Abstract
The evident genetic, pathological and clinical heterogeneity of Alzheimer’s disease (AD) poses challenges for traditional drug development. We conducted a computational drug-repurposing screen for drugs to treat apolipoprotein E4 (APOE4)-related AD. We first established APOE genotype-dependent transcriptomic signatures of AD by analyzing publicly available human brain databases. We then queried these signatures against the Connectivity Map database, which contains transcriptomic perturbations of more than 1,300 drugs, to identify those that best reverse APOE genotype-specific AD signatures. Bumetanide was identified as a top drug for APOE4-related AD. Treatment of APOE4-knock-in mice without or with amyloid β (Aβ) accumulation using bumetanide rescued electrophysiological, pathological or cognitive deficits. Single-nucleus RNA sequencing revealed transcriptomic reversal of AD signatures in specific cell types in these mice, a finding confirmed in APOE4 induced pluripotent stem cell (iPSC)-derived neurons. In humans, bumetanide exposure was associated with a significantly lower AD prevalence in individuals over the age of 65 years in two electronic health record databases, suggesting the effectiveness of bumetanide in preventing AD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (or in its Supplementary Information) or deposited in the GEO and are also available from the corresponding authors’ laboratories. Publicly available datasets used are available in the GEO under the accession number GSE15222 with associated covariate data found on the Myers laboratory website (http://labs.med.miami.edu/myers/LFuN/LFUN/DATA.html) and the associated Google Drive (https://drive.google.com/drive/folders/1ud5F9WN9Xx3oXIkb5xIg1b_zz1nzp3IR) in the ‘samples.covar.ZIP’ file. The CMap database is available in Sage Synapse in the HCC_NEN project from the Bin Chen laboratory under the accession number syn6187678 (https://www.synapse.org/#!Synapse:syn6173892/files), and is linked to the Bin Chen laboratory GitLab repository (also see Code availability below). The publicly available RNA-seq dataset of brains of aging APOE4-KI mice was from Zhao et al. (https://doi.org/10.7303/syn20808171)28. Figs. 4 and 6 and Extended Data Figs. 3, 4, 7 and 8 have associated mouse snRNA-seq data or iPSC-derived human neuron bulk RNA-seq data generated in this study, which are available in the GEO under the accession number GSE182765. The UCSF EHR database and the Mt. Sinai EHR database are not yet available to the general public.
Code availability
The drug-repurposing algorithm can be found in the Bin Chen laboratory GitLab repository (https://github.com/Bin-Chen-Lab/HCC_NEN/). The following packages or software were used either as dependencies to downloading or using packages mentioned in the Methods or in creating the figures in this study: clusterProfiler_3.10.1, pheatmap_1.0.12, vsn_3.48.1, SummarizedExperiment_1.10.1, DelayedArray_0.6.6, BiocParallel_1.14.2, matrixStats_0.54.0, GenomicRanges_1.32.7, GenomeInfoDb_1.16.0, edgeR_3.22.5, mice_3.4.0, lattice_0.20-35, ggbiplot_0.55, scales_1.0.0, plyr_1.8.4, eulerr_5.0.0, VennDiagram_1.6.20, futile.logger_1.4.3, data.table_1.11.8, gridExtra_2.3, GEOquery_2.48.0, qvalue_2.12.0, illuminaHumanv1.db_1.26.0, org.Hs.eg.db_3.6.0, AnnotationDbi_1.42.1, IRanges_2.14.12, S4Vectors_0.18.3, Biobase_2.40.0, BiocGenerics_0.26.0, nf-core/rnaseq_1.4.2, org.Mm.eg.db_3.6.0, raster_2.8.19, FastQC_0.11.8, featureCounts_1.6.2, Data.table_1.12.8, RMySQL_0.10.17, Dplyr_0.8.3, magrittr_1.5, tidyverse_1.3.1 and biomaRt_2_38_0. All custom codes generated in this study will be made available from the corresponding authors’ laboratories upon request.
Change history
12 November 2021
A Correction to this paper has been published: https://doi.org/10.1038/s43587-021-00144-1
References
Golde, T. E., Schneider, L. S. & Koo, E. H. Anti-Aβ therapeutics in Alzheimer’s disease: the need for a paradigm shift. Neuron 69, 203–213 (2011).
Huang, Y. & Mucke, L. Alzheimer mechanisms and therapeutic strategies. Cell 148, 1204–1222 (2012).
Liu, C. C., Kanekiyo, T., Xu, H. & Bu, G. Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat. Rev. Neurol. 9, 106–118 (2013).
Verghese, P. B., Castellano, J. M. & Holtzman, D. M. Apolipoprotein E in Alzheimer’s disease and other neurological disorders. Lancet Neurol. 10, 241–252 (2011).
Mahley, R. W. & Huang, Y. Apolipoprotein E sets the stage: response to injury triggers neuropathology. Neuron 76, 871–885 (2012).
Corder, E. H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261, 921–923 (1993).
