Abstract
The plasma proteomic changes that precede the onset of dementia could yield insights into disease biology and highlight new biomarkers and avenues for intervention. We quantified 4,877 plasma proteins in nondemented older adults in the Atherosclerosis Risk in Communities cohort and performed a proteome-wide association study of dementia risk over five years (n = 4,110; 428 incident cases). Thirty-eight proteins were associated with incident dementia after Bonferroni correction. Of these, 16 were also associated with late-life dementia risk when measured in plasma collected nearly 20 years earlier, during mid-life. Two-sample Mendelian randomization causally implicated two dementia-associated proteins (SVEP1 and angiostatin) in Alzheimer’s disease. SVEP1, an immunologically relevant cellular adhesion protein, was found to be part of larger dementia-associated protein networks, and circulating levels were associated with atrophy in brain regions vulnerable to Alzheimer’s pathology. Pathway analyses for the broader set of dementia-associated proteins implicated immune, lipid, metabolic signaling and hemostasis pathways in dementia pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated in this study are either included in this article (and its Supplementary Information), available upon reasonable request or are available in an online public database. Consistent with a prespecified policy for access of ARIC and AGES data, requests may be submitted to the ARIC and AGES steering committees for review. Requests for clinical or proteomic data from individual investigators will be reviewed to ensure that data can be shared without compromising patient confidentiality or breaching intellectual property restrictions. Reasonable requests will be considered and promptly processed. Participant-level demographic, clinical and proteomic data may be partially restricted based on previously obtained participant consent. Data sharing restrictions may also be applied to ensure consistency with confidentiality or privacy laws and considerations. For information on how to access available data and study protocols, see www2.cscc.unc.edu/aric/. Data from the AGES-Reykjavik study used in this study are available through collaboration (AGES_data_request@hjarta.is) under a data usage agreement with the Icelandic Heart Association. Tissue-specific gene expression data are available at https://www.gtexportal.org/home/. Gene coexpression analyses were conducted using data available at http://www.explainbio.com. Brain gene expression data were derived from the Brain eQTL Almanac (http://www.braineac.org/) and the Functional Mapping and Annotation of Genome-Wide Association Studies105 platform (https://fuma.ctglab.nl/). eQTL gene enrichment was performed using data from the Molecular Signatures database (http://www.broadinstitute.org/msigdb) and the NHGRI-EBI Catalog of Published GWAS (https://www.ebi.ac.uk/gwas/). Functional enrichment of protein networks was conducted using the g:Profiler web tool (https://biit.cs.ut.ee/gprofiler/gost).
Code availability
All software used in this study are publicly available: R v.3.6.2 (https://www.r-project.org/); Stata, v.14 (https://www.stata.com/stata14/); IPA (https://digitalinsights.qiagen.com/products-overview/discovery-insights-portfolio/analysis-and-visualization/qiagen-ipa/); ExplainBio (http://www.explainbio.com/); and GraphPad Prism v.8.4.3. (https://www.graphpad.com/scientific-software/prism/). The code used in this study can be made available from the corresponding author upon reasonable request.
Change history
04 July 2022
In the version of this article initially published, the Supplementary Tables 1–31 file was missing and has now been restored to the online version of the article.
References
Nakamura, A. et al. High-performance plasma amyloid-β biomarkers for Alzheimer’s disease. Nature 554, 249–254 (2018).
Mattsson, N., Cullen, N. C., Andreasson, U., Zetterberg, H. & Blennow, K. Association between longitudinal plasma neurofilament light and neurodegeneration in patients with Alzheimer disease. JAMA Neurol. 76, 791–799 (2019).
Palmqvist, S. et al. Cerebrospinal fluid and plasma biomarker trajectories with increasing amyloid deposition in Alzheimer’s disease. EMBO Mol. Med. https://doi.org/10.15252/emmm.201911170 (2019).
Kim, K. et al. Clinically accurate diagnosis of Alzheimer’s disease via multiplexed sensing of core biomarkers in human plasma. Nat. Commun. 11, 119 (2020).
Soares, H. D. et al. Plasma biomarkers associated with the apolipoprotein E genotype and Alzheimer disease. Arch. Neurol. 69, 1310 (2012).
