Abstract
Therapy resistance represents a major clinical challenge in acute myeloid leukemia (AML). Here we define a ‘MitoScore’ signature, which identifies high mitochondrial oxidative phosphorylation in vivo and in patients with AML. Primary AML cells with cytarabine (AraC) resistance and a high MitoScore relied on mitochondrial Bcl2 and were highly sensitive to venetoclax (VEN) + AraC (but not to VEN + azacytidine). Single-cell transcriptomics of VEN + AraC-residual cell populations revealed adaptive resistance associated with changes in oxidative phosphorylation, electron transport chain complex and the TP53 pathway. Accordingly, treatment of VEN + AraC-resistant AML cells with electron transport chain complex inhibitors, pyruvate dehydrogenase inhibitors or mitochondrial ClpP protease agonists substantially delayed relapse following VEN + AraC. These findings highlight the central role of mitochondrial adaptation during AML therapy and provide a scientific rationale for alternating VEN + azacytidine with VEN + AraC in patients with a high MitoScore and to target mitochondrial metabolism to enhance the sensitivity of AML cells to currently approved therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
LC–MS and IC–MS data present in Fig. 2 have been deposited on MassIVE and are acessible through accession no. MSV000087892 (http://massive.ucsd.edu/ProteoSAFe/status.jsp?task=a2860bbc66924ead868954a0d030f364). RNA-seq data from TUH AML patients presented in Extended Data Fig. 1 and in Fig. 3 are accessible through Gene Expression Omnibus accession no. GSE183329. RNA-seq data from patients with AML presented in Fig. 4 are available through the European Genome–phenome Archive (EGAS00001003820). This dataset is a restrictive dataset from the study ‘Molecular patterns of response and treatment failure after frontline venetoclax combinations in older patients with AML’17. To access these data, contact I. Majewski (majewski@wehi.edu.au). Microarray data from CLDX MOLM14 presented in Fig. 6 are accessible through Gene Expression Omnibus accession no. GSE181792. The single-cell data discussed in Fig. 6 are accessible through Gene Expression Omnibus accession no. GSE178912. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The output bcl2 file was converted to FASTQ format by using CellRanger v.3.0.2 and then loaded in an R session with the Seurat 3.0 toolkit package86 involving the normalization and variance stabilization package sctransform87. For each PDX sample (TUH07 and TUH69), cells were filtered based on number of expressed genes (CTL, 300 < nFeatures < 5,500; AraC, 300 < nFeatures < 6,000; VEN, 300 < nFeatures < 6,500; AraC + VEN, 100 < nFeatures < 3,000). For TUH07 samples, percentage of mitochondrial reads were CTL, VEN, AraC, 0.04 < percent.mito < 0.15; AraC + VEN, 0.01 < percent.mito < 0.1. For TUH69 samples, percentage of mitochondrial reads were CTL, AraC, VEN, 0.02 < percent.mito < 0.15; AraC_VEN, 0.01 < percent.mito < 0.15. Single-Cell Signature Explorer was used for visualization of gene and signatures in the UMAP map88. Clustering was performed using Louvain algorithm implemented in Seurat, with a resolution of 0.1. Cluster markers were defined by Seurat package using a Wilcoxon statistical test. Markers with adjusted P value <0.05 were selected. GSEA for cluster markers were performed using the hypeR package89 using KEGG, Hallmark and GO databases. Area under the curve (AUC) scores for gene signatures were obtained using the R package AUCell, a method that uses the AUC to calculate whether a critical subset of the input gene set is enriched within the expressed genes for each cell42. Next, we ran SCENIC42 analysis, which uses gene regulatory network inference, followed by a refinement step using cis-regulatory information, to generate a set of regulons (transcription factors and their target genes) in the scRNA-seq data. Python implementation, pySCENIC (https://github.com/aertslab/pySCENIC, v.0.9.19) was run using a Nextflow pipeline (https://github.com/aertslab/SCENICprotocol, v.0.2.0)42, which streamlined the main steps of GRN inference and refinement with pySCENIC, as well as the quantification of cellular activity. For each regulon, SCENIC calculates an AUC score for each cell and then defines a binary activity (ON/OFF) by setting up an automatic threshold. SCENIC clustering and heat maps were based on matrix of binary regulon activity. Comparison of regulon activities between clusters were based on continuous AUC score and involved a Mann–Whitney U-test and Benjamini–Hochberg correction for multiple tests. The code used in this study has been deposited in Zenodo and is available at https://zenodo.org/record/5137701#.YP7P1zqxVH4. Single-cell RNA data were processed using cellranger v.3.0.2. (https://www.10XGenomics.com/) and analyzed with the R package Seurat v.3.0 (https://satijalab.org/seurat/). GSEA for cluster markers was performed using the hypeR package11 using KEGG, Hallmark and GO databases. Gene expression signatures were extracted using Single-Cell Signature Explorer (https://fredsoftwares/products/single-cell-signature-explorer). AUC score for genes signatures were obtained using the R package AUCell. pySCENIC (https://github.com/aertslab/pySCENIC, v.0.9.19), was run using a Nextflow pipeline (https://github.com/aertslab/SCENICprotocol, v.0.2.0).
References
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Short, N. J. et al. Advances in the treatment of acute myeloid leukemia: new drugs and new challenges. Cancer Discov. 10, 506–525 (2020).
