantation for adults. Vox. Sang. 91, 195–205 (2006).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Tanaka, H. et al. The NSD2/WHSC1/MMSET methyltransferase prevents cellular senescence-associated epigenomic remodeling. Aging Cell 19, e13173 (2020).
Mosteiro, L. et al. Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science 354, aaf4445 (2016).
Chiche, A. et al. Injury-Induced Senescence Enables In Vivo Reprogramming in Skeletal Muscle. Cell Stem Cell 20, 407–414 (2017).
Boitano, A. E. et al. Aryl hydrocarbon receptor antagonists promote the expansion of human hematopoietic stem cells. Science 329, 1345–1348 (2010).
Fares, I. et al. Cord blood expansion. Pyrimidoindole derivatives are agonists of human hematopoietic stem cell self-renewal. Science 345, 1509–1512 (2014).
Gupta, R. et al. Nov/CCN3 enhances cord blood engraftment by rapidly recruiting latent human stem cell activity. Cell Stem Cell 26, 527–541 (2020).
Carey, B. W. et al. Reprogramming of murine and human somatic cells using a single polycistronic vector. Proc. Natl Acad. Sci. USA 106, 157–162 (2009).
Aarts, M. et al. Coupling shRNA screens with single-cell RNA-seq identifies a dual role for mTOR in reprogramming-induced senescence. Genes Dev. 31, 2085–2098 (2017).
Guerrero, A. et al. Galactose-modified duocarmycin prodrugs as senolytics. Aging Cell 19, e13133 (2020).
Suelves, M. et al. uPA deficiency exacerbates muscular dystrophy in MDX mice. J. Cell Biol. 178, 1039–1051 (2007).
Sacco, A. et al. Short telomeres and stem cell exhaustion model Duchenne muscular dystrophy in mdx/mTR mice. Cell 143, 1059–1071 (2010).
Le, G., Lowe, D. A. & Kyba, M. Freeze injury of the tibialis anterior muscle. Methods Mol. Biol. 1460, 33–41 (2016).
Mitchell, C. & Willenbring, H. A reproducible and well-tolerated method for 2/3 partial hepatectomy in mice. Nat. Protoc. 3, 1167–1170 (2008).
Loforese, G. et al. Impaired liver regeneration in aged mice can be rescued by silencing Hippo core kinases MST1 and MST2. EMBO Mol. Med. 9, 46–60 (2017).
Jung, J. et al. CBX7 induces self-renewal of human normal and malignant hematopoietic stem and progenitor cells by canonical and non-canonical interactions. Cell Rep. 26, 1906–1918 (2019).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Martinez-Zamudio, R. I. et al. AP-1 imprints a reversible transcriptional programme of senescent cells. Nat. Cell Biol. 22, 842–855 (2020).
Acknowledgements
We are grateful to members of J. Gil’s laboratory for reagents, comments and other contributions to this project. We thank members of the Genomics LMS facility (L. Game, K. Rekopoulou and A. Ivan) and the Bioinformatics LMS facility (G. Dharmalingam, M. Karimi and H. Pallikonda) for help with RNA-seq and data processing. We thank O.C. Bing (BRC, A*STAR) for the histopathology scoring of liver sections. For the purpose of open access, the author has applied a Creative Commons Attribution license. Core support from MRC (MC_U120085810), a Development Gap Fund grant from LifeArc and Cancer Research UK (C15075/A28647) funded this research in J. Gil’s laboratory. P.M.-C. acknowledges funding from RTI2018-096068-B-I00, ERC-2016-AdG-741966, La Caixa HR17-00040, UPGRADE-H2020-825825, MWRF, Fundació La Marató-TV3, AFM, MDA and DPP-E. This work was supported by grants from the Deutsche Krebshilfe (to J.J.), the Dutch Cancer Society (to G.d.H.) and the Tekke Huizinga Fund (S.B. and G.d.H.). L.R. was supported by the Pasteur - Paris University International PhD Program and by the Fondation pour la Recherche Médicale. O.B was supported by Fondation ARC pour la Recherche sur le Cancer, INSERM-AGEMED and ANR S-ENCODE - 19-CE13-0017-01. O.B. is a Centre National de la Recherche Scientifique Research Director DR2. T.W. was funded by National Medical Research Council, Singapore through NMRC/OFLCG/003b/2018 and A*STAR through the Central Research Fund for Applied/Translational Research.
