Abstract
The FOXM1 transcription factor exhibits pleiotropic C-terminal transcriptional and N-terminal non-transcriptional functions in various biological processes critical for cellular homeostasis. We previously found that FOXM1 repression during cellular aging underlies the senescence phenotypes, which were vastly restored by overexpressing transcriptionally active FOXM1. Yet, it remains unknown whether increased expression of FOXM1 can delay organismal aging. Here, we show that in vivo cyclic induction of an N-terminal truncated FOXM1 transgene on progeroid and naturally aged mice offsets aging-associated repression of full-length endogenous Foxm1, reinstating both transcriptional and non-transcriptional functions. This translated into mitigation of several cellular aging hallmarks, as well as molecular and histopathological progeroid features of the short-lived Hutchison–Gilford progeria mouse model, significantly extending its lifespan. FOXM1 transgene induction also reinstated endogenous Foxm1 levels in naturally aged mice, delaying aging phenotypes while extending their lifespan. Thus, we disclose that FOXM1 genetic rewiring can delay senescence-associated progeroid and natural aging pathologies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Targeted-transcriptome sequencing data have been deposited in NCBI’s Gene Expression Omnibus and are available under accession number GSE193147. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Kirkwood, T. B. L. Understanding the odd science of aging. Cell 120, 437–447 (2005).
Macedo, J. C., Vaz, S. & Logarinho, E. Mitotic dysfunction associated with aging hallmarks. Adv. Exp. Med. Biol. 1002, 153–188 (2017).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Melo-Pereira, S., Ribeiro, R. & Logarinho, E. Approaches towards longevity: reprogramming, senolysis, and improved mitotic competence as anti-aging therapies. Int. J. Mol. Sci. 20, 938 (2019).
Baker, D. J. et al. BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nat. Genet. 36, 744–749 (2004).
Sieben, C. J. et al. BubR1 allelic effects drive phenotypic heterogeneity in mosaic-variegated aneuploidy progeria syndrome. J. Clin. Invest. 130, 171–188 (2020).
Wan, X. et al. Identification of FoxM1–Bub1b signaling pathway as a required component for growth and survival of rhabdomyosarcoma. Cancer Res. 72, 5889–5899 (2012).
Macedo, J. C. et al. FoxM1 repression during human aging leads to mitotic decline and aneuploidy-driven full senescence. Nat. Commun. 9, 2834 (2018).
Laoukili, J. et al. Activation of FoxM1 during G2 requires cyclin A/Cdk-dependent relief of autorepression by the FoxM1 N-terminal domain. Mol. Cell. Biol. 28, 3076–3087 (2008).
Zona, S., Bella, L., Burton, M. J., Nestal de Moraes, G. & Lam, E. W. F. FOXM1: an emerging master regulator of DNA damage response and genotoxic agent resistance. Biochim. Biophys. Acta 1839, 1316–1322 (2014).
Alvarez-Fernández, M. & Medema, R. H. Novel functions of FoxM1: from molecular mechanisms to cancer therapy. Front. Oncol. 3, 30 (2013).
Black, M. et al. FOXM1 nuclear transcription factor translocates into mitochondria and inhibits oxidative phosphorylation. Mol. Biol. Cell https://doi.org/10.1091/mbc.E19-07-0413 (2020).
Limzerwala, J. F. et al. FoxM1 insufficiency hyperactivates Ect2–RhoA–mDia1 signaling to drive cancer. Nat. Cancer 1, 1010–1024 (2020).
De Sandre-Giovannoli, A. et al. Lamin A truncation in Hutchinson–Gilford progeria. Science 300, 2055 (2003).
Eriksson, M. et al. Recurrent de novo point mutations in lamin A cause Hutchinson–Gilford progeria syndrome. Nature 423, 293–298 (2003).
Worman, H. J. & Foisner, R. The nuclear envelope from basic biology to therapy. Biochem. Soc. Trans. 38, 253–256 (2010).
Ashapkin, V. V., Kutueva, L. I., Kurchashova, S. Y. & Kireev, I. I. Are there common mechanisms between the Hutchinson–Gilford progeria syndrome and natural aging? Front. Genet. 10, 455 (2019).