Doecke, J. D. et al. Blood-based protein biomarkers for diagnosis of Alzheimer disease. Arch. Neurol. 69, 1318 (2012).
Hye, A. et al. Proteome-based plasma biomarkers for Alzheimer’s disease. Brain 129, 3042–3050 (2006).
Cutler, P. et al. Proteomic identification and early validation of complement 1 inhibitor and pigment epithelium-derived factor: two novel biomarkers of Alzheimer’s disease in human plasma. Proteomics. Clin. Appl. 2, 467–477 (2008).
Choi, J., Malakowsky, C. A., Talent, J. M., Conrad, C. C. & Gracy, R. W. Identification of oxidized plasma proteins in Alzheimer’s disease. Biochem. Biophys. Res. Commun. 293, 1566–1570 (2002).
Henkel, A. W. et al. Multidimensional plasma protein separation technique for identification of potential Alzheimer’s disease plasma biomarkers: a pilot study. J. Neural Transm. 119, 779–788 (2012).
Ray, S. et al. Classification and prediction of clinical Alzheimer’s diagnosis based on plasma signaling proteins. Nat. Med. 13, 1359–1362 (2007).
Ijsselstijn, L. et al. Serum levels of pregnancy zone protein are elevated in presymptomatic Alzheimer’s disease. J. Proteome Res. 10, 4902–4910 (2011).
O’Bryant, S. E. et al. A serum protein-based algorithm for the detection of Alzheimer disease. Arch. Neurol. 67, 1077–1081 (2010).
Sattlecker, M. et al. Alzheimer’s disease biomarker discovery using SOMAscan multiplexed protein technology. Alzheimers Dement. 10, 724–734 (2014).
Kiddle, S. J. et al. Candidate blood proteome markers of Alzheimer’s disease onset and progression: a systematic review and replication study. J. Alzheimers Dis. 38, 515–531 (2014).
Sattlecker, M. et al. Longitudinal protein changes in blood plasma associated with the rate of cognitive decline in Alzheimer’s disease. J. Alzheimers Dis. 49, 1105–1114 (2016).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Sharma, K. et al. Cell type- and brain region-resolved mouse brain proteome. Nat. Neurosci. 18, 1819–1831 (2015).
Seyfried, N. T. et al. A multi-network approach identifies protein-specific co-expression in asymptomatic and symptomatic Alzheimer’s disease. Cell Syst. 4, 60–72 (2017).
Ganz, P. et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA 315, 2532–2541 (2016).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Shadlen, M. F. et al. Education, cognitive test scores, and black-white differences in dementia risk. J. Am. Geriatr. Soc. 54, 898–905 (2006).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Harris, T. B. et al. Age, gene/environment susceptibility-Reykjavik study: multidisciplinary applied phenomics. Am. J. Epidemiol. 165, 1076–1087 (2007).
Emilsson, V. et al. Co-regulatory networks of human serum proteins link genetics to disease. Science 361, 769–773 (2018).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Cissé, M. & Checler, F. Eph receptors: new players in Alzheimer’s disease pathogenesis. Neurobiol. Dis. 73, 137–149 (2015).
Shi, L. et al. A decade of blood biomarkers for Alzheimer’s disease research: an evolving field, improving study designs, and the challenge of replication. J. Alzheimers Dis. 62, 1181–1198 (2018).
Palmqvist, S. et al. Discriminative accuracy of plasma phospho-tau217 for Alzheimer disease vs other neurodegenerative disorders. JAMA 324, 772–781 (2020).
Ghosh, S. et al. Sustained interleukin-1 overexpression exacerbates tau pathology despite reduced amyloid burden in an Alzheimer’s mouse model. J. Neurosci. 33, 5053–5064 (2013).
Paolicelli, R. C. et al. TDP-43 depletion in microglia promotes amyloid clearance but also induces synapse loss. Neuron 95, 297–308 (2017).
Varma, S. Network model of normal gene expression predicts gene perturbation fold changes. In Proc. RECOMB/ISCB Conference on Regulatory and Systems Genomics with DREAM Challenges 78–79 (2015).
Savli, H., Szendröi, A., Romics, I. & Nagy, B. Gene network and canonical pathway analysis in prostate cancer: a microarray study. Exp. Mol. Med. 40, 176–185 (2008).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013).