Intlekofer, A. M. et al. Acquired resistance to IDH inhibition through trans or cis dimer-interface mutations. Nature 559, 125–129 (2018).
McMahon, C. M. et al. Clonal selection with RAS pathway activation mediates secondary clinical resistance to selective FLT3 inhibition in acute myeloid leukemia. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-18-1453 (2019).
Farge, T. et al. Chemotherapy-resistant human acute myeloid leukemia cells are not enriched for leukemic stem cells but require oxidative metabolism. Cancer Discov. 7, 716–735 (2017).
Henkenius, K. et al. Maintenance of cellular respiration indicates drug resistance in acute myeloid leukemia. Leuk. Res. 62, 56–63 (2017).
Kuntz, E. M. et al. Targeting mitochondrial oxidative phosphorylation eradicates therapy-resistant chronic myeloid leukemia stem cells. Nat. Med. 23, 1234–1240 (2017).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Bosc, C., Selak, M. A. & Sarry, J.-E. Resistance is futile: targeting mitochondrial energetics and metabolism to overcome drug resistance in cancer treatment. Cell Metab. 26, 705–707 (2017).
Andrzejewski, S. et al. PGC-1α promotes breast cancer metastasis and confers bioenergetic flexibility against metabolic drugs. Cell Metab. 26, e5 (2017).
Lee, K. et al. MYC and MCL1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab. 26, e7 (2017).
Samudio, I. et al. Pharmacologic inhibition of fatty acid oxidation sensitizes human leukemia cells to apoptosis induction. J. Clin. Invest. 120, 142–156 (2010).
Sánchez-Mendoza, S. E. & Rego, E. M. Targeting the mitochondria in acute myeloid leukemia. Appl. Cancer Res. 37, 22 (2017).
Lagadinou, E. D. et al. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 12, 329–341 (2013).
Pan, R. et al. Selective BCL-2 inhibition by ABT-199 causes on-target cell death in acute myeloid leukemia. Cancer Discov. 4, 362–375 (2014).
Konopleva, M. et al. Efficacy and biological correlates of response in a phase ii study of venetoclax monotherapy in patients with acute myelogenous leukemia. Cancer Discov. 6, 1106–1117 (2016).
DiNardo, C. D. et al. Molecular patterns of response and treatment failure after frontline venetoclax combinations in older patients with AML. Blood 135, 791–803 (2020).
Molina, J. R. et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 24, 1036–1046 (2018).
Boyd, A. L. et al. Identification of chemotherapy-induced leukemic-regenerating cells reveals a transient vulnerability of human AML recurrence. Cancer Cell 34, e5 (2018).
Vo, T.-T. et al. Relative mitochondrial priming of myeloblasts and normal HSCs determines chemotherapeutic success in AML. Cell 151, 344–355 (2012).
Chonghaile, T. N. et al. Pretreatment mitochondrial priming correlates with clinical response to cytotoxic chemotherapy. Science 334, 1129–1133 (2011).
Hosseini, M. et al. Targeting myeloperoxidase disrupts mitochondrial redox balance and overcomes cytarabine resistance in human acute myeloid leukemia. Cancer Res. 79, 5191–5203 (2019).
Halestrap, A. P. What is the mitochondrial permeability transition pore? J. Mol. Cell. Cardiol. 46, 821–831 (2009).
Rasola, A. & Bernardi, P. Mitochondrial permeability transition in Ca2+-dependent apoptosis and necrosis. Cell Calcium 50, 222–233 (2011).
Wacquier, B., Combettes, L. & Dupont, G. Dual dynamics of mitochondrial permeability transition pore opening. Sci. Rep. 10, 3924 (2020).
Cannino, G., Ciscato, F., Masgras, I., Sánchez-Martín, C. & Rasola, A. Metabolic plasticity of tumor cell mitochondria. Front. Oncol. 8, 333 (2018).
Griffiths, E. J. & Rutter, G. A. Mitochondrial calcium as a key regulator of mitochondrial ATP production in mammalian cells. Biochim. Biophys. Acta Bioenergy 1787, 1324–1333 (2009).
Bosc, C. et al. Autophagy regulates fatty acid availability for oxidative phosphorylation through mitochondria-endoplasmic reticulum contact sites. Nat. Commun. 11, 4056 (2020).
Tyner, J. W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526–531 (2018).
Jones, C. L. et al. Cysteine depletion targets leukemia stem cells through inhibition of electron transport complex II. Blood 134, 389–394 (2019).
Stuani, L. & Sarry, J.-E. Help from outside: cysteine to survive in AML. Blood 134, 336–338 (2019).
Pollyea, D. A. et al. Venetoclax with azacitidine disrupts energy metabolism and targets leukemia stem cells in patients with acute myeloid leukemia. Nat. Med. 24, 1859–1866 (2018).
Lucantoni, F., Düssmann, H., Llorente-Folch, I. & Prehn, J. H. M. BCL2 and BCL(X)L selective inhibitors decrease mitochondrial ATP production in breast cancer cells and are synthetically lethal when combined with 2-deoxy-D-glucose. Oncotarget 9, 26046–26063 (2018).
Guièze, R. et al. Mitochondrial reprogramming underlies resistance to BCL-2 inhibition in lymphoid malignancies. Cancer Cell 36, e13 (2019).