Carrero, D., Soria-Valles, C. & López-Otín, C. Hallmarks of progeroid syndromes: lessons from mice and reprogrammed cells. Dis. Model. Mech. 9, 719–735 (2016).
Hennekam, R. C. M. Hutchinson–Gilford progeria syndrome: review of the phenotype. Am. J. Med. Genet. A 140, 2603–2624 (2006).
Osorio, F. G. et al. Hutchinson–Gilford progeria: splicing-directed therapy in a new mouse model of human accelerated aging. Sci. Transl. Med. 3, 106ra107–106ra107 (2011).
U.S. Food and Drug Administration. FDA approves first treatment for Hutchinson–Gilford Progeria Syndrome and some progeroid laminopathies. https://www.fda.gov/news-events/press-announcements/fda-approves-first-treatment-hutchinson-gilford-progeria-syndrome-and-some-progeroid-laminopathies (2021).
Gordon, L. B. et al. Association of lonafarnib treatment vs no treatment with mortality rate in patients with Hutchinson–Gilford progeria syndrome. JAMA 319, 1687–1695 (2018).
Gordon, L. B. et al. Impact of farnesylation inhibitors on survival in Hutchinson–Gilford progeria syndrome. Circulation 130, 27–34 (2014).
Misteli, T. Farnesyltransferase inhibition in HGPS. Cell 184, 293 (2021).
Davies, B. S. J. et al. An accumulation of non-farnesylated prelamin A causes cardiomyopathy but not progeria. Hum. Mol. Genet. 19, 2682–2694 (2010).
Beyret, E. et al. Single-dose CRISPR–Cas9 therapy extends lifespan of mice with Hutchinson–Gilford progeria syndrome. Nat. Med. 25, 419–422 (2019).
Santiago-Fernández, O. et al. Development of a CRISPR–Cas9-based therapy for Hutchinson–Gilford progeria syndrome. Nat. Med. 25, 423–426 (2019).
Scaffidi, P. & Misteli, T. Reversal of the cellular phenotype in the premature aging disease Hutchinson–Gilford progeria syndrome. Nat. Med. 11, 440–445 (2005).
Saxena, S. & Kumar, S. Pharmacotherapy to gene editing: potential therapeutic approaches for Hutchinson–Gilford progeria syndrome. GeroScience https://doi.org/10.1007/s11357-020-00167-3 (2020).
Kudlow, B. A., Stanfel, M. N., Burtner, C. R., Johnston, E. D. & Kennedy, B. K. Suppression of proliferative defects associated with processing-defective lamin A mutants by hTERT or inactivation of p53. Mol. Biol. Cell 19, 5238–5248 (2008).
Liu, B. et al. Genomic instability in laminopathy-based premature aging. Nat. Med. 11, 780–785 (2005).
Wang, I. C. et al. Increased expression of FoxM1 transcription factor in respiratory epithelium inhibits lung sacculation and causes Clara cell hyperplasia. Dev. Biol. 347, 301–314 (2010).
Halasi, M. & Gartel, A. L. A novel mode of FoxM1 regulation: positive auto-regulatory loop. Cell Cycle 8, 1966–1967 (2009).
Musich, P. R. & Zou, Y. DNA-damage accumulation and replicative arrest in Hutchinson–Gilford progeria syndrome. Biochem. Soc. Trans. 39, 1764–1769 (2011).
Musich, P. R. & Zou, Y. Genomic instability and DNA damage responses in progeria arising from defective maturation of prelamin A. Aging 1, 28–37 (2009).
Zhang, W., Qu, J., Liu, G. H. & Belmonte, J. C. I. The ageing epigenome and its rejuvenation. Nat. Rev. Mol. Cell Biol. 21, 137–150 (2020).
Arancio, W., Pizzolanti, G., Genovese, S. I., Pitrone, M. & Giordano, C. Epigenetic involvement in Hutchinson–Gilford progeria syndrome: a mini-review. Gerontology 60, 197–203 (2014).
Shumaker, D. K. et al. Mutant nuclear lamin A leads to progressive alterations of epigenetic control in premature aging. Proc. Natl Acad. Sci. USA 103, 8703–8708 (2006).