Jones, L. et al. Convergent genetic and expression data implicate immunity in Alzheimer’s disease. Alzheimers Dement. 11, 658–671 (2015).
Felsky, D. et al. Neuropathological correlates and genetic architecture of microglial activation in elderly human brain. Nat. Commun. https://doi.org/10.1038/s41467-018-08279-3 (2019).
Walker, K. A. et al. The association of mid-to late-life systemic inflammation with white matter structure in older adults: the Atherosclerosis Risk in Communities study. Neurobiol. Aging 68, 26–33 (2018).
Steeland, S. et al. Counteracting the effects of TNF receptor‐1 has therapeutic potential in Alzheimer’s disease. EMBO Mol. Med. https://doi.org/10.15252/emmm.201708300 (2018).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Neumann, H. & Daly, M. J. Variant TREM2 as risk factor for Alzheimer’s disease. N. Engl. J. Med. 368, 182–184 (2013).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Parts, L., Stegle, O., Winn, J. & Durbin, R. Joint genetic analysis of gene expression data with inferred cellular phenotypes. PLoS Genet. 7, e1001276 (2011).
Trimbuch, T. & Rosenmund, C. Should I stop or should I go? The role of complexin in neurotransmitter release. Nat. Rev. Neurosci. 17, 118–125 (2016).
An, S. J., Grabner, C. P. & Zenisek, D. Real-time visualization of complexin during single exocytic events. Nat. Neurosci. 13, 577–583 (2010).
Tannenberg, R. K., Scott, H. L., Tannenberg, A. E. G. & Dodd, P. R. Selective loss of synaptic proteins in Alzheimer’s disease: evidence for an increased severity with APOE ε4. Neurochem. Int. 49, 631–639 (2006).
Nakada, T. A., Russell, J. A., Boyd, J. H., Thair, S. A. & Walley, K. R. Identification of a nonsynonymous polymorphism in the SVEP1 gene associated with altered clinical outcomes in septic shock. Crit. Care Med. 43, 101–108 (2015).
Siddiqui, S., Gurung, R. L., Liu, S., Seet, E. C. P. & Lim, S. C. Genetic polymorphisms and cytokine profile of different ethnicities in septic shock patients, and their association with mortality. Indian J. Crit. Care Med. 23, 135–138 (2019).
Sato-Nishiuchi, R. et al. Polydom/SVEP1 is a ligand for integrin α9β1. J. Biol. Chem. 287, 25615–25630 (2012).
Schwanzer-Pfeiffer, D., Roßmanith, E., Schildberger, A. & Falkenhagen, D. Characterization of SVEP1, KIAA, and SRPX2 in an in vitro cell culture model of endotoxemia. Cell. Immunol. 263, 65–70 (2010).
Richter, V. et al. Circulating vascular cell adhesion molecules VCAM-1, ICAM-1, and E-selectin in dependence on aging. Gerontology 49, 293–300 (2003).
Yousef, H. et al. Aged blood impairs hippocampal neural precursor activity and activates microglia via brain endothelial cell VCAM1. Nat. Med. 25, 988–1000 (2019).
Janelidze, S. et al. CSF biomarkers of neuroinflammation and cerebrovascular dysfunction in early Alzheimer disease. Neurology 91, e867–e877 (2018).
K Ryu, J., P Little, J., Klegeris, A., Jantaratnotai, N. & G McLarnon, J. Actions of the anti-angiogenic compound angiostatin in an animal model of Alzheimer’s disease. Curr. Alzheimers Res. 10, 252–260 (2013).
Jahanshad, N. et al. Genome-wide scan of healthy human connectome discovers SPON1 gene variant influencing dementia severity. Proc. Natl Acad. Sci. USA 110, 4768–4773 (2013).
Bossù, P. et al. Interleukin-18 produced by peripheral blood cells is increased in Alzheimer’s disease and correlates with cognitive impairment. Brain. Behav. Immun. 22, 487–492 (2008).
Li, X., Long, J., He, T., Belshaw, R. & Scott, J. Integrated genomic approaches identify major pathways and upstream regulators in late-onset Alzheimer’s disease. Sci. Rep. 5, 12393 (2015).