Barnes, E. E. & Yaffe, K. Vitamin E and donepezil for the treatment of mild cognitive impairment. N. Engl. J. Med. 353, 951–952 (2005).
Marchant, N. L., King, S. L., Tabet, N. & Rusted, J. M. Positive effects of cholinergic stimulation favor young APOE ε4 carriers. Neuropsychopharmacology 35, 1090–1096 (2010).
Salloway, S. et al. Two phase 3 trials of bapineuzumab in mild-to-moderate Alzheimer’s disease. N. Engl. J. Med. 370, 322–333 (2014).
Cheng, F. et al. Prediction of drug–target interactions and drug repositioning via network-based inference. PLoS Comput. Biol. 8, e1002503 (2012).
Csermely, P., Korcsmaros, T., Kiss, H. J., London, G. & Nussinov, R. Structure and dynamics of molecular network: a novel paradigm of drug discovery: a comprehensive review. Pharmacol. Ther. 138, 333–408 (2013).
Sirota, M. et al. Discovery and preclinical validation of drug indications using compendia of public gene expression data. Sci. Transl. Med. 3, 96ra77 (2011).
Chen, B. et al. Computational discovery of niclosamide ethanolamine, a repurposed drug candidate that reduces growth of hepatocellular carcinoma cells in vitro and in mice by inhibiting cell division cycle 37 signaling. Gastroenterology 152, 2022–2036 (2017).
Chen, B. & Butte, A. J. Leveraging big data to transform target selection and drug discovery. Clin. Pharmacol. Ther. 99, 285–297 (2016).
Cai, X., Chen, Y., Gao, Z. & Xu, R. Explore small molecule-induced genome-wide transcriptional profiles for novel inflammatory bowel disease drug. AMIA Jt Summits Transl. Sci. Proc. 2016, 22–31 (2016).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Webster, J. A. et al. Genetic control of human brain transcript expression in Alzheimer disease. Am. J. Hum. Genet. 84, 445–458 (2009).
Farrer, L. A. et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. JAMA 278, 1349–1356 (1997).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Chen, B. et al. Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets. Nat. Commun. 8, 16022 (2017).
Gharaylou, Z. et al. A preliminary study evaluating the safety and efficacy of bumetanide, an NKCC1 inhibitor, in patients with drug-resistant epilepsy. CNS Drugs 33, 283–291 (2019).
Goubert, E. et al. Bumetanide prevents brain trauma-induced depressive-like behavior. Front. Mol. Neurosci. 12, 12 (2019).
Lemonnier, E. et al. A randomised controlled trial of bumetanide in the treatment of autism in children. Transl. Psychiatry 2, e202 (2012).
Lemonnier, E., Lazartigues, A. & Ben-Ari, Y. Treating schizophrenia with the diuretic bumetanide: a case report. Clin. Neuropharmacol. 39, 115–117 (2016).
Lemonnier, E. et al. Effects of bumetanide on neurobehavioral function in children and adolescents with autism spectrum disorders. Transl. Psychiatry 7, e1056 (2017).
Rahmanzadeh, R. et al. Effect of co-administration of bumetanide and phenobarbital on seizure attacks in temporal lobe epilepsy. Basic Clin. Neurosci. 9, 408–416 (2018).
Sivakumaran, S. & Maguire, J. Bumetanide reduces seizure progression and the development of pharmacoresistant status epilepticus. Epilepsia 57, 222–232 (2016).
Zhao, N. et al. Alzheimer’s risk factors age, APOE genotype, and sex drive distinct molecular pathways. Neuron 106, 727–742 (2020).
Najm, R., Jones, E. A. & Huang, Y. Apolipoprotein E4, inhibitory network dysfunction, and Alzheimer’s disease. Mol. Neurodegener. 14, 24 (2019).
Milior, G. et al. Electrophysiological properties of CA1 pyramidal neurons along the longitudinal axis of the mouse hippocampus. Sci. Rep. 6, 38242 (2016).
Bliss, T. V. & Collingridge, G. L. A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 (1993).
Selkoe, D. J. Alzheimer’s disease is a synaptic failure. Science 298, 789–791 (2002).
Andrews-Zwilling, Y. et al. Apolipoprotein E4 causes age- and tau-dependent impairment of GABAergic interneurons, leading to learning and memory deficits in mice. J. Neurosci. 30, 13707–13717 (2010).
Leung, L. et al. Apolipoprotein E4 causes age- and sex-dependent impairments of hilar GABAergic interneurons and learning and memory deficits in mice. PLoS ONE 7, e53569 (2012).
Knoferle, J. et al. Apolipoprotein E4 produced in GABAergic interneurons causes learning and memory deficits in mice. J. Neurosci. 34, 14069–14078 (2014).
Mucke, L. et al. High-level neuronal expression of Aβ1–42 in wild-type human amyloid protein precursor transgenic mice: synaptotoxicity without plaque formation. J. Neurosci. 20, 4050–4058 (2000).
