Abstract
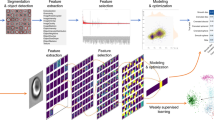
Imaging flow cytometry combines the high-event-rate nature of flow cytometry with the advantages of single-cell image acquisition associated with microscopy. The measurement of large numbers of features from the resulting images provides rich data sets that have resulted in a wide range of novel biomedical applications. In this Primer, we discuss the typical imaging flow instrumentation, the form of data acquired and the typical analysis tools that can be applied to these data. Focusing on the first commercially available imaging flow cytometer, the ImageStream (Luminex), we use examples from the literature to discuss the progression of the analysis methods used in imaging flow cytometry. These methods start from the use of simple single-image features and multiple channel gating strategies, followed by the design and use of custom features for phenotype classification, through to powerful machine and deep-learning methods. For each of these methods, we outline the processes involved in analysing typical data sets and provide details of example applications. Finally, we discuss the current limitations of imaging flow cytometry and the innovations and new instruments that are addressing these challenges.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 1 digital issues and online access to articles
$99.00 per year
only $99.00 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brown, M. & Wittwer, C. Flow cytometry: principles and clinical applications in hematology. Clin. Chem. 46, 1221–1229 (2000).
De Rosa, S. C., Brenchley, J. M. & Roederer, M. Beyond six colors: a new era in flow cytometry. Nat. Med. 9, 112–117 (2003).
Basiji, D. A., Ortyn, W. E., Liang, L., Venkatachalam, V. & Morrissey, P. Cellular image analysis and imaging by flow cytometry. Clin. Lab. Med. 27, 653–670 (2007). Introduction to ImageStream.
Maguire, O., Collins, C., O’Loughlin, K., Miecznikowski, J. & Minderman, H. Quantifying nuclear p65 as a parameter for NF-κB activation: correlation between ImageStream cytometry, microscopy, and Western blot. Cytometry A 79, 461–469 (2011).
Kaur, M. & Esau, L. Two-step protocol for preparing adherent cells for high-throughput flow cytometry. Biotechniques 59, 119–126 (2015).
Stavrakis, S., Holzner, G., Choo, J. & deMello, A. High-throughput microfluidic imaging flow cytometry. Curr. Opin. Biotechnol. 55, 36–43 (2019).
Holzner, G. et al. High-throughput multiparametric imaging flow cytometry: toward diffraction-limited sub-cellular detection and monitoring of sub-cellular processes. Cell Rep. 34, 108824 (2021).
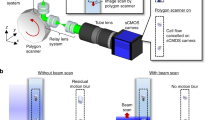
Mikami, H. et al. Virtual-freezing fluorescence imaging flow cytometry. Nat. Commun. 11, 1162 (2020).
Diebold, E. D. et al. Digitally synthesized beat frequency multiplexing for sub-millisecond fluorescence microscopy. Nat. Photonics 7, 806–810 (2013).
George, T. C. et al. Distinguishing modes of cell death using the ImageStream multispectral imaging flow cytometer. Cytometry A 59, 237–245 (2004).
Ortyn, W. E. et al. Extended depth of field imaging for high speed cell analysis. Cytometry A 71, 215–231 (2007).
Haley Renee, P. & Raymond, K. K. Demonstration of high gain mode in combination with imaging flow cytometry for improved EV analysis. Mian Yi Xue Za Zhi 204, 15 (2020).
Vogt, R. F. Jr., Whitfield, W. E., Henderson, L. O. & Hannon, W. H. Fluorescence intensity calibration for immunophenotyping by flow cytometry. Methods 21, 289–296 (2000).
Ortyn, W. E. et al. Sensitivity measurement and compensation in spectral imaging. Cytometry A 69, 852–862 (2006). Compensation of imaging flow cytometry data.
Fazekas de St Groth, B., Zhu, E., Asad, S. & Lee, L. Flow cytometric detection of human regulatory T cells. Methods Mol. Biol. 707, 263–279 (2011).
Holmberg-Thyden, S., Grønbæk, K., Gang, A. O., El Fassi, D. & Hadrup, S. R. A user’s guide to multicolor flow cytometry panels for comprehensive immune profiling. Anal. Biochem. 627, 114210 (2021).
Dominical, V., Samsel, L. & McCoy, J. P. Jr. Masks in imaging flow cytometry. Methods 112, 917 (2017). Discussion of masks in imaging flow cytometry.
Barteneva, N. S. & Vorobjev, I. A. Imaging Flow Cytometry: Methods and Protocols (Humana Press, 2015).
