Abstract
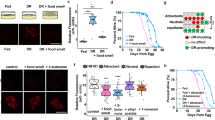
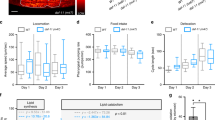
The role of food nutrients in mediating the positive effect of dietary restriction (DR) on longevity has been extensively characterized, but how non-nutrient food components regulate lifespan is not well understood. Here, we show that food-associated odors shorten the lifespan of Caenorhabditis elegans under DR but not those fed ad libitum, revealing a specific effect of food odors on DR-mediated longevity. Food odors act on a neural circuit comprising the sensory neurons ADF and CEP, and the interneuron RIC. This olfactory circuit signals the gut to suppress DR-mediated longevity via octopamine, the invertebrate homolog of norepinephrine, by regulating the energy sensor AMP-activated protein kinase (AMPK) through a Gq-phospholipase Cβ-CaMKK-dependent mechanism. In mouse primary cells, we find that norepinephrine signaling regulates AMPK through a similar mechanism. Our results identify a brain–gut axis that regulates DR-mediated longevity by relaying olfactory information about food abundance from the brain to the gut.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated and analyzed during this study are either included within the manuscript or are available from the corresponding author on reasonable request.
Change history
13 March 2023
In the version of this article initially published, there was a typo in the sentence of the abstract now reading, in part, "This olfactory circuit signals the gut to suppress DR-mediated longevity via octopamine, the invertebrate homolog of norepinephrine," where "invertebrate" replaced "mammalian" in the HTML and PDF versions of the article.
References
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512 (2010).
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span—from yeast to humans. Science 328, 321–326 (2010).
Greer, E. L., Banko, M. R. & Brunet, A. AMP-activated protein kinase and FoxO transcription factors in dietary restriction-induced longevity. Ann. N. Y. Acad. Sci. 1170, 688–692 (2009).
Kapahi, P., Kaeberlein, M. & Hansen, M. Dietary restriction and life span: lessons from invertebrate models. Ageing Res. Rev. 39, 3–14 (2017).
Gendron, C. M. et al. Neuronal mechanisms that drive organismal aging through the lens of perception. Annu. Rev. Physiol. 82, 227–249 (2020).
Alcedo, J. & Kenyon, C. Regulation of C. elegans longevity by specific gustatory and olfactory neurons. Neuron 41, 45–55 (2004).
Apfeld, J. & Kenyon, C. Regulation of life span by sensory perception in Caenorhabditis elegans. Nature 402, 804–809 (1999).
Libert, S. et al. Regulation of Drosophila life span by olfaction and food-derived odors. Science 315, 1133–1137 (2007).
Greer, E. L. & Brunet, A. Different dietary restriction regimens extend life span by both independent and overlapping genetic pathways in C. elegans. Aging Cell 8, 113–127 (2009).
Greer, E. L. et al. An AMPK–FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans. Curr. Biol. 17, 1646–1656 (2007).
Smith, E. D. et al. Age- and calorie-independent life span extension from dietary restriction by bacterial deprivation in Caenorhabditis elegans. BMC Dev. Biol. 8, 49 (2008).
Artan, M. et al. Food-derived sensory cues modulate longevity via distinct neuroendocrine insulin-like peptides. Genes Dev. 30, 1047–1057 (2016).
Bargmann, C. I. Chemosensation in C. elegans. WormBook 25, 1–29 (2006).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R. Soc. Lond. B Biol. Sci. 314, 1–340 (1986).
Kang, L., Gao, J., Schafer, W. R., Xie, Z. & Xu, X. Z. S. C. elegans TRP family protein TRP-4 is a pore-forming subunit of a native mechanotransduction channel. Neuron 67, 381–391 (2010).
Sawin, E. R., Ranganathan, R. & Horvitz, H. R. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron 26, 619–631 (2000).
Li, W., Feng, Z., Sternberg, P. W. & Xu, X. Z. S. A C. elegans stretch receptor neuron revealed by a mechanosensitive TRP channel homologue. Nature 440, 684–687 (2006).