Benayoun, B. A., Pollina, E. A. & Brunet, A. Epigenetic regulation of ageing: linking environmental inputs to genomic stability. Nat. Rev. Mol. Cell Biol. 16, 593–610 (2015).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733 (2016).
Fong, L. G. et al. Heterozygosity for Lmna deficiency eliminates the progeria-like phenotypes in Zmpste24-deficient mice. Proc. Natl Acad. Sci. USA 101, 18111–18116 (2004).
Varga, R. et al. Progressive vascular smooth muscle cell defects in a mouse model of Hutchinson–Gilford progeria syndrome. Proc. Natl Acad. Sci. USA 103, 3250–3255 (2006).
Stehbens, W. E., Delahunt, B., Shozawa, T. & Gilbert-Barness, E. Smooth muscle cell depletion and collagen types in progeric arteries. Cardiovasc. Pathol. 10, 133–136 (2001).
Olive, M. et al. Cardiovascular pathology in Hutchinson–Gilford progeria: correlation with the vascular pathology of aging. Arterioscler. Thromb. Vasc. Biol. 30, 2301–2309 (2010).
Wang, Y. et al. Epidermal expression of the truncated prelamin A causing Hutchinson–Gilford progeria syndrome: effects on keratinocytes, hair and skin. Hum. Mol. Genet. 17, 2357–2369 (2008).
Kurban, R. S. & Bhawan, J. Histologic changes in skin associated with aging. J. Dermatol. Surg. Oncol. 16, 908–914 (1990).
Merideth, M. A. et al. Phenotype and course of Hutchinson–Gilford progeria syndrome. N. Engl. J. Med. 358, 592–604 (2008).
Baker, D. J. et al. Naturally occurring p16 Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Alam, H., Sehgal, L., Kundu, S. T., Dalal, S. N. & Vaidya, M. M. Novel function of keratins 5 and 14 in proliferation and differentiation of stratified epithelial cells. Mol. Biol. Cell 22, 4068–4078 (2011).
Komutrattananont, P., Mahakkanukrauh, P. & Das, S. Morphology of the human aorta and age-related changes: anatomical facts. Anat. Cell Biol. 52, 109–114 (2019).
Kim, P. H. et al. Disrupting the LINC complex in smooth muscle cells reduces aortic disease in a mouse model of Hutchinson–Gilford progeria syndrome. Sci. Transl. Med. 10, eaat7163 (2018).
Turner, M. D., Nedjai, B., Hurst, T. & Pennington, D. J. Cytokines and chemokines: at the crossroads of cell signalling and inflammatory disease. Biochim. Biophys. Acta 1843, 2563–2582 (2014).
Barger, C. J., Branick, C., Chee, L. & Karpf, A. R. Pan-cancer analyses reveal genomic features of FOXM1 overexpression in cancer. Cancers 11, 251 (2019).
Mau, T. & Yung, R. Adipose tissue inflammation in aging. Exp. Gerontol. 105, 27–31 (2018).
Conboy, I. M. & Rando, T. A. Aging, stem cells and tissue regeneration: lessons from muscle. Cell Cycle 4, 407–410 (2005).
Tyrrell, D. J. & Goldstein, D. R. Ageing and atherosclerosis: vascular intrinsic and extrinsic factors and potential role of IL-6. Nat. Rev. Cardiol. 18, 58–68 (2021).
Pisano, C., Balistreri, C. R., Ricasoli, A. & Ruvolo, G. Cardiovascular disease in ageing: an overview on thoracic aortic aneurysm as an emerging inflammatory disease. Mediators Inflamm. 2017, 1274034 (2017).
Helske, S. et al. Complement system is activated in stenotic aortic valves. Atherosclerosis 196, 190–200 (2008).
Tuttle, L. J., Sinacore, D. R. & Mueller, M. J. Intermuscular adipose tissue is muscle specific and associated with poor functional performance. J. Aging Res. 2012, 172957 (2012).
Wiedmer, P. et al. Sarcopenia–molecular mechanisms and open questions. Ageing Res. Rev. 65, 101200 (2021).
Grevendonk, L. et al. Impact of aging and exercise on skeletal muscle mitochondrial capacity, energy metabolism, and physical function. Nat. Commun. 12, 4773 (2021).