Sharon, D. et al. Inhibition of mitochondrial translation overcomes venetoclax resistance in AML through activation of the integrated stress response. Sci. Transl. Med. 11, eaax2863 (2019).
Jones, C. L. et al. Inhibition of amino acid metabolism selectively targets human leukemia stem cells. Cancer Cell 34, e4 (2018).
Pei, S. et al. Monocytic subclones confer resistance to venetoclax-based therapy in patients with acute myeloid leukemia. Cancer Discov. 10, 536–551 (2020).
Zhang, H. et al. Integrated analysis of patient samples identifies biomarkers for venetoclax efficacy and combination strategies in acute myeloid leukemia. Nat. Cancer 1, 826–839 (2020).
Stevens, B. M. et al. Fatty acid metabolism underlies venetoclax resistance in acute myeloid leukemia stem cells. Nat. Cancer https://doi.org/10.1038/s43018-020-00126-z (2020).
Ramsey, H. E. et al. A novel MCL1 inhibitor combined with venetoclax rescues venetoclax-resistant acute myelogenous leukemia. Cancer Discov. 8, 1566–1581 (2018).
Konopleva, M. et al. Mechanisms of apoptosis sensitivity and resistance to the BH3 mimetic ABT-737 in acute myeloid leukemia. Cancer Cell 10, 375–388 (2006).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Daniel, M. G., Pereira, C.-F., Lemischka, I. R. & Moore, K. A. Making a hematopoietic stem cell. Trends Cell Biol. 26, 202–214 (2016).
Bresnick, E. H., Katsumura, K. R., Lee, H.-Y., Johnson, K. D. & Perkins, A. S. Master regulatory GATA transcription factors: mechanistic principles and emerging links to hematologic malignancies. Nucleic Acids Res. 40, 5819–5831 (2012).
Fernandez-Mosquera, L. et al. Mitochondrial respiratory chain deficiency inhibits lysosomal hydrolysis. Autophagy 15, 1572–1591 (2019).
Abbate, F. et al. FBXW7 regulates a mitochondrial transcription program by modulating MITF. Pigment Cell Melanoma Res. 31, 636–640 (2018).
Shin, B. et al. Mitochondrial oxidative phosphorylation regulates the fate decision between pathogenic Th17 and regulatory T cells. Cell Rep. 30, 1898–1909 (2020).
Wei, J. et al. Targeting REGNASE-1 programs long-lived effector T cells for cancer therapy. Nature 576, 471–476 (2019).
Vervliet, T. et al. Modulation of Ca2+ signaling by anti-apoptotic B-cell lymphoma 2 proteins at the endoplasmic reticulum–mitochondrial interface. Front. Oncol. 7, 75 (2017).
Danese, A. et al. Calcium regulates cell death in cancer: roles of the mitochondria and mitochondria-associated membranes (MAMs). Biochim. Biophys. Acta Bioenergy 1858, 615–627 (2017).
Akl, H. & Bultynck, G. Altered Ca(2+) signaling in cancer cells: proto-oncogenes and tumor suppressors targeting IP3 receptors. Biochim. Biophys. Acta 1835, 180–193 (2013).
Bonneau, B., Prudent, J., Popgeorgiev, N. & Gillet, G. Non-apoptotic roles of Bcl-2 family: the calcium connection. Biochim. Biophys. Acta Mol. Cell Res. 1833, 1755–1765 (2013).
Denton, R. M. Regulation of mitochondrial dehydrogenases by calcium ions. Biochim. Biophys. Acta Bioenerg. 1787, 1309–1316 (2009).
Gunter, T. E., Buntinas, L., Sparagna, G., Eliseev, R. & Gunter, K. Mitochondrial calcium transport: mechanisms and functions. Cell Calcium 28, 285–296 (2000).
Green, D. R. & Reed, J. C. Mitochondria and apoptosis. Science 281, 1309–1312 (1998).
Bisaillon, R. et al. Genetic characterization of ABT-199 sensitivity in human AML. Leukemia 34, 63–74 (2020).
Buettner, R. et al. Targeting the metabolic vulnerability of acute myeloid leukemia blasts with a combination of venetoclax and 8-chloro-adenosine. J. Hematol. Oncol. 14, 70 (2021).
Maiti, A. et al. Outcomes of relapsed or refractory acute myeloid leukemia after front-line hypomethylating agent and venetoclax regimens. Haematologica 106, 894–898 (2020).
Pan, R. et al. Synthetic lethality of combined Bcl-2 Inhibition and p53 activation in AML: mechanisms and superior antileukemic efficacy. Cancer Cell 32, e6 (2017).
Savona, M. R. & Rathmell, J. C. Mitochondrial homeostasis in AML and gasping for response in resistance to BCL2 blockade. Cancer Discov. 9, 831–833 (2019).
Chen, X. et al. Targeting mitochondrial structure sensitizes acute myeloid leukemia to venetoclax treatment. Cancer Discov. 9, 890–909 (2019).
Nechiporuk, T. et al. The TP53 apoptotic network is a primary mediator of resistance to BCL2 inhibition in AML cells. Cancer Discov. 9, 910–925 (2019).
Lin, K. H. et al. Systematic dissection of the metabolic-apoptotic interface in AML reveals heme biosynthesis to be a regulator of drug sensitivity. Cell Metab. 29, e7 (2019).