Alkema, M. J., Hunter-Ensor, M., Ringstad, N. & Horvitz, H. R. Tyramine functions independently of octopamine in the Caenorhabditis elegans nervous system. Neuron 46, 247–260 (2005).
Shao, J. et al. Serotonergic neuron ADF modulates avoidance behaviors by inhibiting sensory neurons in C. elegans. Pflugers Arch. 471, 357–363 (2019).
Richmond, J. E., Davis, W. S. & Jorgensen, E. M. UNC-13 is required for synaptic vesicle fusion in C. elegans. Nat. Neurosci. 2, 959–964 (1999).
Speese, S. et al. UNC-31 (CAPS) is required for dense-core vesicle but not synaptic vesicle exocytosis in Caenorhabditis elegans. J. Neurosci. 27, 6150–6162 (2007).
Link, E. et al. Tetanus toxin action: inhibition of neurotransmitter release linked to synaptobrevin proteolysis. Biochem. Biophys. Res. Commun. 189, 1017–1023 (1992).
Hendricks, M., Ha, H., Maffey, N. & Zhang, Y. Compartmentalized calcium dynamics in a C. elegans interneuron encode head movement. Nature 487, 99–103 (2012).
Li, Z., Liu, J., Zheng, M. & Xu, X. Z. Encoding of both analog- and digital-like behavioral outputs by one C. elegans interneuron. Cell 159, 751–765 (2014).
Sokolchik, I., Tanabe, T., Baldi, P. F. & Sze, J. Y. Polymodal sensory function of the Caenorhabditis elegans OCR-2 channel arises from distinct intrinsic determinants within the protein and is selectively conserved in mammalian TRPV proteins. J. Neurosci. 25, 1015–1023 (2005).
Hobert, O. in WormBook (Ed. The C. elegans Research Community) 1–106 (WormBook, 2013).
Blackwell, T. K., Sewell, A. K., Wu, Z. & Han, M. TOR signaling in Caenorhabditis elegans development, metabolism and aging. Genetics 213, 329–360 (2019).
Winston, W. M., Molodowitch, C. & Hunter, C. P. Systemic RNAi in C. elegans requires the putative transmembrane protein SID-1. Science 295, 2456–2459 (2002).
Rera, M., Clark, R. I. & Walker, D. W. Intestinal barrier dysfunction links metabolic and inflammatory markers of aging to death in Drosophila. Proc. Natl Acad. Sci. USA 109, 21528–21533 (2012).
Gelino, S. et al. Intestinal autophagy improves healthspan and longevity in C. elegans during dietary Restriction. PLoS Genet. 12, e1006135 (2016).
Dambroise, E. et al. Two phases of aging separated by the Smurf transition as a public path to death. Sci. Rep. 6, 23523 (2016).
Yoshida, M., Oami, E., Wang, M., Ishiura, S. & Suo, S. Nonredundant function of two highly homologous octopamine receptors in food-deprivation-mediated signaling in Caenorhabditis elegans. J. Neurosci. Res. 92, 671–678 (2014).
Burkewitz, K., Zhang, Y. & Mair, W. B. AMPK at the nexus of energetics and aging. Cell Metab. 20, 10–25 (2014).
Neves, S. R., Ram, P. T. & Iyengar, R. G protein pathways. Science 296, 1636–1639 (2002).
Dimov, I. & Maduro, M. F. The C. elegans intestine: organogenesis, digestion and physiology. Cell Tissue Res. 377, 383–396 (2019).
Lafontan, M. & Berlan, M. Fat cell adrenergic receptors and the control of white and brown fat cell function. J. Lipid Res. 34, 1057–1091 (1993).
Graham, R. M., Perez, D. M., Hwa, J. & Piascik, M. T. alpha 1-adrenergic receptor subtypes. Molecular structure, function and signaling. Circ. Res. 78, 737–749 (1996).
Finger, F. et al. Olfaction regulates organismal proteostasis and longevity via microRNA-dependent signalling. Nat. Metab. 1, 350–359 (2019).
Albrecht, J. et al. Olfactory detection thresholds and pleasantness of a food-related and a non-food odour in hunger and satiety. Rhinology 47, 160–165 (2009).