Koves, T. R. et al. Mitochondrial overload and incomplete fatty-acid oxidation contribute to skeletal muscle insulin resistance. Cell Metab. 7, 45–56 (2008).
Li, X. et al. Direct and indirect effects of growth hormone receptor ablation on liver expression of xenobiotic metabolizing genes. Am. J. Physiol. 305, E942–E950 (2013).
Sousa-Victor, P. et al. Geriatric muscle stem cells switch reversible quiescence into senescence. Nature 506, 316–321 (2014).
Tyshkovskiy, A. et al. Identification and application of gene expression signatures associated with lifespan extension. Cell Metab. 30, 573–593 (2019).
Mu, X. et al. Cytoskeleton stiffness regulates cellular senescence and innate immune response in Hutchinson–Gilford Progeria Syndrome. Aging Cell 19, e13152 (2020).
Rovillain, E. et al. Activation of nuclear factor-kappa B signalling promotes cellular senescence. Oncogene 30, 2356–2366 (2011).
Prokocimer, M., Barkan, R. & Gruenbaum, Y. Hutchinson–Gilford progeria syndrome through the lens of transcription. Aging Cell 12, 533–543 (2013).
Sun, S. et al. Vascular endothelium–targeted Sirt7 gene therapy rejuvenates blood vessels and extends lifespan in a Hutchinson–Gilford progeria model. Sci. Adv. 6, eaay5556 (2020).
Yu, W. et al. Silencing forkhead box M1 promotes apoptosis and autophagy through SIRT7/mTOR/IGF2 pathway in gastric cancer cells. J. Cell. Biochem. 119, 9090–9098 (2018).
Paez‐Ribes, M., González‐Gualda, E., Doherty, G. J. & Muñoz‐Espín, D. Targeting senescent cells in translational medicine. EMBO Mol. Med. 11, e10234 (2019).
Maloberti, A. et al. Vascular aging and disease of the large vessels: role of inflammation. High Blood Press. Cardiovasc. Prev. 26, 175–182 (2019).
Ferreira, F. J., Carvalho, L., Logarinho, E. & Bessa, J. foxm1 modulates cell non-autonomous response in Zebrafish skeletal muscle homeostasis. Cells 10, 1241 (2021).
Hou, Y. et al. The transcription factor Foxm1 is essential for the quiescence and maintenance of hematopoietic stem cells. Nat. Immunol. 16, 810–818 (2015).
Chen, Z. et al. A Cdh1–Foxm1–Apc axis controls muscle development and regeneration. Cell Death Dis. 11, 180 (2020).
Anisimov, V. N. Lifespan extension and cancer risk: myths and reality. Exp. Gerontol. 36, 1101–1136 (2001).
Sotillo, R. et al. Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell 11, 9–23 (2007).
Ustiyan, V. et al. FOXF1 transcription factor promotes lung morphogenesis by inducing cellular proliferation in fetal lung mesenchyme. Dev. Biol. 443, 50–63 (2018).
Kalinichenko, V. V. et al. Ubiquitous expression of the forkhead box M1B transgene accelerates proliferation of distinct pulmonary cell types following lung injury. J. Biol. Chem. 278, 37888–37894 (2003).
Laws, N. & Hoey, A. Progression of kyphosis in mdx mice. J. Appl. Physiol. 97, 1970–1977 (2004).
Papp, A. C. et al. AmpliSeq transcriptome analysis of human alveolar and monocyte-derived macrophages over time in response to Mycobacterium tuberculosis infection. PLoS ONE13, e0198221 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Coppé, J. P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6, 2853–2868 (2008).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660 (2017).
Subramanian, A. et al. Gene-set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Sampaio-Pinto, V. et al. Neonatal apex resection triggers cardiomyocyte proliferation, neovascularization and functional recovery despite local fibrosis. Stem Cell Reports 10, 860–874 (2018).