Rambow, F. et al. Toward minimal residual disease-directed therapy in melanoma. Cell 174, e19 (2018).
Lacroix, M., Riscal, R., Arena, G., Linares, L. K. & Le Cam, L. Metabolic functions of the tumor suppressor p53: Implications in normal physiology, metabolic disorders, and cancer. Mol. Metab. 33, 2–22 (2020).
Haq, R. et al. Oncogenic BRAF regulates oxidative metabolism via PGC1α and MITF. Cancer Cell 23, 302–315 (2013).
Brummer, C. et al. Metabolic targeting synergizes with MAPK inhibition and delays drug resistance in melanoma. Cancer Lett. 442, 453–463 (2019).
Kim, E. M. et al. Nuclear and cytoplasmic p53 suppress cell invasion by inhibiting respiratory complex-I activity via Bcl-2 family proteins. Oncotarget 5, 8452–8465 (2014).
Green, D. R., Galluzzi, L. & Kroemer, G. Metabolic control of cell death. Science 345, 1250256 (2014).
Baccelli, I. et al. Mubritinib targets the electron transport chain complex i and reveals the landscape of OXPHOS dependency in acute myeloid leukemia. Cancer Cell 36, e8 (2019).
Saland, E. et al. A robust and rapid xenograft model to assess efficacy of chemotherapeutic agents for human acute myeloid leukemia. Blood Cancer J. 5, e297 (2015).
Petronilli, V. et al. Transient and long-lasting openings of the mitochondrial permeability transition pore can be monitored directly in intact cells by changes in mitochondrial calcein fluorescence. Biophys. J. 76, 725–734 (1999).
Tosolini, M. et al. Human monocyte recognition of adenosine-based cyclic dinucleotides unveils the A2a Gαs protein-coupled receptor tonic inhibition of mitochondrially induced cell death. Mol. Cell. Biol. 35, 479–495 (2015).
Argüello, R. J. et al. SCENITH: a flow cytometry-based method to functionally profile energy metabolism with single-cell resolution. Cell Metab. 32, e7 (2020).
Chong, J. et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 46, W486–W494 (2018).
Chen, L. et al. HNF4 regulates fatty acid oxidation and is required for renewal of intestinal stem cells in mice. Gastroenterology 158, e9 (2020).
Melamud, E., Vastag, L. & Rabinowitz, J. D. Metabolomic analysis and visualization engine for LC-MS data. Anal. Chem. 82, 9818–9826 (2010).
Su, X., Lu, W. & Rabinowitz, J. D. Metabolite spectral accuracy on orbitraps. Anal. Chem. 89, 5940–5948 (2017).
Spinazzi, M., Casarin, A., Pertegato, V., Salviati, L. & Angelini, C. Assessment of mitochondrial respiratory chain enzymatic activities on tissues and cultured cells. Nat. Protoc. 7, 1235–1246 (2012).
Mookerjee, S. A., Gerencser, A. A., Nicholls, D. G. & Brand, M. D. Quantifying intracellular rates of glycolytic and oxidative ATP production and consumption using extracellular flux measurements. J. Biol. Chem. 292, 7189–7207 (2017).
Chou, T.-C. Drug combination studies and their synergy quantification using the Chou–Talalay method. Cancer Res. 70, 440–446 (2010).
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010).
MacDonald, J. W. hugene20sttranscriptcluster.db: affymetrix hugene20 annotation data (chip hugene20sttranscriptcluster). R package version 8.7.0. (2017).
Hong, F. et al. RankProd: a bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics. 22, 2825–2827 (2006).
Van Noorden, C. J. F. The history of Z-VAD-FMK, a tool for understanding the significance of caspase inhibition. Acta Histochem. 103, 241–251 (2001).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Pont, F., Tosolini, M. & Fournié, J. J. Single-cell signature explorer for comprehensive visualization of single cell signatures across scRNA-seq datasets. Nucleic Acids Res. 47, e133 (2019).
Federico, A. & Monti, S. hypeR: an R package for geneset enrichment workflows. Bioinformatics 36, 1307–1308 (2020).