Bien-Ly, N., Gillespie, A. K., Walker, D., Yoon, S. Y. & Huang, Y. Reducing human apolipoprotein E levels attenuates age-dependent Aβ accumulation in mutant human amyloid precursor protein transgenic mice. J. Neurosci. 32, 4803–4811 (2012).
Wang, C. et al. Gain of toxic apolipoprotein E4 effects in human iPSC-derived neurons is ameliorated by a small-molecule structure corrector. Nat. Med. 24, 647–657 (2018).
Lennon, M. J., Makkar, S. R., Crawford, J. D. & Sachdev, P. S. Midlife hypertension and Alzheimer’s disease: a systematic review and meta-analysis. J. Alzheimers Dis. 71, 307–316 (2019).
Ho, J., Tumkaya, T., Aryal, S., Choi, H. & Claridge-Chang, A. Moving beyond P values: data analysis with estimation graphics. Nat. Methods 16, 565–566 (2019).
Kharod, S. C., Kang, S. K. & Kadam, S. D. Off-label use of bumetanide for brain disorders: an overview. Front. Neurosci. 13, 310 (2019).
Puskarjov, M., Kahle, K. T., Ruusuvuori, E. & Kaila, K. Pharmacotherapeutic targeting of cation–chloride cotransporters in neonatal seizures. Epilepsia 55, 806–818 (2014).
Töpfer, M. et al. Consequences of inhibition of bumetanide metabolism in rodents on brain penetration and effects of bumetanide in chronic models of epilepsy. Eur. J. Neurosci. 39, 673–687 (2014).
Gharaylou, Z. et al. Longitudinal effects of bumetanide on neuro-cognitive functioning in drug-resistant epilepsy. Front. Neurol. 10, 483 (2019).
Sala Frigerio, C. et al. The major risk factors for Alzheimer’s disease: age, sex, and genes modulate the microglia response to Aβ plaques. Cell Rep. 27, 1293–1306 (2019).
Nguyen, A. T. et al. APOE and TREM2 regulate amyloid-responsive microglia in Alzheimer’s disease. Acta Neuropathol. 140, 477–493 (2020).
Griswold, A. J. et al. Increased APOE ε4 expression is associated with the difference in Alzheimer’s disease risk from diverse ancestral backgrounds. Alzheimers Dement. 17, 1179–1188 (2021).
Caselli, R. J. Obstructive sleep apnea, apolipoprotein E e4, and mild cognitive impairment. Sleep Med. 9, 816–817 (2008).
Drogos, L. et al. Evidence of association between sleep quality and APOE ε4 in healthy older adults: a pilot study. Neurology 87, 1836–1842 (2016).
Tranah, G. J. et al. APOEε4 and slow wave sleep in older adults. PLoS ONE 13, e0191281 (2018).
Gozal, D., Capdevila, O. S., Kheirandish-Gozal, L. & Crabtree, V. M. APOE ε4 allele, cognitive dysfunction, and obstructive sleep apnea in children. Neurology 69, 243–249 (2007).
Listos, J. et al. The mechanisms involved in morphine addiction: an overview. Int. J. Mol. Sci. 20, 4302 (2019).
Falcon, E. & McClung, C. A. A role for the circadian genes in drug addiction. Neuropharmacology 56, 91–96 (2009).
Hamanaka, H. et al. Altered cholesterol metabolism in human apolipoprotein E4 knock-in mice. Hum. Mol. Genet. 9, 353–361 (2000).
Sullivan, P. M., Mace, B. E., Maeda, N. & Schmechel, D. E. Marked regional differences of brain human apolipoprotein E expression in targeted replacement mice. Neuroscience 124, 725–733 (2004).
Workman, C. et al. A new non-linear normalization method for reducing variability in DNA microarray experiments. Genome Biol. 3, research0048.0041–research0048.0016 (2002).
Lui, J. H. et al. Radial glia require PDGFD–PDGFRβ signalling in human but not mouse neocortex. Nature 515, 264–268 (2014).
Raju, C. S. et al. Secretagogin is expressed by developing neocortical GABAergic neurons in humans but not mice and increases neurite arbor size and complexity. Cereb. Cortex 28, 1946–1958 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Dijk, D. V. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Acknowledgements
This work was supported by grants AG057683 (Y.H. and M.S.), AG048017 (Y.H.), 1F31AG058439 (A.T.), AG061150 (M.Y.Z.), AG059319 (B.S.G. and M.B.), 1F31AG057150 (E.A.A.J.) and TR001743 and ES028047 (B.C.) from the National Institutes of Health. The grant funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. Data from GSE15222 were downloaded from the NCBI GEO database. We thank R. Thomas for assistance with snRNA-seq analyses and T. Pak for editorial assistance.