Author information
Authors and Affiliations
Contributions
A.G. designed, performed and analyzed the cell culture experiments and wrote the first draft of the manuscript. A.J.I., V.W., M.A. and N.M. designed, performed and analyzed experiments. L.R. performed the ChIP-seq experiments. P.-F.R. analyzed the ChIP-seq and RNA-seq data. O.B. designed and analyzed the ChIP-seq experiments, secured funding and wrote the manuscript. S.C.B., J.J. and A.A. designed, performed and analyzed the UCB experiments. G.d.H. designed and analyzed the UCB experiments, secured funding and wrote the manuscript. L.O. and V.M. designed, performed and analyzed the muscle stem cell experiments. E.P. and P.M.-C. designed, analyzed and wrote the muscle stem cell experiments and secured funding. A.P. designed, performed and analyzed the liver regeneration experiments. T.W. designed, analyzed and wrote the liver regeneration experiments and secured funding. J.G. conceived and designed the project, secured funding and wrote the manuscript, with all authors providing feedback.
Corresponding author
Ethics declarations
Competing interests
J.G. has acted as a consultant for Unity Biotechnology, Geras Bio, Myricx Pharma and Merck KGaA. Pfizer and Unity Biotechnology have funded research in J.G.’s laboratory (unrelated to the work presented here). J.G. owns equity in Geras Bio. J.G. and A.G. are named inventors in an MRC patent and J.G. is a named inventor in other Imperial College patents, both related to senolytic therapies (the patents are not related to the work presented here). T.W. is scientific co-founder of, and holds stakes in, Cargene Therapeutics, which develops nucleic-acid therapeutics for liver diseases (unrelated to the work presented here). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Richard Faragher, Valery Krizhanovsky and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Kriebs, in collaboration with the Nature Aging team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cellular models of senescence induced by oncogene activation and telomere uncapping.
a, IMR90 ER:RAS as a model of OIS. Quantification of immunofluorescence staining for BrdU (left) and p16INK4a (right) of IMR90 ER:RAS cells 4 days after treatment with 4OHT or vehicle (DMSO) (n = 3). b, IMR90 tet-TRF2△B△M as a model of telomere uncapping-induced senescence. c, Left, quantification of immunofluorescence staining for 53BP1 of IMR90 tet-TRF2△B△M cells after treatment with doxycycline or vehicle (DMSO) (n = 3). Right, representative immunofluorescence images. Scale bar, 20 μm. d, Left, quantification of immunofluorescence staining for p21CIP1 (n = 3). Right, representative immunofluorescence images. Scale bar, 50 μm. e, Left, quantification of immunofluorescence staining for p16INK4a (day 7, n = 2). Right, representative immunofluorescence images. Scale bar, 100 μm. f, Left, quantification of immunofluorescence staining for BrdU (day 3, n = 3). Right, representative immunofluorescence images. Scale bar, 50 μm. All statistical significances were calculated using unpaired two-tailed t-tests. All error bars represent mean ± s.d; n represents independent experiments.
Extended Data Fig. 2 Treatment with 3DA alleviates oncogene-induced senescence.
a, Representative immunofluorescence images of p16INK4a (red) in IMR90 ER:RAS cells cells 4 days after treatment with 4OHT and 10 μM 3DA or vehicle (DMSO). Scale bar, 100 μm. b, Quantification (n = 3). The statistical significance was calculated using unpaired two-tailed t-test. c, p21CIP1 protein expression in IMR90 ER:RAS cells treated with DMSO or 4OHT to induce senescence or 4OHT and 10 μM 3DA. Normalized nuclear intensity values and mean values on day 3 (left), day 6 (middle) and day 9 (right panel) are shown (n = 200 cells per condition). The statistical significance was calculated using unpaired two-tailed t-tests. d, Timeline of the experiment (left) and crystal violet staining (right). IMR90 ER:RAS cells were treated with 4OHT continuously to induce senescence. On day 6, DMSO or 10 μM 3DA were added (change of media every 3 days). Cells were fixed on day 14. e, Timeline of the experiment (left) and quantification of immunofluorescence staining for p16INK4a (middle) and BrdU (right). IMR90 ER:RAS cells were treated with DMSO for 4 weeks, or 10 μM 3DA for 2 weeks followed by DMSO for 2 weeks, or 10 μM 3DA for 4 weeks. Media was changed every three days. Treatment was started when cells were at passage 14 and ended when cells were at passage 20. At the end of the experiment, p16INK4a protein expression and BrdU-positive cells were quantified. For p16INK4a, normalized nuclear single-cell intensity values and mean of 200 cells are shown. To assess proliferation, percentage of BrdU-positive cells was measured (n = 3). Statistical significances were calculated using one-way ANOVA. All error bars represent mean ± s.d; n represents independent experiments unless otherwise stated.