Acknowledgements
We thank all members of mice core facilities (UMS006, ANEXPLO, Inserm) in particular M. Lulka, C. Campi and all members of the CREFRE for their support and technical assistance and V. De Mas and E. Delabesse for the management of the Biobank BRC-HIMIP (Biological Resources Centres-Inserm Midi-Pyrénées Cytothèque des hémopathies malignes), which is supported by CAPTOR (Cancer Pharmacology of Toulouse-Oncopole and Région). We thank A. Melotti and IGE3 Genomic Platform (Geneva University) as well as F. Pont and F. Lopez (Pôle Technologique du CRCT, Inserm/U1037) and F. Martins (GET, GENOToul) for bulk DNA/RNA sequencing and single-cell RNA-seq procedures, respectively. We are grateful to the GENOToul Bioinformatics Platform Toulouse Midi-Pyrenees (Bioinfo Genotoul) for providing computing resources. This work was granted access to the HPC resources of CALMIP supercomputing center under the allocation 2019-T19001. 13C-isotope-tracing experiments were carried out at the Rutgers Cancer Institute of New Jersey. Metabolomics was performed at the MetaToul-MetaboHUB core facility (National Infrastructure of Metabolomics and Fluxomics) under supervision of L. Peyriga, F. Bellvert and J.-C. Portais. MetaToul is part of the national infrastructure MetaboHUB-ANR-11-INBS-0010 (French National infrastructure for Metabolomics and Fluxomics; www.metabohub.fr). MetaToul is supported by grants from the Région Midi-Pyrénées, the European Regional Development Fund, the SICOVAL, the Infrastructures en Biologie Santé et Agronomie, the Centre National de la Recherche Scientifique and the Institut National de la Recherche Agronomique Team. J.-E.S. is a member of OPALE Carnot Institute at the Organization for Partnerships in Leukemia. We thank A.-M. Benot, M. Serthelon and S. Nevouet for their daily help on the administrative and financial management of our team. The authors also thank N. Mazure for fruitful discussion about mitochondrial VDAC, M. Konopleva, I. Majewski and M. Selak for critical reading of the manuscript. This work was also supported by grants from the Programme Investissement d’Avenir PSPC (IMODI), the Laboratoire d’Excellence Toulouse Cancer (TOUCAN and TOUCAN2.0; contract ANR11-LABEX), INCA (PLBIO 2020-010, DIALAML), the Fondation Toulouse Cancer Santé, the Fondation ARC, the Ligue National de Lutte Contre le Cancer, the association Prolific and the association GAEL. A.S. is a fellow of the European Regional Development Fund through the Interreg V-A Spain-France-Andorra program, project PROTEOblood (EFA360/19). C.B. has a fellowship from the Fondation ARC.
Author information
Authors and Affiliations
Contributions
C.B. and J.-E.S. conceived the study, designed the experiments and interpreted the results. J.-E.S. designed the research and supervised the study. C.B. developed experimental tools, performed in vitro experiments, performed in vivo treatment studies and analyzed data. E.S. developed experimental tools, performed mitochondria purifications, OxPHOS enzymatic activities and SCENITH analysis. E.S., N.G., M.S., G.C., T.F., E.B., M.G., P.M., C.L., A.S., N.A., L.J. and E.K. contributed to in vivo experiments. E.S. A.S., M.G. and M.P. collected primary samples, collected the patient information for the retrospective analysis. A.S. and M.G. contributed to animal care. Y.W. and X.S. performed LC–MS analysis. L.S. performed IC–MS experiments and analysis and contributed to LC–MS analyses. L.P.P. contributed to LC–MS analyses. L.S. contributed to data interpretation. G.C. performed NMR analysis. M.S. collected MOLM14 in vivo samples for RNA-seq. H.A.L. and C.M. sequenced single-cell RNA-seq and single-cell DNA-seq data. J.T. sequenced RNA-seq data. A.B., M.T., N.P. and T.K. performed bioinformatics analysis. C.J. performed Duolink PLA. R.J. provided SCENITH substrates and help for data analysis. Q.F. and J.K. provided matched patients with AML for metabolic correlations analyses. I.T. and A.H.W. provided RNA-seq and data for clinical patients. J.J. and F.C. provided conceptual input to data interpretation. F.V. and C.R. provided clinical expertise. F.R. and J.C.M developed single-cell tools. N.N. performed Affymetrix experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks Michael Andreeff, Daniel Herranz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AML cells rely on mitochondrially bound BCL2 to sustain OxPHOS-dependent resistance to cytarabine.