Patterson, J. O., Swaffer, M. & Filby, A. An imaging flow cytometry-based approach to analyse the fission yeast cell cycle in fixed cells. Methods 82, 74–84 (2015).
Patterson, J. O., Basu, S., Rees, P. & Nurse, P. CDK control pathways integrate cell size and ploidy information to control cell division. eLife 10, e64592 (2021).
Patterson, J. O., Rees, P. & Nurse, P. Noisy cell-size-correlated expression of cyclin B drives probabilistic cell-size homeostasis in fission yeast. Curr. Biol. 29, 1379–1386.e1374 (2019).
Calvert, M. E. K., Lannigan, J. A. & Pemberton, L. F. Optimization of yeast cell cycle analysis and morphological characterization by multispectral imaging flow cytometry. Cytometry A 73, 825–833 (2008). Cell cycle analysis using imaging flow cytometry using image properties.
Filby, A. et al. An imaging flow cytometric method for measuring cell division history and molecular symmetry during mitosis. Cytometry A 79, 496–506 (2011).
Summers, H. D. et al. Statistical analysis of nanoparticle dosing in a dynamic cellular system. Nat. Nanotechnol. 6, 170–174 (2011).
Bourton, E. C. et al. Multispectral imaging flow cytometry reveals distinct frequencies of γ-H2AX foci induction in DNA double strand break repair defective human cell lines. Cytometry A 81, 130–137 (2012).
Jurgielewicz, B. J., Yao, Y. & Stice, S. L. Kinetics and specificity of HEK293T extracellular vesicle uptake using imaging flow cytometry. Nanoscale Res. Lett. 15, 170 (2020).
Fei, C., Lillico, D. M. E., Hall, B., Rieger, A. M. & Stafford, J. L. Connected component masking accurately identifies the ratio of phagocytosed and cell-bound particles in individual cells by imaging flow cytometry. Cytometry A 91, 372–381 (2017).
Vranic, S. et al. Deciphering the mechanisms of cellular uptake of engineered nanoparticles by accurate evaluation of internalization using imaging flow cytometry. Part. Fibre Toxicol. 10, 2 (2013).
Chivukula, R. R. et al. A human ciliopathy reveals essential functions for NEK10 in airway mucociliary clearance. Nat. Med. 26, 244–251 (2020).
Blasi, T. et al. Label-free cell cycle analysis for high-throughput imaging flow cytometry. Nat. Commun. 7, 10256 (2016). Application of machine learning to imaging flow data.
Nassar, M. et al. Label-free identification of white blood cells using machine learning. Cytometry A 95, 836–842 (2019).
Lippeveld, M. et al. Classification of human white blood cells using machine learning for stain-free imaging flow cytometry. Cytometry A 97, 308–319 (2020).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Hennig, H. et al. An open-source solution for advanced imaging flow cytometry data analysis using machine learning. Methods 112, 201–210 (2017).
Ding, C. & Peng, H. Minimum redundancy feature selection from microarray gene expression data. in Computational Systems Bioinformatics. CSB2003. Proceedings of the 2003 IEEE Bioinformatics Conference (IEEE, 2003).
Mandal, M. & Mukhopadhyay, A. An improved minimum redundancy maximum relevance approach for feature selection in gene expression data. Proc. Technol. 10, 20–27 (2013).
Rees, P., Wills, J. W., Brown, M. R., Barnes, C. M. & Summers, H. D. The origin of heterogeneous nanoparticle uptake by cells. Nat. Commun. 10, 2341 (2019).
Cerveira, J., Begum, J., Di Marco Barros, R., van der Veen, A. G. & Filby, A. An imaging flow cytometry-based approach to measuring the spatiotemporal calcium mobilisation in activated T cells. J. Immunol. Methods 423, 120–130 (2015).
Piasecka, J. et al. Diffusion mapping of eosinophil-activation state. Cytometry A 97, 253–258 (2019).
Eulenberg, P. et al. Reconstructing cell cycle and disease progression using deep learning. Nat. Commun. 8, 463 (2017). Application of deep learning to imaging flow data.
Dunker, S., Boho, D., Wäldchen, J. & Mäder, P. Combining high-throughput imaging flow cytometry and deep learning for efficient species and life-cycle stage identification of phytoplankton. BMC Ecol. 18, 51 (2018).
Dunker, S. et al. Pollen analysis using multispectral imaging flow cytometry and deep learning. New Phytol. 229, 593–606 (2021).