Painter, M. M. et al. TREM2 in CNS homeostasis and neurodegenerative disease. Mol. Neurodegener. https://doi.org/10.1186/s13024-015-0040-9 (2015).
The ARIC Investigators. The Atherosclerosis Risk in Communities (ARIC) study: design and objectives. Am. J. Epidemiol. 129, 687–702 (1989).
Tin, A. et al. Reproducibility and variability of protein analytes measured using a multiplexed modified aptamer assay. J. Appl. Lab. Med. 4, 30–39 (2019).
Kim, C. H. et al. Stability and reproducibility of proteomic profiles measured with an aptamer-based platform. Sci. Rep. 8, 8382 (2018).
Candia, J. et al. Assessment of variability in the SOMAscan assay. Sci. Rep. 7, 14248 (2017).
Williams, S. A. et al. Plasma protein patterns as comprehensive indicators of health. Nat. Med. 25, 1851–1857 (2019).
Muntner, P., He, J., Astor, B. C., Folsom, A. R. & Coresh, J. Traditional and nontraditional risk factors predict coronary heart disease in chronic kidney disease: results from the Atherosclerosis Risk in Communities study. J. Am. Soc. Nephrol. 16, 529–538 (2005).
Knopman, D. S. et al. Mild cognitive impairment and dementia prevalence: the atherosclerosis risk in communities neurocognitive study. Alzheimers Dement. Diagnosis Assess. Dis. Monit. 2, 1–11 (2016).
Hayden, K. M. et al. Cognitive decline in the elderly: an analysis of population heterogeneity. Age Ageing 40, 684–689 (2011).
Wilson, R. S., Li, Y., Bienias, L. & Bennett, D. A. Cognitive decline in old age: separating retest effects from the effects of growing older. Psychol. Aging 21, 774–789 (2006).
McKhann, G. M. et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 7, 263–269 (2011).
American Psychiatric Association. DSM-5: Diagnostic and Statistical Manual of Mental Disorders (American Psychiatric Association, 2013).
Carpenter, C. R., Despain, B., Keeling, T. N., Shah, M. & Rothenberger, M. The six-item screener and AD8 for the detection of cognitive impairment in geriatric emergency department patients. Ann. Emerg. Med. 57, 653–661 (2011).
Galvin, J. E., Roe, C. M., Xiong, C. & Morris, J. C. Validity and reliability of the AD8 informant interview in dementia. Neurology 67, 1942–1948 (2006).
Walker, K. A. et al. Association of midlife to late-life blood pressure patterns with incident dementia. JAMA 322, 535–545 (2019).
Gottesman, R. F. et al. Associations between midlife vascular risk factors and 25-year incident dementia in the Atherosclerosis Risk in Communities (ARIC) cohort. JAMA Neurol. 388, 797–805 (2017).
Diagnostic and Statistical Manual of Mental Disorders: DSM-IV-TR (American Psychiatric Association, 2000).
Knopman, D. S. et al. Vascular imaging abnormalities and cognition: Mediation by cortical volume in nondemented individuals: atherosclerosis risk in communities-neurocognitive study. Stroke 46, 433–440 (2015).
Fischl, B. et al. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–355 (2002).
Dickerson, B. C. et al. Alzheimer-signature MRI biomarker predicts AD dementia in cognitively normal adults. Neurology 76, 1395–1402 (2011).
Raz, L. et al. Thrombogenic microvesicles and white matter hyperintensities in postmenopausal women. Neurology 80, 911–918 (2013).
Gottesman, R. F. et al. The ARIC-PET amyloid imaging study: brain amyloid differences by age, race, sex, and APOE. Neurology 87, 473–480 (2016).
Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37, 658–665 (2013).
Zheng, J. et al. Recent developments in Mendelian randomization studies. Curr. Epidemiol. Reports 4, 330–345 (2017).
Machiela, M. J. & Chanock, S. J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 31, 3555–3557 (2015).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Burgess, S., Foley, C. N., Allara, E., Staley, J. R. & Howson, J. M. M. A robust and efficient method for Mendelian randomization with hundreds of genetic variants. Nat. Commun. 11, 376 (2020).
Staiger, D. & Stock, J. H. Instrumental Variables Regression with Weak Instruments (National Bureau of Economic Research Inc., 1994).