Extended Data Fig. 3 Genetic or chemical inhibition of AHCY alleviates oncogene-induced senescence.
a, Left, representative images of immunofluorescence staining for AHCY (red). Scale bar, 50 μm. Right, single-cell intensities for AHCY (n = 1,000 cells per condition). The statistical significance was calculated using unpaired two-tailed t-test. b, Left, representative images of immunofluorescence staining for γH2AX (red). Right, quantification of immunofluorescence staining for γH2AX (n = 4). c, Quantification of immunofluorescence staining for BrdU of IMR90 ER:RAS cells four days after treatment with 4OHT or vehicle (DMSO) and 2.5 μM DZNep, 10 μM D-eritadenine and 4 μM TGF-β RI kinase inhibitor II as positive control. d, Expression levels of AHCY (n = 4). e, Expression levels of INK4a (encoding for p16INK4a, n = 4). f, Principal component analysis (PCA) for the experiment described in Fig. 3f. g,h, GSEA signatures from the same experiment. All statistical significances were calculated using one-way ANOVA. All error bars represent mean ± s.d; n represents independent experiments unless otherwise stated.
Extended Data Fig. 4 Contribution of histone methyltransferases and H3K36 methylation to senescence induction.
a, Quantification of immunofluorescence staining for BrdU of IMR90 ER:RAS cells 4 days after treatment with 4OHT or vehicle (DMSO) and increasing concentrations of GSK126 (an inhibitor of the H3K27 methylase EZH2), BRD4770 (an inhibitor of the H3 K9 methylase EHMT2) or EPZ004777 (an inhibitor of the H3 K79 methylase DOT1L). T, 4 μM TGF-β RI kinase inhibitor II as positive control (n = 3). All error bars represent mean ± s.d; n represents independent experiments. b, Immunoblot of protein extracts of IMR90 ER:RAS cells after 4OHT induction and treatment with 10 μM 3DA or vehicle (DMSO). Immunoblot of Histone H3 is included as a sample processing control. Immunoblots are a representative experiment out of three. c, Single-cell nuclear intensity values for H3K36me3 in a representative experiment out of 5 (n = 1000 cells per condition). The threshold used to quantify the cells stained for H3K36me3 cells in Fig. 4b, is shown as a red dashed line. d, Left, representative immunofluorescence images of histone H3 staining (red) 4 days after 4OHT induction and treatment with 10 μM 3DA or vehicle (DMSO). Scale bar, 100 μm. Right, single-cell nuclear intensity values for histone H3 in a representative experiment out of 3 (n = 1,000 cells). e, Left, representative immunofluorescence images of H3K36me3 staining (red) 6 days after treatment with 4OHT and 10 μM 3DA or vehicle (DMSO). Scale bar, 100 μm. Right, quantification (n = 4 independent experiments). All statistical significances were calculated using one-way ANOVA. All error bars represent mean ± s.d.
Extended Data Fig. 5 Transcriptional profiling after chemical or genetic inhibition of AHCY.
a, Principal component analysis (PCA) for the RNASeq experiment described in Fig. 4c. b, Principal component analysis (PCA) including data from the experiments in Figs. 3f, 4c. c, GSEA of RNA-Seq data using signatures for oncogene-induced senescence and SASP. d, GSEA of RNA-Seq data using a signature for Hallmark E2F targets. e, GSEA of H3K36me3 ChIP-Seq data using a signature for SASP. f-g, Representative genome browser snapshots showing H3K36me3 normalized signal at IL12RB2 (module 2, f) and CENPF (module 4, g) gene loci for DMSO (orange), DMSO + 4OHT (green) and 3DA + 4OHT (violet) conditions. Data are expressed as normalized counts per million reads (CPM) in 200 bp non-overlapping windows.
Extended Data Fig. 6 H3K36 methylation is needed for establishing oncogene-induced senescence.
a, Expression levels of NSD2 (n = 4). b, Expression levels of NSD3 (n = 3). c, Expression levels of SMYD2 (n = 4). d, Single-cell nuclear intensity values for H3K36me3 staining 7 days after treatment with 4OHT or vehicle (DMSO) of IMR90 ER:RAS cells infected with different pGIPZ shRNAs against NSD2, NSD3, SMYD2, AHCY or the parental pGIPZ vector (n = 1,000 cells per condition for a representative experiment out of 3). The threshold used to quantify the cells stained for H3K36me3 cells in Fig. 5b, is shown as a red dashed line. All statistical significances were calculated using one-way ANOVA. All error bars represent mean ± s.d.
Extended Data Fig. 7 3-deaazadenosine inhibits reprogramming-induced senescence.
a, Senescence induced in IMR90 cells upon expression of reprogramming factors (OSKM). b, crystal violet-stained, IMR90 cells transduced with either and empty vector or OSKM (a vector expressing reprogramming factors OCT4, SOX2, KLF4, cMYC) were treated with 1 μM 3DA or vehicle (DMSO). Images are a representative experiment out of three.