(a, b) Mitochondrial membrane potential (MMP) using TMRE probe (a) and mitochondrial using MTG probe (b), on primary AML sample from Toulouse University Hospital (TUH) at diagnosis and relapse (n=16 patients). **P=0.0013; P=0.1591 (c) Cumulative total cell tumor burden assessed in mice bone marrow and spleen at day 0 (D0) and at day 8 (D8) after AraC treatment. The experiment was performed 10 times: in 9 different PDXs (60 mg/kg) and one CLDX (MOLM14, 30 mg/kg). Each dot is a mouse (MOLM14, CTL n=5 mice, AraC n=7 mice, **P =0.0025; TUH52, CTL n=4 mice, AraC n=7 mice, P=0.4121; TUH19, CTL n=6 mice, AraC n=4 mice, **P=0.0095; TUH57, CTL n=5 mice, AraC n=7 mice, **P=0.0025; TUH11, CTL n=10 mice, AraC n=10 mice, ****P < 0.0001; TUH55, CTL n=4 mice, AraC n=4 mice, *P=0.0286; TUH10, CTL n=7 mice, AraC n=6 mice, **P=0.0012; TUH06, CTL n=5 mice, AraC n=4 mice, *P=0.0159; TUH23, CTL n=5 mice, AraC n=5 mice, **P=0.0079; TUH30, CTL n=7 mice, AraC n=6 mice, **P=0.0012;). PDXs and CLDX are ranked from lower responders (Fold Change, FC, <10) to higher responders (FC > 10). (d) BCL2 expression at diagnosis and relapse after intensive chemotherapy according to Hackl et al. 2011 (n=10 patients). *P=0.0488. (e) BCL2 gene expression in AML cells compared with ‘normal’ hematopoietic stem cells (HSC) and peripheral blood mononuclear cells (PBMC) using Bloopool database (n=44 HSC/PBMC and n=1309 AML patients). ***P=0.0004. (f) Western-Blotting for BCL2, MCL1 and actinin in 2 normal peripheral blood mononuclear cells (PBMC) and 7 different AML cells lines. The experiment was performed twice with independent experiments and a representative example is shown. (g, h) BCL2 expression in MOLM14 CLDX after in vivo AraC treatment (30 mg/kg) (experiment performed twice with independent experiments; representative image shown) (g), and in PDXs after in vivo AraC treatment (60 mg/kg) in NSG mice (experiment was performed in 4 independent PDXs) (h). (i) Western-Blotting for BCL2 and HSP90 after AraC treatment for 24H. n=9 different primary AML samples. (j) Western-Blotting for BCL2 and actinin in MOLM14 transduced with CTL or BCL2 shRNAs. The experiment was performed 4 times and a representative example is shown. (k) Basal oxygen consumption rates of MOLM14 transduced with CTL or BCL2 shRNAs (n=4 independent experiments). *P=0.0286. (l) Citrate, α-ketoglutarate (α-KG), fumarate (Fum) and malate amounts measured by IC/MS in MOLM14 cells transduced with CTL or BCL2 shRNAs. Each dot is an experimental replicate of three independent experiments (n=3). (Citrate, ****P < 0.0001; α-KG, **P=0.0042; Fum, **P=0.0017; Malate, **P=0.0035) (m) EC50 for AraC of MOLM14 transduced with CTL or BCL2 shRNAs. (n=3 independent experiments). P values: *P=0.0139. (n, o) Loss of mitochondrial membrane potential (n) using TMRE staining (*P=0.0138; **P=0.0014; ***P=0.0004) and percent of viable cells (o) (*P=0.0022; ***P=0.0004; ****P < 0.0001), following 24H of AraC treatment in MOLM14 transduced with CTL or BCL2 shRNAs. (n=4 independent experiments). (p) EC50 for venetoclax of patient resistant (R, IC50 > 0.25 μM) vs sensitive (S, IC50 < 0.25 μM) in the two different cohorts TUH (n=22 patients) and BeatAML (n=59 patients). ****P < 0.0001. (q) BCL2 gene expression in TUH (n=7 patients) and BeatAML (n=59 patients) cohorts with resistant (R, IC50 > 0.25 μM) vs sensitive (S, IC50 < 0.25 μM) patients to venetoclax, based on the median sensitivity of the BeatAML cohort. *P=0.0471; ****P < 0.0001. (r, s) Gene Set Enrichment Analysis (GSEA) of gene signatures from Molecular Signature Data Base (MSigDB) and Gene Ontology Biological Processes (GOBP) in patients from the BeatAML cohort (n=59 patients) (r) and TUH cohort (n=7 patients) (s), resistant (R, IC50 > 0.25 μM) vs sensitive (S, IC50 < 0.25 μM) to venetoclax. Data in C, E, K, L, M, N, O, P and Q are displayed as mean ± s.e.m. Data analysis in A, B, D is by two-tailed paired Wilcoxon test and in C, E, K, L, M, N, O, P, Q by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance.
Extended Data Fig. 2 Selective BCL2 inhibitor venetoclax restores sensitivity of AML cells to AraC.
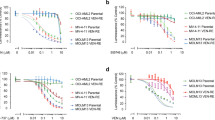
(A,B,C) Concentration response for the MOLM14 (a), U937 (b) and OCI-AML3 (c) AML cell lines upon exposure to simultaneous combinations of venetoclax (VEN) and aracytine (AraC) assessed by annexin V/7AAD staining following 24H of treatment. Scale bar: cell viability 0 to 100%. The experiment was performed 3 times and a representative example is shown. (d–f) Combination index data analysis of the concentration response data in A,B and C panels. Synergy effect correspond to a CI < 1, strong synergy effect CI < 0.3 and antagonist effect CI > 1. (n=3 independent experiments). (g) Percent of viable MOLM14, U937, KG1a and OCI-AML3 cells following treatment with venetoclax (0.5 μM) and aracytine (0.5 μM) for 24H. Values are mean ± s.e.m (n=5 independent experiments). Data analysis is by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance. P values: MOLM14, CTL/VEN **P=0.0079, CTL/AraC *P=0.0159, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC **P=0.0079; U937, CTL/VEN P=0.3095, CTL/AraC *P=0.0317, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.3095; KG1a, CTL/VEN **P=0.0079, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC **P=0.0079; OCIAML3, CTL/VEN *P=0.0159, CTL/AraC P=0.3095, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.2222. (h, i) Immunoblotting of cell lines (H) and primary AML cells (I) with the indicated antibodies following 24 h of treatment with venetoclax (Cell lines: 0.5 μM; Primary cells: 50 nM) and aracytine (Cell lines: 0.5 μM, Primary cells :25 μM). The experiment was performed twice with independent experiments and a representative example is shown.
Extended Data Fig. 3 Venetoclax disrupts mitochondrially bound BCL2 and decreases mitochondrial activities without affecting mitochondrial protein expression levels.