Acknowledgements
We thank the personnel at i3S Scientific Platforms for technical support: animal facility (S. Lamas), CCGEN (P. Magalhães), bioimaging (M. Lázaro), genomics (A. M. Rocha), advanced light microscopy (P. Sampaio) and biosciences screening (A. Maia). The Genomics platform is part of the GenomePT project (POCI-01-0145-FEDER-022184). Bioimaging, ALM and BS platforms are members of the Portuguese Platform of Bioimaging (PPBI-POCI-01-0145-FEDER-022122). We thank B. Wen and E. Li (Cincinnati Children’s Hospital Medical Center) and A. Águas (ICBAS, Porto University) for valuable reagents and technical support in histology. We are grateful to all laboratory members for discussions during this work. The laboratory of E.L. was supported by: Portuguese funds through Fundação para a Ciência e a Tecnologia (FCT) in the framework of the project PTDC/MED-OUT/2747/2020; a grant from the Progeria Research Foundation (PRF 2020-78); FEDER (Fundo Europeu de Desenvolvimento Regional) funds through the COMPETE 2020–Operational Programme for Competitiveness and Internationalization (POCI), Portugal 2020 and by Portuguese funds through FCT, I. P., in the framework of the project POCI-01-0145-FEDER-031120 (PTDC/BIA-CEL/31120/2017); and by POCI-01-0145-FEDER-007274 i3S framework project co-funded by COMPETE 2020/PORTUGAL 2020 through FEDER. V.N.G. was supported by grants from the National Institutes of Health. R.R. was supported by an FCT fellowship (PD/BD/128000/2016). E.L. was supported by grants IF/00916/2014 and CEECIND/00654/2020. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: E.L.; methodology: all authors; formal analysis: R.R., R.N.G., V.U., A.V.S., A.T., J.P.C., F.V.-N., V.V.K., and E.L.; investigation: all authors; resources: A.T., D.S.N., V.N.G., T.V.K., V.V.K., and E.L.; data curation: R.R.; writing–original draft: R.R. and E.L.; writing–review and editing: all authors; supervision: E.L.; project administration: E.L.; funding acquisition: E.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Jan Deursen, David Sinclair and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Improvement of proliferation fitness of progeroid fibroblasts following FoxM1-dNdK short-term induction (related to Fig. 1).
(a) RT-qPCR analysis of endogenous FoxM1 transcript levels (2-ΔΔCt) following 2- and 4-days induction with doxycycline (dox; 2d, 4d). (b) RT-qPCR analysis of FOXM1-dNdK mRNA levels (2-ΔCt) in LAKI-FoxM1 fibroblasts. (c-f) RT-qPCR analysis of relative CcnB1 (c), Plk1 (d), p21/Cdkn1a (e) and p16/Cdkn2a (f) mRNA levels (2-ΔΔCt). (g) Western blot analysis of progerin/LMNA/C protein levels. Vinculin was used as loading control. (h) Quantification of progerin levels from western blot analysis. (i) Percentage of cells staining positive for the Annexin V-FITC apoptosis marker. (j) Western blot analysis of total FOXM1 protein levels in LAKI MAFs treated with dox for 2 and 4 consecutive days. (k) Quantification of total FOXM1 protein levels. Tubulin was used as loading control and levels were normalized to untreated cells (0d). (l-q) Immunofluorescence analysis of cellular aging phenotypes in LAKI MAFs following dox-treatment. (l) Percentage of proliferative cells (Ki67 + ). (m) Percentage of cells with DNA damage (γH2AX + ). (n) Percentage of cells staining positive for SA-β-gal activity assay. (o) Quantification of the fluorescence intensity levels of H3K9me3 and H4K20me3 epigenetic marks. (p) Percentage of cells with nuclear blebbing. (q) Quantification of the nuclear area. Error bars represent s.d. “n” in all graphs refer to the number of independent experiments, except for (o, q) where n = number of cells. Tbp and Gapdh were used as reference genes and expression levels were normalized to untreated LAKI-FoxM1 fibroblasts (0d). Statistics were performed by ordinary one-way ANOVA with Tukey’s multiple comparison correction (a-f, h, i, k), two-sided Fisher’s exact (l-n, p) and two-sided Kruskal-Wallis with Dunn’s multiple comparison correction (o, q) statistical tests for the indicated comparisons.
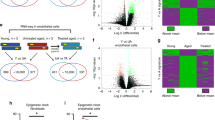
Extended Data Fig. 2 Improvement of aging molecular signature in progeroid fibroblasts under FOXM1-dNdK cyclic induction (related to Fig. 2).