Burkewitz, K. et al. Neuronal CRTC-1 governs systemic mitochondrial metabolism and life span via a catecholamine signal. Cell 160, 842–855 (2015).
Greer, E. L. et al. The energy sensor AMP-activated protein kinase directly regulates the mammalian FOXO3 transcription factor. J. Biol. Chem. 282, 30107–30119 (2007).
Weir, H. J. et al. Dietary restriction and AMPK increase life span via mitochondrial network and peroxisome remodeling. Cell Metab. 26, 884–896 (2017).
Riera, C. E. et al. The sense of smell impacts metabolic health and obesity. Cell Metab. 26, 198–211 (2017).
Piggott, B. J., Liu, J., Feng, Z., Wescott, S. A. & Xu, X. Z. S. The neural circuits and synaptic mechanisms underlying motor initiation in C. elegans. Cell 147, 922–933 (2011).
Ching, T. T. & Hsu, A. L. Solid plate-based dietary restriction in Caenorhabditis elegans. J. Vis. Exp. https://doi.org/10.3791/2701 (2011).
Zhang, B. et al. Brain–gut communications via distinct neuroendocrine signals bidirectionally regulate longevity in C. elegans. Genes Dev. 32, 258–270 (2018).
Wang, X., Li, G., Liu, J., Liu, J. & Xu, X. Z. S. TMC-1 mediates alkaline sensation in C. elegans through nociceptive neurons. Neuron 91, 146–154 (2016).
Jun, H. et al. An immune-beige adipocyte communication via nicotinic acetylcholine receptor signaling. Nat. Med. 24, 814–822 (2018).
Qiao, X. et al. Protein arginine methyltransferase 1 interacts with pgc1α and modulates thermogenic fat activation. Endocrinology 160, 2773–2786 (2019).
Hwang, A. B. et al. Feedback regulation via AMPK and HIF-1 mediates ROS-dependent longevity in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 111, E4458–E4467 (2014).
Xiao, R. et al. A genetic program promotes C. elegans longevity at cold temperatures via a thermosensitive TRP channel. Cell 152, 806–817 (2013).
Zhang, B. et al. Environmental temperature differentially modulates C. elegans longevity through a thermosensitive TRP channel. Cell Rep. 11, 1414–1424 (2015).
Acknowledgements
We thank J. Gong, S. Zhang and H. Chen for assistance. Some strains were obtained from the Caenorhabditis Genetics Center. J.L. received funding support from the National Natural Science Foundation of China (81872945 and 81720108031). J.W. received funding support from the National Institute of Diabetes and Digestive and Kidney Diseases (R01DK107583). X.Z.S.X. received funding support from the National Institute of General Medical Sciences (R35GM126917).
Author information
Authors and Affiliations
Contributions
B.Z. conducted the experiments and analyzed the data. H.J., B.Z. and J.W. performed mouse experiments and analyzed the data. B.Z., J.L. and X.Z.S.X. wrote the paper with assistance from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks Andrew Dillin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional data related to food odor suppression of DR longevity without increasing food ingestion.
a, b, Food odors do not change the pumping rate of worms under DR. Pumping rate was counted at 24 hr (a), and 96 hr (b), after worms were transferred to the DR assaying plates. 2 × DR: twice amount of bacterial food, which stimulated the pumping rate. n = 30 (DR), 30 (DR + odor), 16 (2xDR) and 16 (2xDR + odor) biologically independent animals in (a). n = 30 (DR), 30 (DR + odor), 15 (2xDR) and 14 (2xDR + odor) biologically independent animals in (b). Data are presented as mean ± s.e.m. P values were calculated with one-way ANOVA with Bonfferroni’s test. See Fig. 1c for data of the pumping rate counted at 1 hr post-transfer. c, Feeding worms with twice amount of bacteria food (2x DR) does not affect DR longevity, but food odors can still suppress the lifespan of these worms.