(a) Purified mitochondria from U937, MOLM14, KG1A immunoblotted with the indicated antibodies. The experiment was performed once for 3 different AML cell lines. (b) Spare respiratory capacity of MOLM14, U937, KG1A and OCIAML3. OCIAML3, CTL/VEN *P=0.0305, CTL/AraC P=0.0635, CTL/VEN+AraC *P=0.0248, AraC/VEN+AraC **P=0.0019; U937, CTL/VEN *P=0.0203, CTL/AraC *P=0.0309, CTL/VEN+AraC P=0.5490, AraC/VEN+AraC *P=0.0329; MOLM14, CTL/VEN **P=0.0052, CTL/AraC *P=0.0361, CTL/VEN+AraC P=0.1431, AraC/VEN+AraC *P=0.0125; KG1A, CTL/VEN *P=0.0169, CTL/AraC *P=0.0106, CTL/VEN+AraC ***P=0.0009, AraC/VEN+AraC **P=0.0027. (c,d) Western-Blotting for ETC complexes and HSP90 in cell lines (c) and primary AML samples (d). Two independent experiments were performed and a representative example is shown. (e,f) Mitochondrial mass using MTG probe in cell lines (e) and in PDXs (f). OCIAML3, CTL/AraC **P=0.0079, CTL/VEN+AraC P=0.6825, AraC/VEN+AraC P=0.8413; U937, CTL/AraC P=0.1270, CTL/VEN+AraC P=0.1270, AraC/VEN+AraC P=0.3095; MOLM14, CTL/AraC P=0.1270, CTL/VEN+AraC P=0.1270, AraC/VEN+AraC P=0.8413; KG1A, CTL/AraC P=0.6825, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC **P=0.0079; TUH07, D0/AraC P=0.0519, D0/VEN+AraC P=0.0823, AraC/VEN+AraC P=0.0931; TUH35, D0/AraC **P=0.0022, D0/VEN+AraC ***P=0.0007, AraC/VEN+AraC P=0.0926; TUH30, D0/AraC **P=0.0016, D0/VEN+AraC **P=0.0016, AraC/VEN+AraC P=0.2222. (g,h) Mitochondrial membrane potential using TMRE staining in 4 different cell lines (g) and in 3 different PDXs (h). OCIAML3, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC *P=0.0159; U937, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P > 0.9999; MOLM14, CTL/AraC P=0.1270, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.1508; KG1A, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.8413; TUH07, D0/AraC P=0.3853, D0/VEN+AraC P=0.7554, AraC/VEN+AraC P=0.6688; TUH35, D0/AraC **P=0.0087, D0/VEN+AraC ***P=0.0007, AraC/VEN+AraC P=0.0813; TUH30, D0/AraC **P=0.0031, D0/VEN+AraC **P=0.0016, AraC/VEN+AraC P=0.1508. (i) Mitochondrial calcium content using rhodamin2 probe. OCIAML3, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.6905; U937, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.5476; MOLM14, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC P=0.8413; KG1A, CTL/AraC **P=0.0079, CTL/VEN+AraC **P=0.0079, AraC/VEN+AraC **P=0.0079. In B for OCIAML3 and KG1A n=3 independent experiments, for U937 and MOLM14 n=4 independent experiments. In E, G and I n=5 independent experiments. Mice number in F and H: TUH07, D0 n=5, AraC n=6, VEN+AraC n=6; for TUH35, D0 n=6, AraC n=5, VEN+AraC n=8; TUH30, D0 n=8, AraC n=5, VEN+AraC n=5. Data in B, E, F, G, H and I are displayed as mean ± s.e.m. Data analysis in B, E, F, G, H and I is by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance.
Extended Data Fig. 4 Venetoclax blocks AraC-induced OXPHOS and metabolic phenotypes.
(a) An integrated analysis of IC/MS data based on Metaboanalyst software (pathway tool) for a simplified view of contributing pathways of MOLM14 cells. (b,c) The glycolytic metabolites: glucose 6-phosphate (G6P) (n=6 independent experiments), fructose 6-phosphate (F6P), 2,3-bisphosphoglycerate (2/3PG), amounts (n=3 independent experiments) (b), and the pentose phosphate pathway (PPP) metabolites (n=3 independent experiments): 6-phosphogluconate (6PG), ribose 1-phosphate (Rib1P), sedoheptulose 7-phosphate (Sed7P) amounts (c) measured by IC/MS in MOLM14 cells. Data are represented in fold change to control. P values: G6P, CTL/VEN P=0.3648, CTL/AraC *P=0.0338, CTL/VEN+AraC P=0.1367, AraC/VEN+AraC *P=0.0261; F6P, nsP > 0.1111; 2/3PG, nsP > 0.1111; 6PG, nsP > 0.1111; Rib1P, nsP > 0.1111; Sed7P, nsP > 0.1111; Aspartate, CTL/VEN P=0.0877, CTL/AraC *P=0.0106, CTL/VEN+AraC *P=0.0267, AraC/VEN+AraC *P=0.0170; Glutamate, CTL/VEN P=0.3735, CTL/AraC *P=0.0186, CTL/VEN+AraC P=0.4883, AraC/VEN+AraC P=0.0837. (d) Aspartate and glutamate amount measured by IC/MS in MOLM14 cells. Data are represented in fold change to control. (n=4 independent experiments). (e,f,g) Basal and maximal JATP production in KG1A (e), OCIAML3 (f) and U937 (g) cells, divided into JATPglyc and JATPox using a Seahorse XF24 Extracellular Analyzer. Aggregate data from 4 independent experiments. (h) The contribution of glucose to aspartate and glutamate production in MOLM14 cells (n=2 independent experiments). Data in A, B, C, D, E, F, G, and H are assessed 24H post in vitro treatment with venetoclax (VEN, 0.5 μM) and/or cytarabine (AraC, 0.5 μM). Data in B, C, D, E, F, G, and H are displayed as mean ± s.e.m. Data analysis in B, C and D is by two-tailed paired t-test. *P < 0.05, **P < 0.01, ****P < 0.0001, ns:non-significant.