Author information
Authors and Affiliations
Contributions
A.T., M.S. and Y. Huang designed and coordinated the study. A.T. carried out most of the studies and data analyses. M.Y.Z. performed brain slice electrophysiological recordings and data analysis. Y. Hao prepared snRNA-seq and bulk RNA-seq samples. K.C. helped analyze the publicly available bulk RNA-seq dataset (https://doi.org/10.7303/syn20808171) from APOE4-KI mice. K.C. and B.G. helped with snRNA-seq data analyses of J20/E4-KI mice. M.E.B. cultured human iPSC-derived neurons. P.N., S.Y.Y. and E.A.A.J. performed genetic screening of mice and behavioral tests. K.A.Z., S.P., B.C., N.C., N.K., C.W., W.C. and A.A. helped with tissue sectioning, immunostaining or data analyses. I.K., T.O. and M.S. led study design and data analysis for the UCSF EHR study. M.B., F.C., I.P., J.D.F. and B.G. led study design and analysis for the Mt. Sinai EHR study. A.T., K.A.Z., M.S. and Y. Huang wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y. Huang is a cofounder and scientific advisory board member of ESCAPE Bio, GABAeron and Mederon Bio. M.S. is on the advisory board of Aria Pharmaceuticals. A pending patent application related to this work has been filed on which Y. Huang, A.T., M.S. and P.N. were listed as inventors. The other authors declare no competing financial interests.
Additional information
Peer review information Nature Aging thanks Na Zhao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bumetanide is also predicted to rescue the transcriptomic signature of aging in apoE4-KI mouse cortex.
a, PCA plot of top 500 variable genes in apoE4-KI mouse cortex shows a distinct effect of age, with 3 month-old brains grouping separately from 12 and 24 month-old brains. b, Venn diagrams of the overlapping DE genes (logFC > 2, unadjusted P < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) and DE pathways (unadjusted P < 0.05 by bespoke enrichment method, kegga function (limma v 3.36.5)) between 12 vs 3 month-old apoE4-KI brains and 24 vs 3 month-old apoE4-KI brains. c, Graphs of compounds ordered by CMap score against DE genes (logFC > 2, unadjusted P < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 12 vs 3 month-old apoE4-KI brains (see Methods for details). Bumetanide has a negative CMap score in the 7th percentile of all drugs in the CMap. d, Graphs of compounds ordered by CMap score against DE genes (logFC > 2, unadjusted P < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 24 vs 3 month-old apoE4-KI brains. Bumetanide has a negative CMap score in the 8th percentile of all drugs in the CMap. e, Histogram of the rank of FC of the DE genes (logFC > 2, unadjusted P < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 12 vs 3 month-old apoE4-KI brains, which were also measured in the CMap database after bumetanide treatment. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the dashed red line and the mean FC rank of the down-regulated genes (colored blue in histogram) is denoted by the blue dashed line. P-value of significance of the “flip” of up- and down-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P = 0.056). This p-value does not reach significance even while the magnitude of the “flip” is quite large. f, Histogram of the rank of FC of the DE genes (logFC > 2, unadjusted p-value < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 24 vs 3 month-old apoE4-KI brains which were also measured in the CMap database after bumetanide treatment. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the dashed red line and the mean FC rank of the down-regulated genes (colored blue in histogram) is denoted by the blue dashed line. P-value of significance of the “flip” of up- and down-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P = 0.064). This p-value does not reach significance even while the magnitude of the “flip” is quite large.
Extended Data Fig. 2 Bumetanide treatment does not affect swim speed or visible trial performance in aged apoE4-KI mice and does not affect behavioral performance in wildtype (WT) mice.
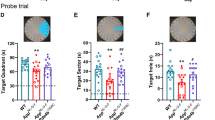
a, Bumetanide did not significantly affect swim speed during hidden platform trials of apoE4-KI and apoE3-KI mice (n = 11 for each group) at 24 month of age over learning days 1–5. b, There was no significant difference between any groups in visible trials (measured by 2-way ANOVA) of apoE4-KI and apoE3-KI mice (n = 11 for each group), indicating there were no motor or vision impairment in any of the groups. c, Escape latency of vehicle (n = 16) and bumetanide (n = 15) treated WT mice during learning days 1–5 did not differ. d, Bumetanide did not significantly affect swim speed during hidden platform trials in WT mice (n = 15) as compared to vehicle treated WT controls (n = 16). e, In the 24-hour probe trial, both vehicle (n = 16, two way ANOVA with Bonferroni’s multiple comparisons test P < 0.0001) and bumetanide (n = 15, two way ANOVA with Bonferroni’s multiple comparisons test P = 0.0001) treated WT mice spent significantly more time in the target quadrant versus average percent time in the other quadrants. f, In the 72-hour probe trial, both vehicle (n = 16, two way ANOVA with Bonferroni’s multiple comparisons test P = 0.0054) and bumetanide (n = 15, two way ANOVA with Bonferroni’s multiple comparisons test P < 0.0001) treated WT mice spent more time in the target quadrant than the other quadrants. Values are mean ± SEM in a-f.