Burgess, S. & Thompson, S. G. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur. J. Epidemiol. 32, 377–389 (2017).
Verbanck, M., Chen, C. Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Cochran, W. The combination of estimates from different experiments. Biometrics 10, 101–129 (1954).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife https://doi.org/10.7554/eLife.34408 (2018).
Burgess, S., Scott, R. A., Timpson, N. J., Davey Smith, G. & Thompson, S. G. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552 (2015).
Bowden, J. et al. Improving the visualization, interpretation and analysis of two-sample summary data Mendelian randomization via the Radial plot and Radial regression. Int. J. Epidemiol. 47, 1264–1278 (2018).
Watanabe, K., Taskesen, E., Van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. https://doi.org/10.1038/s41467-017-01261-5 (2017).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Ramasamy, A. et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat. Neurosci. 17, 1418–1428 (2014).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Lonsdale, J. et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Varma, V. R. et al. α-2 macroglobulin in Alzheimer’s disease: a marker of neuronal injury through the RCAN1 pathway. Mol. Psychiatry 22, 13–23 (2017).
Hage, C. et al. Inflammatory biomarkers predict heart failure severity and prognosis in patients with heart failure with preserved ejection fraction: a holistic proteomic approach. Circ. Cardiovasc. Genet. https://doi.org/10.1161/CIRCGENETICS.116.001633 (2017).
Helleman, J., Smid, M., Jansen, M. P. H. M., van der Burg, M. E. L. & Berns, E. M. J. J. Pathway analysis of gene lists associated with platinum-based chemotherapy resistance in ovarian cancer: the big picture. Gynecol. Oncol. 117, 170–176 (2010).
Krämer, A., Green, J., Pollard, J. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Raudvere, U. et al. G:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Gene, T. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25 (2000).
The Gene Ontology, C. et al. The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res. https://doi.org/10.17863/CAM.36439 (2019).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Bohler, A. et al. Reactome from a WikiPathways perspective. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1004941 (2016).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, 1–11 (2010).
Seddighi, S. et al. SPARCL1 accelerates symptom onset in Alzheimer’s disease and influences brain structure and function during aging. J. Alzheimers Dis. 61, 401–414 (2018).
Acknowledgements
We thank the staff and participants of the ARIC study for their important contributions. We also thank B. Chen for her valuable assistance with aspects of the manuscript. The ARIC study has been funded in whole or in part with Federal funds from the National Heart, Lung and Blood Institute (NHLBI), NIH, Department of Health and Human Services (contract nos. HHSN268201700001I, HHSN268201700002I, HHSN268201700003I, HHSN268201700004I, HHSN268201700005I, R01HL087641 and R01HL086694); National Human Genome Research Institute contract U01HG004402; and NIH contract HHSN268200625226C. Neurocognitive data are collected by U01 2U01HL096812, 2U01HL096814, 2U01HL096899, 2U01HL096902, 2U01HL096917 from the NIH (NHLBI, National Institute of Neurological Disorders and Stroke (NINDS), NIA and National Institute on Deafness and Other Communication Disorders (NIDCD)) and with previous brain MRI examinations funded by R01-HL70825 from the NHLBI. The ARIC-PET study is funded by the NIA (R01AG040282). Infrastructure was partly supported by grant no. UL1RR025005, a component of the NIH and NIH Roadmap for Medical Research. This study was also supported by contracts K23 AG064122 (to K.A.W.), K24 AG052573 (to R.F.G.) and U01-AG052409 (to M.F.) from NIA; and R01-HL134320 (to C.M.B.) from NHLBI. Avid Radiopharmaceuticals provided the florbetapir isotope for the study but had no role in the study design or interpretation of results. The AGES-Reykjavik study was supported by the Icelandic Heart Association, the NIA (N01-AG-12100 and HHSN271201200022C), the Intramural Program at the NIA, the Althingi (the Icelandic Parliament), the Icelandic Centre for Research grant 141101-051 and the Novartis Institute for Biomedical Research. SomaLogic provided assays as an in-kind contribution in a data flex change collaboration agreement. This research was supported in part by the Intramural Research Program of the NIH, NIA. Funders had no control over the publication.
