Abstract
Understanding the physiological origins of age-related cognitive decline is of critical importance given the rising age of the world’s population1. Previous work in animal models has established a strong link between cognitive performance and the microbiota2,3,4,5, and it is known that the microbiome undergoes profound remodeling in older adults6. Despite growing evidence for the association between age-related cognitive decline and changes in the gut microbiome, the mechanisms underlying such interactions between the brain and the gut are poorly understood. Here, using fecal microbiota transplantation (FMT), we demonstrate that age-related remodeling of the gut microbiota leads to decline in cognitive function in mice and that this impairment can be rescued by transplantation of microbiota from young animals. Moreover, using a metabolomic approach, we found elevated concentrations of δ-valerobetaine, a gut microbiota-derived metabolite, in the blood and brain of aged mice and older adults. We then demonstrated that δ-valerobetaine is deleterious to learning and memory processes in mice. At the neuronal level, we showed that δ-valerobetaine modulates inhibitory synaptic transmission and neuronal network activity. Finally, we identified specific bacterial taxa that significantly correlate with δ-valerobetaine levels in the brain. Based on our findings, we propose that δ-valerobetaine contributes to microbiota-driven brain aging and that the associated mechanisms represent a promising target for countering age-related cognitive decline.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Microbiome profiling (16S rRNA-seq) raw data are accessible at the European Nucleotide Archive under accession number PRJEB46041. Data on TwinsUK twin participants are available to bona fide researchers under managed access due to governance and ethical constraints. Raw data should be requested via the website (http://twinsuk.ac.uk/resources-for-researchers/access-our-data/), and requests are reviewed by the TwinsUK Resource Executive Committee regularly. Other data that support the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
R scripts used for microbiome profiling are available online at https://github.com/OmarMossad/Mossad_FMT_NatAging.
References
Livingston, G. et al. Dementia prevention, intervention, and care. Lancet 390, 2673–2734 (2017).
Desbonnet, L., Clarke, G., Shanahan, F., Dinan, T. G. & Cryan, J. F. Microbiota is essential for social development in the mouse. Mol. Psychiatry 19, 146–148 (2014).
Hoban, A. E. et al. The microbiome regulates amygdala-dependent fear recall. Mol. Psychiatry 23, 1134–1144 (2018).
Savignac, H. M., Tramullas, M., Kiely, B., Dinan, T. G. & Cryan, J. F. Bifidobacteria modulate cognitive processes in an anxious mouse strain. Behav. Brain Res. 287, 59–72 (2015).
Magnusson, K. R. et al. Relationships between diet-related changes in the gut microbiome and cognitive flexibility. Neuroscience 300, 128–140 (2015).
Claesson, M. J. et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl Acad. Sci. USA 108, 4586–4591 (2011).
Koistinen, V. M. et al. Contribution of gut microbiota to metabolism of dietary glycine betaine in mice and in vitro colonic fermentation. Microbiome 7, 103 (2019).
Koeth, R. A. et al. γ-butyrobetaine is a proatherogenic intermediate in gut microbial metabolism of l-carnitine to TMAO. Cell Metab. 20, 799–812 (2014).
Shin, S. Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46, 543–550 (2014).
Long, T. et al. Whole-genome sequencing identifies common-to-rare variants associated with human blood metabolites. Nat. Genet. 49, 568–578 (2017).
Vaadia, E. et al. Dynamics of neuronal interactions in monkey cortex in relation to behavioural events. Nature 373, 515–518 (1995).
Courtin, J. et al. Prefrontal parvalbumin interneurons shape neuronal activity to drive fear expression. Nature 505, 92–96 (2014).
Prezioso, M. et al. Spike-timing-dependent plasticity learning of coincidence detection with passively integrated memristive circuits. Nat. Commun. 9, 5311 (2018).
Dejean, C. et al. Prefrontal neuronal assemblies temporally control fear behaviour. Nature 535, 420–424 (2016).
Yang, Y. & Mailman, R. B. Strategic neuronal encoding in medial prefrontal cortex of spatial working memory in the T-maze. Behav. Brain Res. 343, 50–60 (2018).
Morici, J. F., Bekinschtein, P. & Weisstaub, N. V. Medial prefrontal cortex role in recognition memory in rodents. Behav. Brain Res. 292, 241–251 (2015).