Extended Data Fig. 3 Violin plots of marker genes for 18 cell clusters and their properties identified by snRNA-seq in the hippocampus of aged apoE4-KI mice.
a, Violin plots of expression of marker genes for each of the 18 cell clusters identified by snRNA-seq in the hippocampus of aged apoE4-KI mice with and without bumetanide treatment. Y-axis is average imputed expression of a marker gene across all cells in a cluster (see Methods for details), x-axis denotes each cell cluster. b, snRNA-seq analysis of the hippocampus of aged apoE4-KI mice with and without bumetanide treatment identifies 18 unique cell clusters. c, Number of cells per cluster. d, Average number of genes identified per cell in each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 58 cells in any cluster). e, Average nUMI per cell for each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 58 cells in any cluster). f, Average % mitochondrial genes per cell in each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 58 cells in any cluster).
Extended Data Fig. 4 Histograms of FC rank changes of human apoE4/4-specific AD signature genes in cell clusters 1, 4, 6, 7, 9–18 in aged apoE4-KI mice.
a–n, Histograms of the human apoE4/4-specific transcriptomic signature of AD geneset that was also detected by DE analysis of snRNA-seq in the apoE4-KI mouse hippocampus after bumetanide treatment as compared to controls in cell clusters 1, 4, 6, 7, 9–18. The rank of the FC of these genes in each cluster following bumetanide treatment, as compared to vehicle treatment, is plotted. The mean rank of all genes in this geneset is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the red dashed line and the mean FC rank of the down-regulated genes (colored blue in histogram) is denoted by the blue dashed line. P-value of the significance of the “flip” of up- and down-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P < 0.05 considered significant). Cell clusters 1, 4, 6, 9, 10, 13, 15, and 16, which include all excitatory neurons, mixed neurons, and endothelial/fibroblast-like cells have a significant “flip” of human apoE4/4-specific AD signiture genes, whereas cell clusters 7, 11, 12, 14, 17 and 18, which include oligodendrocytes, VIP-interneurons, OPC’s, RELN-interneurons, astrocytes, choroid plexus, are not significant. o, Histogram of the human apoE4/4-specific transcriptomic signature of AD geneset that was also detected by DE analysis of snRNA-seq in the apoE4-KI mouse hippocampus after bumetanide treatment as compared to controls in combined data from all neuronal clusters that exhibited a significant “flip” of human apoE4/4 AD genes (Clusters 1, 2, 3, 4, 5, 6, 8, 9, 10, 13, and 16 combined). The rank of the FC of these genes in the combined neuronal cells following bumetanide treatment, as compared to vehicle treatment, is plotted. The mean rank of all genes in this geneset is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the red dashed line and the mean FC rank of the down-regulated genes (colored blue in histogram) is denoted by the blue dashed line. P-value of the significance of the “flip” of up- and down-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P < 0.05 considered significant).
Extended Data Fig. 5 The fold change size and directionality of all DE genes after bumetanide treatment in the five large excitatory neuronal cell types in aged apoE4-KI mouse hippocampus mimicked the fold change size and directionality after bumetanide treatment in PC3 cells in the CMap database.
a, Scatterplot of average number of cells per cell cluster (of clusters with > 50 cells) in apoE4-KI hippocampi versus the percentile of CMap score against the DE genes in those clusters after bumetanide treatment (see Methods for details). The top 300 DE genes by p-value of the first five cell clusters (all excitatory neuronal cells) have a CMap score above the top 90 percentile of all drugs in the CMap database. b, Graphs of compounds ordered by CMap score against DE genes in Dentate Gyrus Granule Cells (see Methods for details). Bumetanide has one of the highest positive scores, suggesting that the signature in these cells in vivo is similar to the signature in the CMap database. c, Correlation analysis plot of rank of FC of genes in apoE4-KI Dentate Gyrus Granule Cells after bumetanide treatment versus rank of FC of genes in the CMap database after bumetanide treatment. There is a positive correlation (by the “lm” linear model method, geom_smooth function with default parameters (Ggplot2 v_3.2.1)) with an R2 = 0.08829 and an unadjusted P-value = 3.6 ×10−12, indicating that the FC of genes in these two signatures of DE genes after bumetanide treatment mimic each other. The shaded region represents the 95% confidence interval for predictions from the linear model. d, Graphs of compounds ordered by CMap score against DE genes in apoE4-KI CA1 neurons (see Methods for details). Bumetanide has one of the highest positive scores, suggesting that the signature in these cells in vivo is similar to the signature in the CMap database. e, Correlation analysis plot of rank of FC of genes in apoE4-KI CA1 neurons after bumetanide treatment versus rank of FC of genes in the CMap database after bumetanide treatment. There is a positive correlation (by the “lm” linear model method, geom_smooth function with default parameters (Ggplot2 v_3.2.1)) with an R2 = 0.08091 and an unadjusted P-value = 1.9 ×10−10, indicating that the FC of genes in these two signatures of DE genes after bumetanide treatment mimic each other. The shaded region represents the 95% confidence interval for predictions from the linear model. f, Graphs of compounds ordered by CMap score against DE genes in apoE4-KI CA 2/3 Neurons (see Methods for details). Bumetanide has one of the highest positive scores, suggesting that the signature in these cells in vivo is similar to the signature in the CMap database. g, Correlation analysis plot of rank of FC of genes in apoE4-KI CA2/3 neurons after bumetanide treatment versus rank of FC of genes in the CMap database after bumetanide treatment. There is a positive correlation (by the “lm” linear model method, geom_smooth function with default parameters (Ggplot2 v_3.2.1)) with an R2 = 0.06221 and an unadjusted P-value = 9.9 ×10−7, suggesting that the FC of genes in these two signatures of DE genes after bumetanide treatment mimic each other. The shaded region represents the 95% confidence interval for predictions from the linear model.