(a) RT-qPCR analysis of endogenous FoxM1 transcript levels (2-ΔΔCt) in LAKI-FoxM1 MAFs under cyclic dox treatment as indicated. (b) RT-qPCR analysis of absolute FOXM1-dNdK transcript levels (2-ΔCt). (c-e) RT-qPCR analysis of Ccnb1 (c), Plk1 (d) and p16/Cdkn2a (e) transcript levels (2-ΔΔCt). (f) Western blot analysis of total FOXM1 protein levels following short-term induction of FOXM1-dNdK expression under serum starvation. (g) Quantification of total FOXM1 protein levels from western blot analysis. Tubulin was used as loading control. (h,i) Quantification of Ki67+ proliferating cells (h) and cells staining positive for SA-β-gal activity assay (i). Scale bar: 10 µm. Error bars represent s.d. “n” in all graphs refer to the number of independent experiments. Tbp and Gapdh were used as reference genes and expression levels were normalized to untreated LAKI-FoxM1 fibroblasts (0d). Statistics were performed by ordinary one-way ANOVA with Tukey’s multiple comparison correction (a-e, g) and two-sided Fisher’s exact (h, i) statistical tests for the indicated comparisons.
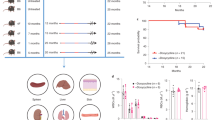
Extended Data Fig. 3 Lifespan extension and improvement of cardiovascular dysfunction in HGPS mice upon FOXM1 transgene induction (related to Fig. 3).
(a) Survival curve of LAKI C57Bl6 parental strain (ref20.) and LAKI C57Bl6xFVB strain (this study). (b,c) Survival curves of LAKI and LAKI-FoxM1 males (b) and females (c) under no-dox diet. (d,e) Survival curves of LAKI and LAKI-FoxM1 males (d) and females (e) under cyclic dox-diet. (f) Mean lifespan (weeks) of the littermates used for survival curve analysis. (g-i) Electrophysiological analysis of ejection fraction (EF) (g), stroke volume (SV) (h) and ratio between left ventricular early (E) to late (A) filling velocities (E/A) (i). (j) Representative images of Masson’s trichrome staining and (k) Sirius Red/Fast Green staining of heart sections of WT, LAKI, and LAKI-FoxM1 littermates. (l) Quantification of interstitial fibrosis. Scale bars: 1 mm (j) and 10 μm (k). Error bars represent s.d. “n” represents individual mice throughout. Statistics were performed by log-rank (Mantel-Cox) (a-e), unpaired two-sided t-test (f), two-sided Kruskal-Wallis with Dunn’s multiple comparison correction (g, i, l) and ordinary one-way ANOVA with Tukey’s multiple comparison correction (h) statistical tests for the indicated comparisons.
Extended Data Fig. 4 Improvement of HGPS histopathological features and associated molecular signatures upon FOXM1-dNdK cyclic induction (related to Fig. 4).
(a) Quantification of the epidermis layer thickness of telogenic skin. (b) Histological analysis of anagenic skin of 16 weeks-old WT, LAKI, and LAKI-FoxM1 littermates as indicated. Dashed lines delimitate the hypodermal layer. (c,d) Quantification of hypodermis (c) and epidermis (d) layers’ thickness in anagenic skin. (e) RT-qPCR analysis of FOXM1-dNdK expression in the skin (2-ΔCt). (f) RT-qPCR analysis of FoxM1 expression in the skin (2-ΔΔCt). (g-i) Western blot analysis of total FOXM1 (g) and p21 (h,i) protein levels in skin extracts. GAPDH and Vinculin were used as loading controls. (j,k) Immunostaining and quantification of cytokeratin-5 (K5)-expressing keratinocytes in skin. (l) RT-qPCR analysis of FOXM1-dNdK expression in gonadal fat (2-ΔCt). (m) RT-qPCR analysis of FoxM1 expression in gonadal fat (2-ΔΔCt). (n,o) Masson’s Trichrome of aorta sections of the indicated mice and respective quantification. (p) RT-qPCR analysis of FOXM1-dNdK expression in aorta (2-ΔCt). (q) RT-qPCR analysis of FoxM1 expression in the aorta (2-ΔΔCt). (r) 3D modeling of the cortical tibia (diaphysis). (s) Quantification of micro-CT analysis of cortical bone thickness. (t) Cytokine levels in serum samples. (u) RT-qPCR analysis of FOXM1-dNdK expression in gonadal fat (2-ΔCt). (v) RT-qPCR analysis of FoxM1 expression in gonadal fat (2-ΔΔCt). (x) RT-qPCR analysis of senescence markers in the kidney. Scale bars: 50 μm (b, n), 25 μm (j) and 5 mm (r). Tbp and Gapdh were used as reference genes and expression levels were normalized to WT mice (2-ΔΔCt). Error bars represent s.d. “n” represents independent experiments, except for (a, c, d, k, o, s) where n = individual mice. Statistics were performed by two-sided Kruskal-Wallis test with Dunn’s multiple comparison correction (a, d, o) and ordinary one-way ANOVA with Tukey’s multiple comparison correction (c, e, f, i, k-m, p, q, s-x) statistical tests for the indicated comparisons.
