Abstract
Senescence is a cell fate that contributes to multiple aging-related pathologies. Despite profound age-associated changes in skeletal muscle (SkM), whether its constituent cells are prone to senesce has not been methodically examined. Herein, using single-cell and bulk RNA sequencing and complementary imaging methods on SkM of young and old mice, we demonstrate that a subpopulation of old fibroadipogenic progenitors highly expresses p16Ink4a together with multiple senescence-related genes and concomitantly, exhibits DNA damage and chromatin reorganization. Through analysis of isolated myofibers, we also detail a senescence phenotype within a subset of old cells, governed instead by p21Cip1. Administration of a senotherapeutic intervention to old mice countered age-related molecular and morphological changes and improved SkM strength. Finally, we found that the senescence phenotype is conserved in SkM from older humans. Collectively, our data provide compelling evidence for cellular senescence as a hallmark and potentially tractable mediator of SkM aging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The scRNA-seq (GSE172410), myofiber RNA-seq data (GSE172254) and SkM RNA-seq data (GSE184348) generated in this study are deposited in the Gene Expression Omnibus. Human SkM RNA-seq data are publicly available at the Gene Expression Omnibus (GSE97084).
An interactive website for the SkM scRNA-seq dataset can be found at https://mayoxz.shinyapps.io/Muscle/.
Code availability
Codes and all other data are available from the corresponding author upon reasonable request.
References
Fielding, R. A. et al. Sarcopenia: an undiagnosed condition in older adults. Current consensus definition: prevalence, etiology, and consequences. International working group on sarcopenia. J. Am. Med. Dir. Assoc. 12, 249–256 (2011).
Campisi, J. & di Fagagna, F. D. Cellular senescence: when bad things happen to good cells. Nat. Rev. Mol. Cell Biol. 8, 729–740 (2007).
Coppe, J. P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6, 2853–2868 (2008).
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–U221 (2013).
Khosla, S., Farr, J. N., Tchkonia, T. & Kirkland, J. L. The role of cellular senescence in ageing and endocrine disease. Nat. Rev. Endocrinol. 16, 263–275 (2020).
Hernandez-Segura, A., Nehme, J. & Demaria, M. Hallmarks of cellular senescence. Trends Cell Biol. 28, 436–453 (2018).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Welle, S. et al. Skeletal muscle gene expression profiles in 20–29-year-old and 65–71-year-old women. Exp. Gerontol. 39, 369–377 (2004).
Kayo, T., Allison, D. B., Weindruch, R. & Prolla, T. A. Influences of aging and caloric restriction on the transcriptional profile of skeletal muscle from rhesus monkeys. Proc. Natl Acad. Sci. USA 98, 5093–5098 (2001).
Edwards, M. G. et al. Gene expression profiling of aging reveals activation of a p53-mediated transcriptional program. BMC Genomics 8, 80 (2007).
Dungan, C. M. et al. In vivo analysis of γH2AX+ cells in skeletal muscle from aged and obese humans. FASEB J. 34, 7018–7035 (2020).
Shoji, H. & Miyakawa, T. Age-related behavioral changes from young to old age in male mice of a C57BL/6J strain maintained under a genetic stability program. Neuropsychopharmacol. Rep. 39, 100–118 (2019).
Hewitt, G. et al. Telomeres are favoured targets of a persistent DNA damage response in ageing and stress-induced senescence. Nat. Commun. 3, 708 (2012).
Giordani, L. et al. High-dimensional single-cell cartography reveals novel skeletal muscle-resident cell populations. Mol. Cell 74, 609 (2019).
Sousa-Victor, P. et al. Geriatric muscle stem cells switch reversible quiescence into senescence. Nature 506, 316 (2014).
Groen, B. B. L. et al. Skeletal muscle capillary density and microvascular function are compromised with aging and type 2 diabetes. J. Appl. Physiol. 116, 998–1005 (2014).
Coppe, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Anerillas, C., Abdelmohsen, K. & Gorospe, M. Regulation of senescence traits by MAPKs. Geroscience 42, 397–408 (2020).
Tominaga, K. & Suzuki, H. I. TGF-β signaling in cellular senescence and aging-related pathology. Int. J. Mol. Sci. 20, 5002 (2019).
Wiley, C. D. & Campisi, J. From ancient pathways to aging cells-connecting metabolism and cellular senescence. Cell Metab. 23, 1013–1021 (2016).