Kells, P. A., Gautam, S. H., Fakhraei, L., Li, J. & Shew, W. L. Strong neuron-to-body coupling implies weak neuron-to-neuron coupling in motor cortex. Nat. Commun. 10, 1575 (2019).
Okun, M. et al. Diverse coupling of neurons to populations in sensory cortex. Nature 521, 511–515 (2015).
Sweeney, Y. & Clopath, C. Population coupling predicts the plasticity of stimulus responses in cortical circuits. eLife 9, e56053 (2020).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Blank, T., Goldmann, T. & Prinz, M. Microglia fuel the learning brain. Trends Immunol. 35, 139–140 (2014).
Parkhurst, C. N. et al. Microglia promote learning-dependent synapse formation through brain-derived neurotrophic factor. Cell 155, 1596–1609 (2013).
Wu, Y., Dissing-Olesen, L., MacVicar, B. A. & Stevens, B. Microglia: dynamic mediators of synapse development and plasticity. Trends Immunol. 36, 605–613 (2015).
Boehme, M. et al. Microbiota from young mice counteracts selective age-associated behavioral deficits. Nat Aging 1, 666–676 (2021).
Dagher, N. N. et al. Colony-stimulating factor 1 receptor inhibition prevents microglial plaque association and improves cognition in 3×Tg-AD mice. J. Neuroinflammation 12, 139 (2015).
Lawton, M. P. et al. Health, valuation of life, and the wish to live. Gerontologist 39, 406–416 (1999).
Gazzaley, A., Cooney, J. W., Rissman, J. & D’Esposito, M. Top–down suppression deficit underlies working memory impairment in normal aging. Nat. Neurosci. 8, 1298–1300 (2005).
Grady, C. L. et al. Age-related reductions in human recognition memory due to impaired encoding. Science 269, 218–221 (1995).
Jermakowicz, W. J. & Casagrande, V. A. Neural networks a century after Cajal. Brain Res. Rev. 55, 264–284 (2007).
Cohen, M. R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Averbeck, B. B., Latham, P. E. & Pouget, A. Neural correlations, population coding and computation. Nat. Rev. Neurosci. 7, 358–366 (2006).
Rolls, E. T., Treves, A. & Tovee, M. J. The representational capacity of the distributed encoding of information provided by populations of neurons in primate temporal visual cortex. Exp. Brain Res. 114, 149–162 (1997).
Shew, W. L., Yang, H., Yu, S., Roy, R. & Plenz, D. Information capacity and transmission are maximized in balanced cortical networks with neuronal avalanches. J. Neurosci. 31, 55–63 (2011).
Goossens, T., Vercammen, C., Wouters, J. & van Wieringen, A. Aging affects neural synchronization to speech-related acoustic modulations. Front. Aging Neurosci. 8, 133 (2016).
Biagi, E. et al. Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 5, e10667 (2010).
Langille, M. G. et al. Microbial shifts in the aging mouse gut. Microbiome 2, 50 (2014).
Rath, S., Rud, T., Pieper, D. H. & Vital, M. Potential TMA-producing bacteria are ubiquitously found in Mammalia. Front. Microbiol. 10, 2966 (2019).
Karkkainen, O. et al. Whole grain intake associated molecule 5-aminovaleric acid betaine decreases β-oxidation of fatty acids in mouse cardiomyocytes. Sci. Rep. 8, 13036 (2018).
Mezo, C. et al. Different effects of constitutive and induced microbiota modulation on microglia in a mouse model of Alzheimer’s disease. Acta Neuropathol. Commun. 8, 119 (2020).
Spowart-Manning, L. & van der Staay, F. J. The T-maze continuous alternation task for assessing the effects of putative cognition enhancers in the mouse. Behav. Brain Res. 151, 37–46 (2004).
Evans, A. M., DeHaven, C. D., Barrett, T., Mitchell, M. & Milgram, E. Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal. Chem. 81, 6656–6667 (2009).
Rath, S., Heidrich, B., Pieper, D. H. & Vital, M. Uncovering the trimethylamine-producing bacteria of the human gut microbiota. Microbiome 5, 54 (2017).