Extended Data Fig. 6 Bumetanide treatment flips apoE4-mediated murine transcriptomic signature of aging in specific neuron subtypes in the hippocampus of aged apoE4-KI mice.
a, Number of upregulated (119) and downregulated (3) genes (logFC > 2, unadjusted P < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 24 vs 3 month-old apoE4-KI cortex. b, Number of overlapping genes that were detected in clusters 1–18 in apoE4-KI hippocampi with and without bumetanide treatment. Due to sequencing depth and gene drop out, none of the 3 downregulated genes were detected in any cell cluster in aged apoE4-KI hippocampi. c, P-value of the “flip” of apoE4/4 specific transcriptomic signatures of AD in humans is plotted on the x-axis versus the P-value of the “flip” of aging signature of upregulated DE genes in 24 vs 3 month-old apoE4-KI hippocampus for each of the 18 cell clusters after bumetanide treatment in apoE4-KI mouse hippocampus, as calculated by Monte-Carlo simulation. The black dashed lines denotes P = 0.05. Cell clusters 1, 2 and 4 have a significant P-value in each analysis, while Clusters 1–6 and cluster 13 have P-values either trending towards or reaching significance in both analyses, suggesting that most exitatory neuronal clusters experience a “flip” of both aging and apoE4/4 AD signatures when exposed to bumetanide in apoE4-KI hippocampi. d, Histogram of the rank of FC after bumetanide treatment in apoE4-KI Dentate Gyrus Granule Cell (Cluster 1) of the aging DE genes (logFC > 2, unadjusted P-value < 0.05 by Wald test with default parameters (DESeq2 v 1.20.2)) in 24 vs 3 month-old apoE4-KI hippocampus. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the dashed red line. P-value of significance of the “flip” of up-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P = 0.015). e, Histogram of the rank of FC after bumetanide treatment in apoE4-KI Dentate Gyrus Granule Cell (Cluster 2) of the aging DE genes (logFC > 2, unadjusted P-value < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 24 vs 3 month-old apoE4-KI hippocampus. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the dashed red line. P-value of significance of the “flip” of up-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P = 0.031). f, Histogram of the rank of FC after bumetanide treatment in apoE4-KI CA 2/3 Neurons (Cluster 4) of the aging DE genes (logFC > 2, unadjusted P-value < 0.05 by Wald test with default parameters (DESeq2 v 1.30.2)) in 24 vs 3 month-old apoE4-KI hippocampus. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the dashed red line. P-value of significance of the “flip” of up-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P = 0.007).
Extended Data Fig. 7 Violin plots of marker genes for 25 cell clusters and their properties identified by snRNA-seq in the hippocampus of J20/E4-KI mice.
a, Violin plots of expression of marker genes for each of the 25 cell clusters identified by snRNA-Seq in the hippocampus of J20/E4-KI mice with and without bumetanide treatment. Y-axis is average imputed expression of a marker gene across all cells in a cluster (see Methods for details), x-axis denotes each cell cluster. b, snRNA-seq analysis of the hippocampus of aged apoE4-KI mice with and without bumetanide treatment identifies 25 unique cell clusters. c, Number of cells per cluster. d, Average number of genes identified per cell in each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 177 cells in any cluster). e, Average nUMI per cell for each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 177 cells in any cluster). f, Average % mitochondrial genes per cell in each cluster (± SEM). Number of cells (n) for each cell cluster can be found in c (> 177 cells in any cluster).
Extended Data Fig. 8 snRNA-seq analysis of the transcriptomic perturbation signature of bumetanide in the hippocampus of J20/E4-KI mice.