Avelar, R. A. et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. 21, 91 (2020).
Tront, J. S., Hoffman, B. & Liebermann, D. A. Gadd45a suppresses Ras-driven mammary tumorigenesis by activation of c-Jun NH2-terminal kinase and p38 stress signaling resulting in apoptosis and senescence. Cancer Res. 66, 8448–8454 (2006).
Wei, Z. et al. Pan-senescence transcriptome analysis identified RRAD as a marker and negative regulator of cellular senescence. Free Radic. Biol. Med. 130, 267–277 (2019).
Anderson, G. et al. RUNX-mediated growth arrest and senescence are attenuated by diverse mechanisms in cells expressing RUNX1 fusion oncoproteins. J. Cell. Biochem. 119, 2750–2762 (2018).
Casella, G. et al. Transcriptome signature of cellular senescence. Nucleic Acids Res. 47, 7294–7305 (2019).
Lemster, B. H. et al. Induction of CD56 and TCR-independent activation of T cells with aging. J. Immunol. 180, 1979–1990 (2008).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660 (2017).
Sasaki, M., Miyakoshi, M., Sato, Y. & Nakanuma, Y. Modulation of the microenvironment by senescent biliary epithelial cells may be involved in the pathogenesis of primary biliary cirrhosis. J. Hepatol. 53, 318–325 (2010).
Marques, J. C. M. T. et al. Identification of new genes associated to senescent and tumorigenic phenotypes in mesenchymal stem cells. Sci. Rep. 7, 17837 (2017).
Lopez-Dominguez, J. A. et al. Cdkn1a transcript variant 2 is a marker of aging and cellular senescence. Aging 13, 13380–13392 (2021).
Robinson, M. M. et al. Enhanced protein translation underlies improved metabolic and physical adaptations to different exercise training modes in young and old humans. Cell Metab. 25, 581–592 (2017).
Heredia, J. E. et al. Type 2 Innate signals stimulate fibro/adipogenic progenitors to facilitate muscle regeneration. Cell 153, 376–388 (2013).
Kuswanto, W. et al. Poor repair of skeletal muscle in aging mice reflects a defect in local, interleukin-33-dependent accumulation of regulatory T cells. Immunity 44, 355–367 (2016).
Lukjanenko, L. et al. Aging disrupts muscle stem cell function by impairing matricellular WISP1 secretion from fibro-adipogenic progenitors. Cell Stem Cell 24, 433 (2019).
Biferali, B., Proietti, D., Mozzetta, C. & Madaro, L. Fibro-adipogenic progenitors cross-talk in skeletal muscle: the social network. Front. Physiol. 10, 1074 (2019).
Saito, Y., Chikenji, T. S., Matsumura, T., Nakano, M. & Fujimiya, M. Exercise enhances skeletal muscle regeneration by promoting senescence in fibro-adipogenic progenitors. Nat. Commun. 11, 889 (2020).
Hall, B. M. et al. p16(Ink4a) and senescence-associated β-galactosidase can be induced in macrophages as part of a reversible response to physiological stimuli. Aging 9, 1867–1884 (2017).
Fry, C. S. et al. Inducible depletion of satellite cells in adult, sedentary mice impairs muscle regenerative capacity without affecting sarcopenia. Nat. Med. 21, 76–80 (2015).
Keefe, A. C. et al. Muscle stem cells contribute to myofibres in sedentary adult mice. Nat. Commun. 6, 7087 (2015).
Englund, D. A. et al. Depletion of resident muscle stem cells negatively impacts running volume, physical function, and muscle fiber hypertrophy in response to lifelong physical activity. Am. J. Physiol. Cell Physiol. 318, C1178–C1188 (2020).
Perez, K. et al. Single nuclei profiling identifies cell specific markers of skeletal muscle aging, sarcopenia and senescence. Preprint at medRxiv https://doi.org/10.1101/2021.01.22.21250336 (2021).
Takeuchi, S. et al. Intrinsic cooperation between p16INK4a and p21Waf1/Cip1 in the onset of cellular senescence and tumor suppression in vivo. Cancer Res. 70, 9381–9390 (2010).
Alcorta, D. A. et al. Involvement of the cyclin-dependent kinase inhibitor p16 (INK4a) in replicative senescence of normal human fibroblasts. Proc. Natl Acad. Sci. USA 93, 13742–13747 (1996).