Extended Data Fig. 2 Additional data related to the olfactory circuit.
a, ADF dendrite, soma and axon from DR, but not AL worms, all respond robustly to medium containing food odors. GCaMP6f was expressed as a transgene in ADF neurons under tph-1(L) promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). b, NSM neurons (dendrite/soma/axon) from DR and AL worms do not respond to medium containing food odors. GCaMP6f was expressed as a transgene in the NSM neurons using tph-1(s) promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). c, Bar graph summarizing data in (a) and (b). n = 11 (ADF dendrite - AL), 10 (ADF dendrite - DR), 8 (ADF soma - AL), 9 (ADF soma - DR), 11 (ADF axon - AL), 10 (ADF axon - DR), 11 (NSM soma - AL), 14 (NSM soma - DR), 13 (NSM processes - AL), and 10 (NSM processes - DR) biologically independent animals. d, CEP dendrite from DR worms, but not AL worms, responds robustly to medium containing food odors. GCaMP6f was expressed as a transgene in CEP neurons using dat-1 promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). e, CEP soma and axon from both DR and AL worms respond to medium containing food odors, showing no specificity towards DR. Shades along the traces represent error bars (SEM). f, Bar graph summarizing data in (d) and (e). n = 6 (CEP dendrite - AL), 8 (CEP dendrite - DR), 13 (CEP soma - AL), 20 (CEP soma - DR), 12 (CEP axon - AL) and 17 (CEP axon - DR) biologically independent animals. g, h, ADE and PDE neurons (soma and processes) from DR and AL worms do not respond to medium containing food odors. GCaMP6f was expressed as a transgene in ADE and PDE neurons using dat-1 promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). i, Bar graph summarizing data in (g) and (h). n = 11 (ADE soma - AL), 10 (ADE soma - DR), 13 (ADE processes - AL), 15 (ADE processes - DR), 10 (PDE soma - AL), 11 (PDE soma - DR), 10 (PDE processes - AL) and 11 (PDE processes - DR) biologically independent animals. j, RIC soma and processes from DR worms, but not AL worms, respond robustly to medium containing food odors. GCaMP6f was expressed as a transgene in RIC neurons under tbh-1 promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). The soma traces are duplicates of those presented in Fig. 2i, as these experiments were performed at the same time. k, RIM soma and axon from both DR and AL worms respond to medium containing food odors, showing no specificity towards DR. GCaMP6f was expressed as a transgene in RIM neurons using cex-1 promoter. DsRed was co-expressed to enable ratiometric imaging. Shades along the traces represent error bars (SEM). l, Bar graph summarizing data in (j) and (k). n = 11 (RIC soma - AL), 14 (RIC soma - DR), 11 (RIC processes - AL), 12 (RIC processes - DR), 12 (RIM soma - AL), 12 (RIM soma - DR), 12 (RIM processes - AL) and 10 (RIM processes - DR) biologically independent animals. m, n, RNAi knockdown of odr-3 and ocr-2 specifically in ADF neurons eliminates food odor-evoked calcium responses in these neurons. dsRNA against odr-3 and ocr-2 gene was expressed as a transgene specifically in ADF neurons using the bas-1(prom7) promoter. (m) Calcium imaging traces. Shades along the traces represent error bars (SEM). (n) Bar graph summarizing the data in (m). n = 10 (WT - AL), 9 (WT - DR), 10 (ADF odr-3(RNAi) - AL), 10 (ADF odr-3(RNAi) - DR), 10 (ADF ocr-2(RNAi) - AL) and 10 (ADF ocr-2(RNAi) - DR) biologically independent animals. (o) NSM neurons are not required for food odors to suppress DR longevity. tph-1(s) promoter was used to drive the expression of TeTx transgene specifically in NSM neurons. (p) Blocking the output of RIC neuron shortens DR longevity. tbh-1 promoter was used to drive the expression of TeTx transgene in RIC neurons. Data are presented as mean ± s.e.m. P values in c, f, i and l: two-tailed student’s t test. P values in n: one-way ANOVA with Bonfferroni’s test.
Extended Data Fig. 3 Other serotonin receptors are not required for food odors to suppress DR longevity.
Food odors can still suppress DR longevity in ser-1 (a), ser-4 (b), ser-7 (c) and mod-1 (d) mutant worms. a-d, share the same control group, as these experiments were performed at the same time.
Extended Data Fig. 4 Other dopamine receptors are not required for food odors to suppress DR longevity.