Extended Data Fig. 5 FoxM1 downregulation in the gut, heart, and brain tissues of naturally aged mice (related to Fig. 6).
(a-c) Gene expression analysis of endogenous FoxM1 transcript levels (2-ΔΔCt) in 4-week-old vs. 2-year-old WT mice. Tbp and Gapdh were used as reference genes and expression levels were normalized to 4w-old WT mice. Error bars represent s.d. “n” represents individual mice. Statistics were performed by unpaired two-sided t-test.
Extended Data Fig. 6 Lifespan extension upon widespread expression of FOXM1-dNdK transgene (related to Fig. 6).
(a,b) Survival curves of Rosa26 and FoxM1-dNdK males (a) and females (b) under no-dox diet. (c,d) Survival curves of Rosa26 and FoxM1-dNdK males (c) and females (d) under cyclic dox-diet. (e) Mean lifespan (weeks) of the littermates used for survival curve analysis. (f) RT-qPCR analysis of endogenous FoxM1 expression in tissues of 88-week-old (±1.7 y) FoxM1-dNdK (2-ΔΔCt). Tbp and Gapdh were used as reference genes and expression levels were normalized to those in Rosa26 mice tissues. (g) Western blot analysis of FOXM1-dNdK protein levels in extracts of skin, kidney, and liver of 88-week-old mice under dox-diet cyclic induction. Vinculin was used as loading control. (h) Immunostaining of FOXM1-dNdK on skin, kidney and liver sections of the mentioned mice. Scale bars: 100 μm (kidney insets, 50 μm). Error bars represent s.d. “n” represents individual mice throughout, except for (f) where n = independent experiments. Statistics were performed by log-rank (Mantel-Cox) (a-d) and unpaired two-sided t-test (e, f) statistical tests.
Extended Data Fig. 7 Tumor incidence analysis on cohorts under long-term FOXM1 transgene induction (related to Fig. 6).
(a) Representative photographs of 88-week-old Rosa26 and FoxM1-dNdK littermates’ necropsy. (b) Post-mortem solid tumor incidence on the indicated cohorts. “n” represents individual mice. (c) Summarization of spontaneous solid tumor formation observed in cohorts indicated in (b). (d) Representative images of liver and kidney histological sections for exclusion of lymphocyte infiltration in n = 3 animals per genotype. Scale bars: 1 cm (a) and 200 μm (d). Error bars represent s.d. Statistics were performed by two-sided Fisher’s exact statistical test for the indicated comparisons.
Extended Data Fig. 8 Improvement of senescence-associated histopathological aging features upon long-term FOXM1-dNdK induction (related to Fig. 6).
(a,b) Immunostaining and quantification of PAX7-positive muscle stem cells (arrowheads) in sections of the gastrocnemius muscle of Rosa26 and FoxM1-dNdK mice. (c) Grip-strength test in 14-16w-old Rosa26 and in 88w-old Rosa26 and FoxM1-dNdK littermates (n = 3 animals per genotype). Dashed lines interconnect values of 3 measurements in the same animal obtained with 5 min interval. (d,e) Masson’s Trichrome staining of aorta sections of Rosa26 and FoxM1-dNdK littermates and respective quantification. (f-h) Exclusion of sex bias effect in improved phenotypes upon FOXM1 transgene induction. (i-l) RT-qPCR analysis of senescence markers in the skin (i), kidney (j), gonadal fat (k), and gastrocnemius muscle (l) of 88-week-old Rosa26 and FoxM1-dNdK littermates. Tbp and Gapdh were used as reference genes and expression levels were normalized to Rosa26 mice (2-ΔΔCt). Scale bars: 50 μm (a, d) and 2.5 μm (inset in a). Error bars represent s.d. “n” is the number of mice except in (i-l) where n = independent experiments. Statistics were performed by two-sided Fisher’s exact (b, e), unpaired two-sided t-test (c, i-l), and ordinary one-way ANOVA with Tukey’s multiple comparison correction (f-h) statistical tests for the indicated comparisons.