Takahashi, S., Tomita, J., Nishioka, K., Hisada, T. & Nishijima, M. Development of a prokaryotic universal primer for simultaneous analysis of bacteria and archaea using next-generation sequencing. PLoS ONE 9, e105592 (2014).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Foster, Z. S. L., Sharpton, T. J. & Grünwald, N. J. Metacoder: an R package for visualization and manipulation of community taxonomic diversity data. PLoS Comput. Biol. 13, e1005404 (2017).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
T.B., O.M., D.E., T.L., E.N. and M.P. were supported by the DFG (SFB/TRR167 ‘NeuroMac’). M.P. was further supported by the Sobek Foundation, the Ernst-Jung Foundation, the DFG (SFB 992, SFB1160, Reinhart Koselleck grant, Gottfried Wilhelm Leibniz prize), the Alzheimer Forschung Initiative e.V. (AFI) and the Ministry of Science, Research and Arts, Baden-Wuerttemberg (Sonderlinie ‘Neuroinflammation’). This study was supported by the DFG under Germany’s Excellence Strategy (CIBSS, EXC-2189, project ID-390939984). We are grateful to M. Gomez De Agüero and S.C. Ganal-Vonarburg (Macpherson laboratory, Bern, Switzerland) for providing us with GF mice. TwinsUK is funded by the Wellcome Trust, the Medical Research Council, the European Union, the National Institute for Health Research-funded BioResource, the Clinical Research Facility and the Biomedical Research Centre based at Guy’s and St. Thomas’ NHS Foundation Trust in partnership with King’s College London. We are grateful to Plexxikon for providing the CSF1R inhibitor PLX5622. We thank A. Dumas for proofreading the article.
Author information
Authors and Affiliations
Contributions
O.M. conducted the majority of experiments and data analysis; E.N., T.L. and J.M.B. performed targeted metabolomics; S.W. and M.V. performed microbiome profiling; S.F. and J.F.S. performed electrophysiological recordings; D.S. and P.S. contributed to the FMT experimental paradigm; J.F.S., D.E., T.L., B.S., A.S., M.B. and M.P. designed experiments and interpreted results; T.B. initiated and conceived the project and designed experiments; O.M. and T.B. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
T.B. is an author on a patent application, which is pending. All authors declare no other competing interests.
Additional information
Peer review information Nature Aging thanks Sven Pettersson, Pieter Dorrestein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effects of antibiotics and fecal microbiota transplantation.
(a) Gating on DAPI- Syto9+ live bacteria of fecal samples from SPF and ABX-treated mice. Representative plot for the flow cytometry gating. (b) Percentage of live Gram+ and Gram- bacteria and (c) Quantification of live bacteria per mg fecal sample. (d) NMDS plot of Beta diversity (Bray-Curtis distance matrix) of microbiota in fecal pellets. Samples collected from start and end of the experiments are shown as triangles and circles, respectively. (e) Abundance of cecal microbiota composition profiles at the phylum level (each color represents one bacterial phylum). (f, g) Alpha diversity in cecum measured by (f) Shannon and (g) Simpson indices. Box plots; centre = median, upper and lower “hinges” correspond to the first and third quartiles (the 25th and 75th percentiles), upper whisker extends from the hinge to the highest value that is within 1.5 * IQR of the hinge, and lower whisker extends from the hinge to the lowest value within 1.5 * IQR of the hinge, where IQR is the inter-quartile range, or distance between the first and third quartiles. Statistical analysis with (b, c) Mann–Whitney U test (two-tailed) (f, g) Wilcoxon test followed by Holm post-hoc test (two-tailed) (*p < 0.05, **p < 0.01, ***p < 0.001, ns = not significant). Exact p-values are reported in the source data.
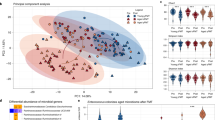
Extended Data Fig. 2 Age-associated microbiota alterations on genus level.
(a,b) Differentially abundant taxa in (a) aged vs young animals, and (b) oFMTiAged vs yFMTiAged animals with Padj<0.05. Each dot represents one ASV. Genus classification is depicted on y-axis, and log2FoldChange on x-axis. Taxa with unclassified genus were filtered out. Color represents different phyla.
Extended Data Fig. 3 δ-Valerobetaine enhances inhibitory postsynaptic currents and population synchronization in the medial prefrontal cortex.