Luo, S. et al. Deep learning-enabled imaging flow cytometry for high-speed Cryptosporidium and Giardia detection. Cytometry A 99, 1123–1133 (2021).
Rodrigues, M. A., Beaton-Green, L. A., Kutzner, B. C. & Wilkins, R. C. Automated analysis of the cytokinesis-block micronucleus assay for radiation biodosimetry using imaging flow cytometry. Radiat. Environ. Biophys. 53, 273–282 (2014).
Rodrigues, M. A., Probst, C. E., Beaton-Green, L. A. & Wilkins, R. C. Optimized automated data analysis for the cytokinesis-block micronucleus assay using imaging flow cytometry for high throughput radiation biodosimetry. Cytometry A 89, 653–662 (2016).
Verma, J. R. et al. Evaluation of the automated MicroFlow® and MetaferTM platforms for high-throughput micronucleus scoring and dose response analysis in human lymphoblastoid TK6 cells. Arch. Toxicol. 91, 2689–2698 (2017).
Rodrigues, M. A. Automation of the in vitro micronucleus assay using the Imagestream imaging flow cytometer. Cytometry A 93, 706–726 (2018).
Rodrigues, M. A. et al. The in vitro micronucleus assay using imaging flow cytometry and deep learning. npj Syst. Biol. Appl. 7, 20 (2021).
Wills, J. W. et al. Inter-laboratory automation of the in vitro micronucleus assay using imaging flow cytometry and deep learning. Arch. Toxicol. 95, 3101–3115 (2021).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun. ACM 60, 84–90 (2017).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. in 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (IEEE, 2016).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. Preprint at https://doi.org/10.48550/arXiv.1409.1556 (2014).
Mochalova, E. N., Kotov, I. A., Lifanov, D. A., Chakraborti, S. & Nikitin, M. P. Imaging flow cytometry data analysis using convolutional neural network for quantitative investigation of phagocytosis. Biotechnol. Bioeng. 119, 626–635 (2022).
Probst, C., Zayats, A., Venkatachalam, V. & Davidson, B. Advanced characterization of silicone oil droplets in protein therapeutics using artificial intelligence analysis of imaging flow cytometry data. J. Pharm. Sci. 109, 2996–3005 (2020).
Spidlen, J., Breuer, K. & Brinkman, R. Preparing a minimum information about a flow cytometry experiment (MIFlowCyt) compliant manuscript using the International Society for Advancement of Cytometry (ISAC) FCS file repository (FlowRepository.org). Curr. Protoc. Cytom. 61, 10.18.1–10.18.26 (2012).
Filby, A. & Davies, D. Reporting imaging flow cytometry data for publication: why mask the detail? Cytometry A 81, 637–642 (2012).
Caicedo, J. C. et al. Data-analysis strategies for image-based cell profiling. Nat. Methods 14, 849–863 (2017).
Montero Llopis, P. et al. Best practices and tools for reporting reproducible fluorescence microscopy methods. Nat Methods 18, 1463–1476 (2021).
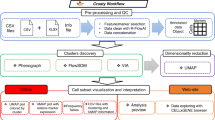
Doan, M. et al. Deepometry, a framework for applying supervised and weakly supervised deep learning to imaging cytometry. Nat. Protoc. 16, 3572–3595 (2021).
Demont, Y. Tools for Imaging Flow Cytometry [R package IFC version 0.1.2] (2021).
Schraivogel, D. et al. High-speed fluorescence image–enabled cell sorting. Science 375, 315320 (2022).
Gu, Y. et al. Machine learning based real-time image-guided cell sorting and classification. Cytometry A 95, 499–509 (2019).
Nitta, N. et al. Intelligent image-activated cell sorting. Cell 175, 266–276.e213 (2018). Image-based flow cytometry sorting.
Nawaz, A. A. et al. Intelligent image-based deformation-assisted cell sorting with molecular specificity. Nat. Methods 17, 595–599 (2020).
Merola, F. et al. Tomographic flow cytometry by digital holography. Light Sci. Appl. 6, e16241 (2017).
Kebschull, J. M. & Zador, A. M. Cellular barcoding: lineage tracing, screening and beyond. Nat. Methods 15, 871–879 (2018).
Rees, P. et al. Nanoparticle vesicle encoding for imaging and tracking cell populations. Nat. Methods 11, 1177–1181 (2014).
Han, Y. & Lo, W.-H. Imaging cells in flow cytometer using spatial-temporal transformation. Sci. Rep. 5, 13267 (2015).