Sapieha, P. & Mallette, F. A. Cellular senescence in postmitotic cells: beyond growth arrest. Trends Cell Biol. 28, 595–607 (2018).
Jurk, D. et al. Postmitotic neurons develop a p21-dependent senescence-like phenotype driven by a DNA damage response. Aging Cell 11, 996–1004 (2012).
Anderson, R. et al. Length-independent telomere damage drives post-mitotic cardiomyocyte senescence. EMBO J. 38, e100492 (2019).
Farr, J. N. et al. Targeting cellular senescence prevents age-related bone loss in mice. Nat. Med. 23, 1072 (2017).
Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med. 24, 1246–1256 (2018).
Zhu, Y. et al. The Achilles’ heel of senescent cells: from transcriptome to senolytic drugs. Aging Cell 14, 644–658 (2015).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Li, Y., Lee, Y. I. & Thompson, W. J. Changes in aging mouse neuromuscular junctions are explained by degeneration and regeneration of muscle fiber segments at the synapse. J. Neurosci. 31, 14910–14919 (2011).
Schafer, M. J. et al. Exercise prevents diet-induced cellular senescence in adipose tissue. Diabetes 65, 1606–1615 (2016).
LeBrasseur, N. K. et al. Myostatin inhibition enhances the effects of exercise on performance and metabolic outcomes in aged mice. J. Gerontol. A Biol. Sci. Med. Sci. 64, 940–948 (2009).
Mayeuf-Louchart, A. et al. MuscleJ: a high-content analysis method to study skeletal muscle with a new Fiji tool. Skelet. Muscle 8, 25 (2018).
da Silva, P. F. L. et al. The bystander effect contributes to the accumulation of senescent cells in vivo. Aging Cell 18, e12848 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Franzen, O., Gan, L. M. & Bjorkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA-sequencing data. Database 2019, baz046 (2019).
Ouyang, J. F., Kamaraj, U. S., Cao, E. Y. & Rackham, O. J. L. ShinyCell: simple and sharable visualization of single-cell gene expression data. Bioinformatics 37, 3374–3376 (2021).
Yu, G. C., Wang, L. G., Han, Y. Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Marinkovic, M. et al. Fibro-adipogenic progenitors of dystrophic mice are insensitive to NOTCH regulation of adipogenesis. Life Sci. Alliance 2, e201900437 (2019).
Gallot, Y. S., Hindi, S. M., Mann, A. K. & Kumar, A. Isolation, culture, and staining of single myofibers. Bio Protoc. 6, e1942 (2016).
Acknowledgements
We are grateful for the support of the National Institutes of Health, National Institute on Aging for grants P01 AG062413, R01 AG055529 and R56 AG060907 to N.K.L. and R01 AG068048 and UG3CA 268103 to J.F.P. and T32 AG049672 to D.A.E. This work was also supported by the Glenn Foundation for Medical Research and the Pritzker Foundation (N.K.L.). X.Z. was supported by a Robert and Arlene Kogod Center on Aging Career Development Award. L.H. was supported by NIHR Newcastle Biomedical Research Centre grant awarded to the Newcastle upon Tyne Hospitals NHS Foundation Trust and Newcastle University. R.A.F. and D.A.R. are partially supported by the US Department of Agriculture under agreement no. 58-8050-9-004 and by and by National Institutes of Health Boston Claude D Pepper Center (OAIC; 1P30AG031679). Any opinions, findings, conclusions or recommendations expressed in this publication are those of the authors and do not necessarily reflect the view of the US Department of Agriculture. We also thank members of P01 AG062413 (PI, S.K.) for helpful discussions, J.M. Cunningham and E. Wieben within the Mayo Clinic Genome Analysis Core for RNA-seq, Y. Li within the Division of Computational Biology and Department of Quantitative Health Sciences for assistance with bioinformatic analyses and staff at the Optical Microscopy Core within the Mayo Clinic Center for Cell Signaling in Gastroenterology (P30 DK084567) for guidance and use of imaging equipment.