Author information
Authors and Affiliations
Contributions
K.A.W., S.L.Z., R.F.G., T.H.M., E.B., C.M.B. and J. Coresh conceptualized and designed the study. M.G., N.D., R.C.H., C.M.B., V.E., L.J.L., L.L.J., V.G. and J. Coresh contributed to data acquisition. K.A.W., J. Chen, J.Z., M.F., Y.Y., L.Z., A.W. and E.F.G. analyzed data. K.A.W., M.F., Y.Y., M.G., A.T., P.G., N.C., R.F.G. and J. Coresh contributed to interpretation of data. K.A.W., M.F., Y.Y., M.G., A.T., R.C.H., A.W., K.J.S., P.G., L.J.L., R.F.G., C.M.B. and J. Coresh drafted and provided substantial revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.C.H. has received grants and consulting fees from Denka Seiken outside the scope of the current research study. A.W. received fees from the Analysis Group as a consultant outside the scope of the current research study. P.G. is a member of the SomaLogic Medical Advisory board, for which he receives no remuneration of any kind. L.L.J. is an employee and stockholder of Novartis. R.F.G. received fees from the American Academy of Neurology for her role as an Associate Editor for the journal Neurology. Proteomic assays in ARIC were conducted free of charge as part of a data exchange agreement with SomaLogic. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks Sebastian Palmqvist and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 P-values for dementia-associated proteins in mid-life replication and AGES-Reykjavik external replication analyses.
a, P-values (two-sided) for the mid-life replication analysis (index visit 1993-1995, ages 49-73) of the 38 dementia-associated proteins identified in older adults (y-axis) plotted against P-values for each dementia-associated protein derived from the primary analysis (index visit 2011-2013) on the x-axis. The horizontal dotted red line represents the Bonferroni-corrected threshold for statistical significance in the mid-life replication analysis (0.05/38; P < 0.0013). The vertical dotted black line represents the Bonferroni-corrected threshold for statistical significance in the primary analysis (0.05/4,877; P < 1.03×10-5). b, P-values (two-sided) for the AGES-Reykjavik (index visit 2002-2006) replication of proteins that were significantly associated with dementia risk in both the primary and the mid-life replication analysis (y-axis) plotted against P-values for each dementia-associated protein derived from the primary analysis (index visit 2011-2013) on the x-axis. The horizontal dotted red line represents the Bonferroni-corrected threshold for statistical significance in the AGES-Reykjavik replication analysis (0.05/13; P < 0.0038). The vertical dotted black line represents the Bonferroni-corrected threshold for statistical significance in the primary analysis (0.05/4,877; P < 1.03×10-5).
Extended Data Fig. 2 Prediction of incident dementia using proteins, demographic and clinical variables, and their combination measured at late-life baseline (2011-2013).
Elastic net machine learning with Cox proportional hazards regression was used to select the best combination of proteins from the top 50 proteins in each model. This table shows results from 10-fold cross validated analyses. Two-sided P-values were calculated for the C statistic comparisons. No corrections for multiple comparisons were performed. a Protein combination defined using elastic net machine learning algorithm. b Includes age, sex, race-center, education, and APOEε4. c Includes body mass index, diabetes, hypertension, smoking status, and eGFR-creatinine. Abbreviations: C stat. Δ, change in C statistic with the addition of elastic net proteins.
Extended Data Fig. 3 Gene expression in whole blood, brain, heart, kidney, spleen, and adipose tissue of genes coding for dementia-associated proteins.
Using gene expression data available from postmortem samples in the GTEx database, this heatmap shows the expression of genes coding for dementia-associated proteins (in transcripts per million) in whole blood, select brain regions, and other selected tissue. Hierarchical cluster analysis was used to group dementia-associated proteins based on gene expression across multiple tissues.
Extended Data Fig. 4 Functional profiling of protein Network 1 identified among the set of dementia-associated proteins.
Protein networks were assembled based on evidence of known gene/molecule interactions in the Ingenuity Knowledge Base. The g:Profiler toolkit106 was used to analyze the proteins in each dementia-associated protein network for functional enrichment. We defined the biological pathways/processes associated with each protein set using the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and WikiPathways databases. Images were generated using the g:Profiler web tool: https://biit.cs.ut.ee/gprofiler/. Abbreviations: GO:BP, gene ontology biological process; GO:MF, gene ontology molecular function.