Lee, K. C. M. et al. Quantitative phase imaging flow cytometry for ultra-large-scale single-cell biophysical phenotyping. Cytometry A 95, 510–520 (2019).
Ouk, C., Jayat-Vignoles, C., Donnard, M. & Feuillard, J. Both CD62 and CD162 antibodies prevent formation of CD36-dependent platelets, rosettes, and artefactual pseudoexpression of platelet markers on white blood cells: a study with ImageStream®. Cytometry A 79, 477–484 (2011).
Ugawa, M. et al. In silico-labeled ghost cytometry. eLife 10, e67660 (2021).
Rosenburg, C. A. Exploring dyserythropoiesis in patients with myelodysplasticsyndrome by imaging flow cytometry and machine-learning assisted morphometrics. Cytometry 100, 554–567 (2021).
Acknowledgements
P.R. and H.D.S. acknowledge the UK Engineering and Physical Sciences Research Council (EP/N013506/1) and UK Biotechnology and Biological Sciences Research Council (BB/P026818/1) for supporting this work. A.E.C. acknowledges the National Science Foundation (DBI 1458626) and the NIH (R35 GM122547) for supporting this work.
Author information
Authors and Affiliations
Contributions
Introduction (H.D.S., A.E.C., M.D. and P.R.); Experimentation (H.D.S., A.E.C. and P.R.); Results (H.D.S., A.F., A.E.C. and P.R.); Applications (H.D.S., A.F., A.E.C. and P.R.); Reproducibility and data deposition (H.D.S., A.F., A.E.C. and P.R.); Limitations and optimizations (H.D.S., A.E.C., M.D. and P.R.); Outlook (H.D.S., A.E.C., M.D. and P.R.).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Method Primers thanks Maik Herbig and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
FigShare: https://figshare.com/
FlowRepository: https://flowrepository.org/
GitHub: https://github.com/
Luminex: https://www.luminexcorp.com/eu/imaging-flow-cytometry/
Supplementary information
Glossary
- Brightfield
-
The simplest form of microscopy, in which the image is formed by white light that is transmitted through the sample and then captured on a detector.
- Darkfield
-
In the context of imaging flow cytometry, the darkfield image is formed when light scattered from the cell is collected on the detector perpendicular to the excitation direction.
- Gating
-
A range of bins for the histogram or a polygon for the scatter plot. This process selects cells for further analysis. The gating process can be repeated to define phenotypes that require more than two markers for identification.
- Multi-spectral images
-
An image data set in which the same field of view is imaged in different spectral bands.
- Masks
-
A mask is a binary image that defines the extent of the object in an image; the pixel values in the image are 1 inside the object perimeter and 0 elsewhere to represent the background.
- Raw maximum pixel
-
A feature in the ImageStream Data Exploration and Analysis Software that returns the maximum pixel value in an image acquired by the detector before any compensation. This is often used to set the laser excitation intensity to ensure that the pixel values are not saturated.
- Building block
-
Suggested feature scatter plots and gating strategies to help the user with simple analysis and preprocessing tasks, such as determining in-focus cells in the ImageStream Data Exploration and Analysis software.
- Aspect ratio
-
The ratio of the minor axis and the major axis. The major axis is the longest line that can be drawn through the shape, and the minor axis is the shortest line that can be drawn through the shape at right angles to the major axis.
- AND mask operation
-
The AND operator applied to two masks delivers the overlapped shared area between the masks.
- NOT mask operation
-
The NOT operator is a logic operator that delivers the inverse of a mask, that is, 0s become 1s.
- Confusion matrix
-
A confusion matrix is used to compare the predicted outcome of a machine-learning algorithm with the known classes of the data. Diagonal elements represent the number of correct classifications. Off-diagonal elements can be used to assess misclassifications.
- t-Distributed stochastic neighbour embedding
-
An algorithm used to visualize high-dimensional data sets in two or three dimensions. Nonlinear dimensional reduction of the data to the 2D–3D coordinate system is used to preserve the distances between similar and dissimilar data points.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rees, P., Summers, H.D., Filby, A. et al. Imaging flow cytometry. Nat Rev Methods Primers 2, 86 (2022). https://doi.org/10.1038/s43586-022-00167-x
Accepted:
Published:
DOI: https://doi.org/10.1038/s43586-022-00167-x
This article is cited by
-
Light-field flow cytometry for high-resolution, volumetric and multiparametric 3D single-cell analysis
Nature Communications (2024)
-
Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Nature Communications (2023)