a, Transcripts in 47,619 single nuclei from the hippocampus of bumetanide- and vehicle-treated J20/E4-KI mice (n = 3 mice per group) were sequenced. b, Clustering and visualization by t-SNE identifies 25 distinct cell clusters which are color-coded according to cell-type. c, Cell clusters color-coded by treatment groups. d, Histogram of the rank of FC of the human apoE4/4-specific transcriptomic signature of AD genes that were also detected by snRNA-seq in J20/E4-KI mouse hippocampi (calculated via FindMarkers with default parameters, Wilcoxon rank sum test, Seurat v_3.1.5.9005) in four representative cell clusters. The rank of the FC of these genes in a dentate gyrus granule cell cluster (1), a subiculum neuronal cluster (5), a microglial cluster (10), and an astrocyte cluster (17) in J20/E4-KI mouse hippocampi following bumetanide treatment as compared to vehicle treatment were plotted. The mean rank of all genes in this gene set is denoted by the black line, the average mean FC rank of up-regulated genes (colored red in histogram) is denoted by the red dashed line and the mean FC rank of down-regulated genes (colored blue in histogram) is denoted by the blue dashed line. P-value of significance of the “flip” of up- and down-regulated FC rank means away from the rank mean of all genes as calculated by Monte-Carlo simulation is shown (P < 0.05 considered significant. e, Heatmap of genes from apoE4/4-specific transcriptomic signature of AD, rank ordered and color coded (red for up, blue for down) by estimated FC in human apoE4/4 AD (top), then re-ordered by FC rank after bumetanide treatment in four representative cell types (clusters 1, 5, 10 and 17) in the J20/E4-KI mouse hippocampus. Bumetanide treatment flips the expression rank of both up- and down-regulated genes of the human apoE4/4-specific transcriptomic signature of AD in these four cell types. f, P-value of the “flip” of the apoE4/4-specific transcriptomic signature of AD, as calculated by Monte-Carlo simulation, is plotted on the y-axis versus the number of DE genes in each cell cluster on the x-axis. The red dashed line denotes P = 0.05. Dentate gyrus granule cells, subiculum neurons, OPCs, microglia, and astrocytes exhibit a significant “flip” of the apoE4/4-specific transcriptomic signature of genes in AD despite varying number of DE genes. g, Heatmap of the p-values of enriched ontological pathways in all cell clusters exhibiting the “flip” behavior of the apoE4/4-specific transcriptomic signatures of AD reveals 37 pathways that are affected in at least one of these cell types (unadjusted P < 0.005 by bespoke enrichment method, kegga function (limma v 3.36.5)). Pathway names highlighted in red (n = 7) are those shared with the apoE4/4-specific signature pathways of AD (see Fig. 1g and Supplementary Table 6 for human pathways).
Extended Data Fig. 9 Analyses of overlapping enriched ontological pathways among bumetanide-treated apoE4-KI mice, J20/E4-KI mice, human iPSC-derived neurons, and human apoE4/4-specific transcriptomic signature of AD.
a, 22 overlapping enriched ontological pathways (Supplementary Table 17) in bumetanide-flipped cell clusters in apoE4-KI mice vs J20/E4-KI mice. b, Six overlapping enriched ontological pathways (Supplementary Table 18) in bumetanide-flipped cell clusters in apoE4-KI mice vs J20/E4-KI mice vs E4/4-hiPSC neurons. c, Three overlapping enriched ontological pathways in bumetanide-flipped cell clusters in apoE4-KI mice vs J20/E4-KI mice vs E4/4-hiPSC neurons vs human E4/4 signature of AD, which include GABAergic Synapse, Circadian Entrainment, Morphine Addiction pathways.
Extended Data Fig. 10 Bumetanide exposure is associated with a significantly lower AD prevalence in individuals over the age of 65 in two independent EHR databases.
We evaluated two large-scale EHR databases (UCSF EHR and Mt. Sinai EHR) in a cross-sectional manner to test the association of bumetanide exposure with AD prevalence in individuals with the age of 65 or above using a propensity score matching approach to control cohort creation. a, AD prevalence in bumetanide-exposed cohort is significantly lower than those in all 10 randomly selected non-bumetanide-exposed cohorts in the UCSF EHR database. All 10 randomly selected 1:2 control cohorts were matched on propensity score which included age, sex, race, and hypertension and edema diagnosis. Two-sided χ2 test, df = 1, χ2 values shown, all P < 0.05, unadjusted p-values shown. b, AD prevalence in bumetanide-exposed cohort is significantly lower than those in all 10 randomly selected non-bumetanide-exposed cohorts in the Mt. Sinai EHR database. All 10 randomly selected 1:2 control cohorts were matched on propensity score which included age, sex, race, and hypertension and edema diagnosis. Two-sided χ2 test, df = 1, χ2 values shown, all P < 0.0001, unadjusted p-values shown. c, AD prevalence in bumetanide-exposed cohort is significantly lower than those in 8 out of 10 randomly selected non-bumetanide-exposed cohorts controlled for non-bumetanide diuretic drug use for hypertension and edema treatment in the UCSF EHR database. Two-sided χ2 test, df = 1, χ2 values shown, 8 out of 10 P < 0.05, unadjusted p-values shown. d, AD prevalence in bumetanide-exposed cohort is significantly lower than those in all 10 randomly selected non-bumetanide-exposed cohorts controlled for non-bumetanide diuretic drug use for hypertension and edema treatment in the Mt. Sinai EHR database. Two-sided χ2 test, df = 1, χ2 values shown, all P < 0.0001, unadjusted p-values shown.
Supplementary information
Supplementary Table 2
DE genes in human APOE4/APOE4 AD versus the APOE4/APOE4 control. Includes DE genes from each of the ten permutations and DE genes from the union of all ten permutations.
Supplementary Table 3
DE genes in human APOE3/APOE3 AD versus the APOE3/APOE3 control. Includes DE genes from each of the ten permutations and DE genes from the union of all ten permutations.