Extended Data Fig. 5 Functional profiling of protein Network 2 and Network 3 identified among the set of dementia-associated proteins.
Protein networks were assembled based on evidence of known gene/molecule interactions in the Ingenuity Knowledge Base. The g:Profiler toolkit106 was used to analyze the proteins in each dementia-associated protein network for functional enrichment. We defined the biological pathways/processes associated with each protein set using the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and WikiPathways databases. Images were generated using the g:Profiler web tool: https://biit.cs.ut.ee/gprofiler/. Abbreviations: GO:BP, gene ontology biological process; GO:MF, gene ontology molecular function.
Extended Data Fig. 6 Functional profiling of protein Network 4, Network 5, and Network 6 identified among the set of dementia-associated proteins.
Protein networks were assembled based on evidence of known gene/molecule interactions in the Ingenuity Knowledge Base. The g:Profiler toolkit106 was used to analyze the proteins in each dementia-associated protein network for functional enrichment. We defined the biological pathways/processes associated with each protein set using the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and WikiPathways databases. Images were generated using the g:Profiler web tool: https://biit.cs.ut.ee/gprofiler/. Abbreviations: GO:BP, gene ontology biological process; GO:MF, gene ontology molecular function.
Extended Data Fig. 7 Functional profiling of protein Network 7, Network 8, and Network 10 identified among the set of dementia-associated proteins.
Protein networks were assembled based on evidence of known gene/molecule interactions in the Ingenuity Knowledge Base. The g:Profiler toolkit106 was used to analyze the proteins in each dementia-associated protein network for functional enrichment. We defined the biological pathways/processes associated with each protein set using the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and WikiPathways databases. Images were generated using the g:Profiler web tool: https://biit.cs.ut.ee/gprofiler/. No enriched biological pathways/processes were found for Network 9. Abbreviations: GO:BP, gene ontology biological process; GO:MF, gene ontology molecular function.
Extended Data Fig. 8 Functional profiling of protein Network 11 and Network 12 identified among the set of dementia-associated proteins.
Protein networks were assembled based on evidence of known gene/molecule interactions in the Ingenuity Knowledge Base. The g:Profiler toolkit106 was used to analyze the proteins in each dementia-associated protein network for functional enrichment. We defined the biological pathways/processes associated with each protein set using the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and WikiPathways databases. Images were generated using the g:Profiler web tool: https://biit.cs.ut.ee/gprofiler/. Abbreviations: GO:BP, gene ontology biological process; GO:MF, gene ontology molecular function.
Extended Data Fig. 9 PEER factors associated with incident dementia after visit 5.
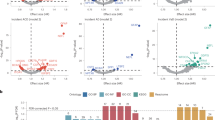
a, Volcano plot showing the hazard ratio (x-axis) and two-sided P-value (y-axis) for the association of 110 PEER factors with incident dementia. PEER factors above the horizontal dotted line were significantly associated with incident dementia after Bonferroni correction (0.05/110; P < 0.00045). P-values for dementia-associated PEER factors were 1.52E-06, 2.24E-05, and 2.00E-04 for PEER factors 9, 88, and 3, respectively. b, Spearman correlations between dementia-associated protein level and each PEER factors associated with dementia risk.
Supplementary information
Supplementary Information
Supplementary Methods and Figs. 1 and 2.
Supplementary Tables
Supplementary Tables 1–31
Rights and permissions
About this article
Cite this article
Walker, K.A., Chen, J., Zhang, J. et al. Large-scale plasma proteomic analysis identifies proteins and pathways associated with dementia risk. Nat Aging 1, 473–489 (2021). https://doi.org/10.1038/s43587-021-00064-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00064-0
This article is cited by
-
Associations between genetically predicted plasma protein levels and Alzheimer’s disease risk: a study using genetic prediction models
Alzheimer's Research & Therapy (2024)
-
Integration of polygenic and gut metagenomic risk prediction for common diseases
Nature Aging (2024)
-
Blood protein markers predict 15-year risk of dementia
Nature Aging (2024)
-
Plasma proteomic profiles predict future dementia in healthy adults
Nature Aging (2024)
-
Associations of plasma proteomics and age-related outcomes with brain age in a diverse cohort
GeroScience (2024)