Extended Data Fig. 9 Impact of FOXM1 transgene induction in tissues from young animals (related to Fig. 6).
(a) Histological analysis of the aorta in littermate mice as indicated and respective quantification (b) of vascular smooth muscle cells (VSMC) nuclei density and (c) of aortic wall thickness. (d) Histological analysis of telogenic skin. Dashed lines delimitate the hypodermal layer. (e,f) Quantification of hypodermal and epidermal layers’ thickness in telogenic skin. (g) Histological analysis of gonadal fat tissue and (h) quantification of adipocyte cross-sectional area and (i) crown-like structures. (j) Histological analysis of gastrocnemius and (k) quantification of muscle fiber cross-sectional area. Scale bar: 10 μm (a) and 50 μm (d, g, j). Error bars represent s.d. “n” represents the number of individual mice. Statistics were performed by unpaired two-sided t-test.
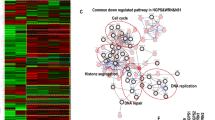
Extended Data Fig. 10 Bioinformatic analysis of transcriptomic changes in the aorta and gastrocnemius produced by FOXM1-dNdK induction (related to Fig. 6).
(a) Principal component analysis (PCA) of aorta (circles) and muscle (triangles) samples from 88-week-old Rosa26 (black) and FoxM1-dNdK (green) mice under cyclic dox-diet. (b,c) MSigDB pathways found similarly (b) and oppositely (c) altered in the aorta and muscle. (d) KEGG pathway analysis for the aorta signature. (e) GSEA-analysis in the gastrocnemius using pathways from MSigDB databases. (f) KEGG pathway analysis for the gastrocnemius signature. (g) GSEA enrichment plot for ‘SASP’ custom pathway in the FoxM1-dNdK gastrocnemius. (h) Association of transcriptomic changes induced by FOXM1-dNdK in the muscle with signatures of lifespan-extending interventions (LEI) (GH - growth hormone; Common LEI - common gene expression changes across different interventions) and aging (muscle and murine multi-tissue). (i) Correlation matrix of aggregated gene expression profiles across interventions in comparison with signatures from the aorta and gastrocnemius muscle under long-term induction of FOXM1-dNdK. (j) Pathways altered in lifespan-extending interventions, aging signatures and FOXM1-dNdK overexpressing tissues. Significance scores of functional enrichment analyses (d-h) are described in Methods.
Supplementary information
Supplementary Information
Supplementary Table 1 and Supplementary Notes
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Ribeiro, R., Macedo, J.C., Costa, M. et al. In vivo cyclic induction of the FOXM1 transcription factor delays natural and progeroid aging phenotypes and extends healthspan. Nat Aging 2, 397–411 (2022). https://doi.org/10.1038/s43587-022-00209-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-022-00209-9
This article is cited by
-
Hesperetin activates CISD2 to attenuate senescence in human keratinocytes from an older person and rejuvenates naturally aged skin in mice
Journal of Biomedical Science (2024)
-
FOXO1-mediated lipid metabolism maintains mammalian embryos in dormancy
Nature Cell Biology (2024)
-
DNA methylation abnormalities induced by advanced maternal age in villi prime a high-risk state for spontaneous abortion
Clinical Epigenetics (2023)
-
Metabolomics analysis of dietary restriction results in a longer lifespan due to alters of amino acid levels in larval hemolymph of Bombyx mori
Scientific Reports (2023)
-
Cellular rejuvenation: molecular mechanisms and potential therapeutic interventions for diseases
Signal Transduction and Targeted Therapy (2023)