Supplementary Table 4
DE genes in human APOE3/APOE4 AD versus the APOE3/APOE4 control. Includes DE genes from each of the ten permutations and DE genes from the union of all ten permutations.
Supplementary Table 5
DE genes in all AD samples versus all control samples. Includes DE genes from each of the ten permutations and DE genes from the union of all ten permutations.
Supplementary Table 6
DE pathways in human AD by APOE genotype. Includes DE pathways in APOE4/APOE4 AD, APOE3/APOE4 AD and APOE3/APOE3 AD.
Supplementary Table 7
DE genes in the cortex of old versus young APOE4/APOE4-KI mice. Includes DE genes in the cortex of 12- versus 3-month-old APOE4-KI mice and DE genes in the cortex of 24- versus 3-month-old APOE4-KI mice from a publicly available bulk RNA-seq dataset.
Supplementary Table 8
DE pathways in the cortex of old versus young APOE4/APOE4-KI mice. Includes DE pathways in the cortex of 12- versus 3-month-old APOE4-KI mice and DE pathways in the cortex of 24- versus 3-month-old APOE4-KI mice from a publicly available bulk RNA-seq dataset.
Supplementary Table 9
Marker genes for all cell clusters in the snRNA-seq dataset from the APOE4-KI mouse hippocampus. Includes marker genes for every cell cluster in the APOE4-KI mouse hippocampal dataset.
Supplementary Table 10
DE genes in hippocampi of bumetanide-treated APOE4-KI mice versus vehicle-treated APOE4-KI mice. Includes DE genes in every cell cluster in the APOE4-KI mouse hippocampus after bumetanide treatment.
Supplementary Table 11
DE pathways in hippocampi of bumetanide-treated APOE4-KI mice versus vehicle-treated APOE4-KI mice. Includes DE pathways in every cell cluster in the APOE4-KI mouse hippocampus after bumetanide treatment.
Supplementary Table 12
Marker genes for all cell clusters in the snRNA-seq dataset from the J20/E4-KI mouse hippocampus. Includes marker genes for every cell cluster in the J20/E4-KI mouse hippocampal dataset.
Supplementary Table 13
DE genes in hippocampi of bumetanide-treated J20/E4-KI mice versus vehicle-treated J20/E4-KI mice. Includes DE genes in every cell cluster in the J20/E4-KI mouse hippocampus after bumetanide treatment.
Supplementary Table 14
DE pathways in hippocampi of bumetanide-treated J20/E4-KI mice versus vehicle-treated J20/E4-KI mice. Includes DE pathways in every cell cluster in the J20/E4-KI mouse hippocampus after bumetanide treatment.
Supplementary Table 15
DE genes in bumetanide-treated APOE4/APOE4 hiPSC-derived neurons versus vehicle-treated APOE4/APOE4 hiPSC-derived neurons. Includes DE genes in APOE4/APOE4 hiPSC-derived neurons after bumetanide treatment.
Supplementary Table 16
DE pathways in bumetanide-treated APOE4/APOE4 hiPSC-derived neurons versus vehicle-treated APOE4/APOE4 hiPSC-derived neurons. Includes DE pathways in APOE4/APOE4 hiPSC-derived neurons after bumetanide treatment.
Supplementary Table 17
DE pathways shared in hippocampi of bumetanide-treated APOE4-KI and J20/E4-KI mice. Includes all DE pathways shared by hippocampi of bumetanide-treated APOE4-KI and J20/E4-KI mice.
Supplementary Table 18
DE pathways shared in hippocampi of bumetanide-treated APOE4-KI and J20/E4-KI mice as well as bumetanide-treated APOE4/APOE4 iPSC-derived neurons. Includes all DE pathways shared by hippocampi of bumetanide-treated APOE4-KI and J20/E4-KI mice as well as bumetanide-treated APOE4/APOE4 iPSC-derived neurons.
Source data
Source Data Fig. 3
Statistical source data for Fig. 3a–e.
Source Data Fig. 5
Statistical source data for Fig. 5a–c,f–i.
Source Data Fig. 5
Image source data for Fig. 5d,e.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2a–f.
Rights and permissions
About this article
Cite this article
Taubes, A., Nova, P., Zalocusky, K.A. et al. Experimental and real-world evidence supporting the computational repurposing of bumetanide for APOE4-related Alzheimer’s disease. Nat Aging 1, 932–947 (2021). https://doi.org/10.1038/s43587-021-00122-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00122-7
This article is cited by
-
Leveraging electronic health records and knowledge networks for Alzheimer’s disease prediction and sex-specific biological insights
Nature Aging (2024)
-
APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia
Nature (2024)
-
Neuronal APOE4 removal protects against tau-mediated gliosis, neurodegeneration and myelin deficits
Nature Aging (2023)
-
Clinical relevance of animal models in aging-related dementia research
Nature Aging (2023)
-
Insights into Alzheimer’s disease from single-cell genomic approaches
Nature Neuroscience (2023)