Extended Data Fig. 8 3DA rejuvenates geriatric satellite cells.
a-f, Analysis of the experiment described in Fig. 6a. a, Expression levels for mouse Cdkn1a mRNA (encoding for p21) in young (2–3 months, n = 7) versus geriatric satellite cells (28–31 months, n = 8). b, Quantification of γH2Ax intensity (arbitrary units: a.u.; n = 75–91 cells). c, Representative images of γH2Ax. d, Quantification of BrdU staining of young (2–3 months, n = 5) versus geriatric satellite cells (28–31 months, n = 7). e, f, Expression levels for mouse Myog (e) and Myh3 mRNA (f) in young (2–3 months, n = 6) versus geriatric (28–31 months, n = 5) satellite cells after the indicated treatments. g, Experimental design. Tibialis anterior muscles were injected with cardiotoxin to induce damage and regeneration. Mice were treated with vehicle or 3DA daily (10 mg/kg, i.p.) and sacrificed at 4 days post muscle injury to assess SA-β-gal activity. h, Left, quantification of SA- β -gal+ cells in the damaged area (n = 4 mice per group). Right, representative images of SA- β-gal staining in cryosections of tibialis anterior muscle. Scale bars 10 μm in c and 50 μm in h. All error bars represent mean ± s.d; n represents number of mice unless otherwise stated. Statistical significances were calculated using two-tailed unpaired t test. This figure was partly generated using Servier Medical Art, provided by Servier, licensed under a Creative Commons Attribution 3.0 unported license.
Extended Data Fig. 9 3DA treatment improves liver regeneration in aged mice.
a, Schematic representation of the experiment. Two-year old mice were treated 3 and 1 days before partial hepatectomy (PH) with 3DA or vehicle. The resected liver material was used for γH2AX staining and histopathology. 48 h post PH, the rest of the liver was harvested and proliferation level was determined by Ki67 staining. b, Right side shows representative photographs of IF staining with antibody against γH2AX and fluorescent DNA stain (DAPI). The inlay shows a magnification of positive nuclei from the respective main photograph. Left side shows the quantification. A significantly higher amount (p < 0.05) of γH2AX positive hepatocytes was detected in the control group (vehicle, n = 4) compared to experiment (3DA, n = 3), indicating a reduction in senescent cells. c, Right side shows representative photographs of IF staining with antibody against Ki67 and fluorescent DNA stain (DAPI). Left side shows the quantification. A significantly higher amount (p < 0.05) of Ki67 positive hepatocytes were detected in experimental group (3DA, n = 3) compared to control (vehicle, n = 3), indicating that a reduction in senescent hepatocytes is associated with improved proliferation. Statistical significance was calculated using the unpaired two-tailed Student’s t test. Error bars are represented as mean ± SEM; n represents number of mice. d-g, Pathological score (quantified blindly in a scale from 0–5) for the indicated parameters were assigned to H&E-stained liver sections from the experimental group (3DA, n = 4) and control group (vehicle, n = 4). Statistical significance was calculated using the unpaired two-tailed Student’s t test. Error bars are represented as mean ± SEM; n represents number of mice. This figure was partly generated using Servier Medical Art, provided by Servier, licensed under a Creative Commons Attribution 3.0 unported license.
Extended Data Fig. 10 3DA improves the engraftment of umbilical cord blood cells.
a-d, GSEA signatures for the cord blood RNA-seq experiment. e, Cord blood-derived human hematopoietic stem and progenitor (CD34 + ) cells were treated at day 1, 4 and 7 with 10 μM 3DA or DMSO and analyzed before xenotransplantation at day nine. Absolute cell numbers were determined by manual cell counting. Data are represented as mean ± SD, n = 4 independent experiments. Statistical significance was calculated using one-tailed Student’s t test. f, 1.5 × 106 Cells from the experiment described in a were transplanted into NSG mice. The engraftment of human (CD45+) cells and the percentage of primitive cells (CD34 + CD38−) in the bone marrow was analyzed by flow cytometry. For e and f, each shape (open cicle, closed circle, star, square or triangle) represents a different cord blood sample. Each shape is the average of 2‐3 transplanted mice with that cord blood sample (n = 4 independent cord blood samples). Statistical significance was calculated using one-tailed Student’s t test. g, Gating strategy for the experiment shown in Fig. 7f. h, Gating strategy for the experiment shown in Fig. 7g.
Supplementary information
Supplementary Table 1
ChIP-seq data.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guerrero, A., Innes, A.J., Roux, PF. et al. 3-Deazaadenosine alleviates senescence to promote cellular fitness and cell therapy efficiency in mice. Nat Aging 2, 851–866 (2022). https://doi.org/10.1038/s43587-022-00279-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-022-00279-9
This article is cited by
-
Healthspan versus lifespan: new medicines to close the gap
Nature Aging (2022)