(a) Dose-dependent increase in δ-valerobetainein the brain represented by area under the curve (AUC) of the mass-spec peaks.Error bars represent SEM. (b) T-maze (c,d) Familiarization (c) and Testing sessions (d) with a 6 h gap for the Novel object recognition test (NOR) (n=5). Statistical analysis with one-way ANOVA compared to the control group (0 mg/kg) followed by Bonferroni post-hoctest (*p < 0.05, **p < 0.01, ***p < 0.001). (e) Examples of spontaneous excitatory postsynaptic currents (EPSCs, left) and spontaneous inhibitory postsynaptic currents (IPSCs, right) of prefrontal pyramidal cells under control conditions (black) and after pre-incubation with 10 µM δ-valerobetaine (green). Data arefrom patch-clamp recordings in acute brain slices. (f) δ-valerobetaine increased the frequency (p=0.006, unpairedt-test) but not the amplitude of spontaneous IPSCs (p=0.174, unpaired t-test) while EPSC properties were unaltered (p=0.089 and p=0.641 for frequency andamplitude, respectively). n=10 control and 9 δ-valerobetaine-treated neurons. (a-d,f) Error bars represent SEM (g) Single unit recordings from them PFC to assess the effect of δ-valerobetaine on population activity in vivo. Example traces show a shortsegment of the summed population firing rate from all units of one recordingsession (black, binned in 100 ms time epochs, see Methods) and the binned spikerate of one neuron (bottom) before (left) and after (right) δ-valerobetaineinjection. Note that the unit showed increased coupling to the populationactivity upon δ-valerobetaine injection, indicating enhanced synchronization inthe prefrontal network. (h) Quantification of spike rates and population coupling of all neurons revealedno effect of δ-valerobetaine on average spike frequency but a significant increase in population coupling of the recorded cells (control: n=77 units from 4 mice, δ-valerobetaine: n=56 units from 4, Welch’s tests). Black bars indicatemeans. Statistical analysis (f,h) with Mann–Whitney U test (two-tailed)(**p < 0.01, ns = not significant).
Extended Data Fig. 4 Beneficial cognitive effect of young FMT is independent of microglia.
(a) Schematic diagram of the experimental setting: as in (Fig.1a) with the addition of the mice receiving either control AIN-76A standard chow (NC) or supplemented with PLX5622 (PLX, 1200 mg/kg) to deplete CSF1R-dependent cells, including microglia. (b) Representative immunofluorescence images of Iba-1+ (red), DAPI (blue) in cortex and hippocampus of PLX5622-treated mice upon sacrifice after behavioral testing. Scale bar, 150μm. (c) Quantification of Iba1+ cell density in cortex and hippocampus as one ROI upon PLX5622 treatment. Error bars represent SEM. (d)T-maze (e,f) Familiarization (e) and Testing sessions (f) with a 6 h gap for Novel object recognition test (NOR) (young (NC), aged (NC), aged (PLX), yFMTiAged (NC), yFMTiAged (PLX), oFMTiAged; n=4, 7, 8, 7, 8, 7, respectively). (a-f) Data represent two independent experiments. Errorbars represent SEM. Statistical analysis with (c-f) one-way ANOVA followed by Tukey post-hoc test (*p < 0.05, **p < 0.01, ***p < 0.001, ns = not significant). Exact p-values are reported in the source data.
Extended Data Fig. 5 Graphical abstract.
Heterochronic microbiota transplantation ameliorates the age-related cognitive decline through modulating the levels of the microbiota-dependent metabolite, δ-valerobetaine.
Supplementary information
Supplementary Tables
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Mossad, O., Nent, E., Woltemate, S. et al. Microbiota-dependent increase in δ-valerobetaine alters neuronal function and is responsible for age-related cognitive decline. Nat Aging 1, 1127–1136 (2021). https://doi.org/10.1038/s43587-021-00141-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00141-4
This article is cited by
-
Body and mind: how somatic feedback signals shape brain activity and cognition
Pflügers Archiv - European Journal of Physiology (2023)
-
Inconsistent nomenclature of microbiota-associated metabolites hampers progress of the field
Nature Metabolism (2022)
-
Healthy aging and the human gut microbiome: why we cannot just turn back the clock
Nature Aging (2022)
-
Gut microbiota drives age-related oxidative stress and mitochondrial damage in microglia via the metabolite N6-carboxymethyllysine
Nature Neuroscience (2022)