Food odors can still suppress DR longevity in dop-1 (a) dop-2 (b) dop-3 (c) dop-4 (d) and dop-5 (e) mutant worms. (a) and (c) share the same control group, as these experiments were performed at the same time. (b), (d) and (e) share the same control group, as these experiments were performed at the same time.
Extended Data Fig. 5 Additional data related to regulation of DR longevity by AMPK and octopamine signaling.
a, DR can extend the lifespan of raga-1 mutant worms. b, Food odors can suppress DR longevity in raga-1 mutant worms. c, DR can extend the lifespan of rict-1 mutant worms. d, Food odors can suppress DR longevity in rict-1 mutant worms. e, Pan-neuronal expression of aak-2 gene only has a slight rescue effect on the longevity defect of aak-2 mutant worms. This aak-2 neuronal transgene also does not rescue the odor sensitivity defect of aak-2 mutant worms. rgef-1 promoter was used to drive the expression of aak-2 cDNA in neurons. f-g, Food odors can still suppress DR longevity in ser-6 (f) and octr-1 (g) mutant worms. (f) and (g) share the same control group, as these experiments were performed at the same time. h, Intestine-specific knock-down of par-4/LKB1 by dsRNA transgene (Pges-1::par-4(RNAi)) does not prevent food odors from suppressing DR longevity, though it partially inhibits DR longevity. i, Intestine-specific knock-down of mom-4/TAK1 by dsRNA transgene (Pges-1::mom-4(RNAi)) does not prevent food odors from suppressing DR longevity; nor does it affect DR longevity. j–l, Mutations in egl-30 (j), egl-8 (k), and ckk-1 (l) abolish the ability of food odors to suppress DR longevity, a defect that is rescued by transgenic expression of corresponding wild-type genes in the intestine using ges-1 promoter.
Extended Data Fig. 6 AAK-2/AMPK-dependent lifespan extension requires DAF-16/FOXO.
a, Lifespan extension mediated by intestinal expression of aak-2 requires daf-16. daf-16 RNAi blocked the lifespan-extension effect of the intestinal aak-2 transgene. b-d, Intestinal expression of aak-2 promotes sod-3 gene expression in multiple tissues in a daf-16-dependent manner. sod-3::gfp is a transgene reporting the expression level of sod-3 gene. (b) Sample images showing a low level of sod-3::gfp expression. Left: bright field image. Right: fluorescent image. (c) Sample images showing that the Pges-1::aak-2 transgene increased the expression of sod-3::gfp. Top left: bright field image. Top right: fluorescent image. Bottom: zoomed-in images showing sod-3::gfp expression in multiple tissues, including pharynx (head), neurons (head), body-wall muscles, vulval muscles (mid-body), intestine, etc. Scale Bar: 100 μm. (d) Bar graph summarizing the data in (b) and (c). n = 24 (WT), 20 (Pges-1::aak-2), 43 (daf-16(RNAi)) and 22 (daf-16(RNAi); Pges-1::aak-2) biologically independent animals. Data are presented as mean ± s.e.m. P values were calculated with one-way ANOVA with Bonfferroni’s test.
Supplementary information
Supplementary Table 1
Lifespan statistics related to Figs. 1–5.
Source data
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, B., Jun, H., Wu, J. et al. Olfactory perception of food abundance regulates dietary restriction-mediated longevity via a brain-to-gut signal. Nat Aging 1, 255–268 (2021). https://doi.org/10.1038/s43587-021-00039-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00039-1
This article is cited by
-
BPIFB4 and its longevity-associated haplotype protect from cardiac ischemia in humans and mice
Cell Death & Disease (2023)
-
Age-associated decline in RAB-10 efficacy impairs intestinal barrier integrity
Nature Aging (2023)
-
Olfactory chemosensation extends lifespan through TGF-β signaling and UPR activation
Nature Aging (2023)
-
Dopamine D1- and D2-like receptors oppositely regulate lifespan via a dietary restriction mechanism in Caenorhabditis elegans
BMC Biology (2022)
-
Serotonin and dopamine modulate aging in response to food odor and availability
Nature Communications (2022)