Author information
Authors and Affiliations
Contributions
Conceptualization was carried out by X.Z. and N.K.L. X.Z., L.H., K.S.N., M.J.S., J.F.P. and N.K.L. were responsible for the methodology. Investigation was conducted by X.Z., L.H., Z.A., Y.E.N., A.E.S., M.M.R., D.A.R., S.D., V.M.P., T.A.W., A.J.H., A.B.L., S.K.J., D.A.E. and M.J.S. Resources were the responsibility of M.R., S.D., I.R.L., R.A.F. and K.S.N. Writing of the original draft was conducted by X.Z., L.H., Z.A., J.F.P. and N.K.L. Reviewing and editing was carried out by X.Z., L.H., Z.A., A.E.S., M.M.R., D.A.R., S.D., V.M.P., T.A.W., Y.E.N., A.B.L., S.K.J., D.A.E., A.G., A.A.S., D.J., I.R.L., S.K., R.A.F., K.S.N., M.J.S., J.F.P. and N.K.L. A.G., A.A.S., D.J., J.F.P. and N.K.L. were responsible for supervision. J.F.P. and N.K.L. were responsible for funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks William Keyes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The age-related loss of skeletal mass, strength, and function.
a–c, Comparisons of body weight (a), lean mass (%) (b), and quadriceps (quad) muscle weight (c) between young (n = 9) and old (n = 5) mice. d, e, Representative images of quad muscle cross-sections stained for Laminin (d) and quantification of fiber size and distribution for young and old female mice (n = 4 per group) (e). f–h, Comparison of grip strength normalized to body weight (f), treadmill exercise capacity (g), and Rotarod endurance (h) between young (n = 9) and old (n = 5) mice. Two-tailed unpaired t-test was used; error bars represent s.e.m. **, and *** denote p < 0.01 and 0.001, respectively.
Extended Data Fig. 2 The age-related increase of lipofuscin in skeletal muscle (SkM).
a,b, Representative images of Sudan Black B (SBB) staining (a) and quantification of the numbers of positive foci per quadriceps section (b) from young and old female mice (n = 4 per age group). Two-tailed unpaired t test was used; error bars represent s.e.m. * and ** denote p < 0.05 and 0.01, respectively.
Extended Data Fig. 3 Identification of skeletal muscle (SkM) cell populations by single cell RNA-sequencing.
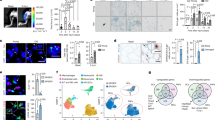
a, Heat map of the genes delineating 10 distinct cell populations in young and old female mice (n = 3 per group). b,c, UMAP plot (b) and dot plot (c) showing the main markers of each cell type. d, Relative abundance of the distinct SkM cell populations in individual mice.
Extended Data Fig. 4 Gene Ontology analysis of the upregulated and downregulated genes in high p16-expressing FAP cluster 3 compared to other FAPs.
BP: biological process; MF: molecular function; CC: cellular component. Benjamini–Hochberg Procedure was used to calculate the FDR adjusted p value.
Extended Data Fig. 5 Gene Ontology analysis of the upregulated and downregulated genes in old p21high myofibers compared to old p21low myofibers.
BP: biological process; MF: molecular function; CC: cellular component. Benjamini–Hochberg Procedure was used to calculate the FDR adjusted p value.
Supplementary information
Supplementary information
Key Resources Table.
Source data
Source Data Fig. 1
Statistical Source Data for Fig. 1.
Source Data Fig. 2
Statistical Source Data for Fig. 2.
Source Data Fig. 4
Statistical Source Data for Fig. 4.
Source Data Fig. 5
Statistical Source Data for Fig. 5.
Source Data Fig. 5
Uncropped western blot gel image for Fig. 5b.
Source Data Fig. 6
Statistical Source Data for Fig. 6.
Source Data Fig. 7
Statistical Source Data for Fig. 7.
Source Data Extended Data Fig. 1
Statistical Source Data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical Source Data for Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Zhang, X., Habiballa, L., Aversa, Z. et al. Characterization of cellular senescence in aging skeletal muscle. Nat Aging 2, 601–615 (2022). https://doi.org/10.1038/s43587-022-00250-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-022-00250-8
This article is cited by
-
Pro-ferroptotic signaling promotes arterial aging via vascular smooth muscle cell senescence
Nature Communications (2024)
-
Telomeres, cellular senescence, and aging: past and future
Biogerontology (2024)
-
Translatability of mouse muscle-aging for humans: the role of sex
GeroScience (2024)
-
Senolytic treatment does not mitigate oxidative stress-induced muscle atrophy but improves muscle force generation in CuZn superoxide dismutase knockout mice
GeroScience (2024)
-
Single-cell Mayo Map (scMayoMap): an easy-to-use tool for cell type annotation in single-cell RNA-sequencing data analysis
BMC Biology (2023)