Extended Data Fig. 5 Venetoclax improves anti-AML effects of AraC without additional in vivo toxicity.
(a) Percent of loss of mitochondrial membrane potential in human AML cells in the bone marrow of leukemic mice were assessed by flow cytometry using fluorescent TMRE probe staining. P values: TUH10, D0/AraC P=0.9898, D0/VEN+AraC ****P < 0.0001, AraC/VEN+AraC **** P < 0.0001; TUH28, D0/AraC P=0.0513, D0/VEN+AraC ***P=0.0006, AraC/VEN+AraC P=0.0734; TUH35, D0/AraC P=0.8182, D0/VEN+AraC ***P=0.0013, AraC/VEN+AraC *P=0.0127; TUH59, D0/AraC *P=0.0317, D0/VEN+AraC *P=0.0159, AraC/VEN+AraC **P=0.0079; TUH30, D0/AraC P=0.2844, D0/VEN+AraC *P=0.0451, AraC/VEN+AraC P=0.1508; TUH07, D0/AraC P=0.5368, D0/VEN+AraC **P=0.0043, AraC/VEN+AraC P=0.1320. (b,c) Change in mice weight (P values: nsP > 0.1111) (b) and murine CD45 count in bone marrow and spleen (P values: nsP > 0.1111, ****P < 0.0001) (c). (d) Change in peripheral blood indices (hematocrite, white blood cells, red blood cells and platelets) in mice during the different treatments of PDX TUH35. P values: Hematocrite and WBC, D0/AraC **P=0.0022, D0/VEN+AraC ***P=0.0007, AraC/VEN+AraC P=0.3233; RBC, nsP > 0.1111; Platelets, D0/AraC *P=0.0411, D0/VEN+AraC **P=0.0013, AraC/VEN+AraC *P=0.0293. In A, B, C and D for TUH10, D0 n=12 mice, AraC n=13 mice, VEN+AraC n=13 mice; for TUH28, D0 n=7 mice, AraC n=6 mice, VEN+AraC n=7 mice; for TUH35, D0 n=6 mice, AraC n=6 mice, VEN+AraC n=8 mice; TUH59, D0 n=4 mice, AraC n=5 mice, VEN+AraC n=5 mice; for TUH30, D0 n=8 mice, AraC n=5 mice, VEN+AraC n=5 mice; for TUH07, D0 n=5 mice, AraC n=6 mice, VEN+AraC n=6 mice. Data in A, B, C and D are assessed post in vivo treatment with venetoclax (100 mg/kg) and aracytine (30 mg/kg). (e,f) Apoptosis induction post 24H of in vitro treatment with venetoclax (50 nM) and aracytine (25 μM) (E) and immunoblotting before treatments with the indicated antibodies (F) of primary AML cells (n=18 patients). Data in A, B, C, and D are displayed as mean ± s.e.m. Data analysis in A, B, C, and D is by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance. *P < 0.05, **P < 0.01, ****P < 0.0001, ns:non-significant.
Extended Data Fig. 6 Patients with a high MitoScore are predicted to be high responders to VEN plus AraC and not to VEN plus AZA doublet therapy.
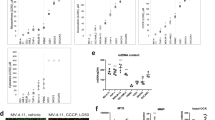
(a) Gene set enrichment analysis (GSEA) of the gene ontology biological processes (GOBP) data base at diagnosis in patient receiving the duplet therapy venetoclax plus low dose acracytine (LDAC) from the study VIALE-C (NCT03069352). RNA-seq data available through the European Genome-phenome Archive (EGAS00001003820). Dotted line at −1.3 (log10(0.05)) indicates the threshold limit of the q-value below which the gene signatures are significantly enriched. (n=19 patients). (b–d) Kaplan-Meier plots of overall survival (OS) for patients with AML treated with venetoclax in association with LDAC (n=19 patients) based on two different OxPHOS score category (MOOTHA_MITOCHONDRIA geneset, B; FARGE_HIGH_OXPHOS geneset, C, and FAO score category (FATTY_ACID_METABOLISM geneset, D). Data analysis is by log-rank Mantel-Cox’s t-test. (e) Heatmap of gene-expression values depicting CPT1a and BCL2 family members in AML patients treated with venetoclax in association with LDAC (n=19 patients) and classified either by their MitoScore (Low vs High) or by their initial response (CR+CRi vs RD). Low to high expression is represented by a change of colour from orange to green, respectively. (f) OncoPrint of genetic alterations in patients with AML treated with venetoclax in association with either LDAC (VIALE-C, n=19 patients) or HMA (VIALE-A, n=12 patients) classified in MitoScoreLow and MitoScoreHigh. (g) Data of oxygen consumption rate (OCR) in the primary sample TUH161. (n=1 with 3 technical replicates). Representation of the spare respiratory capacity (SRC) calculation.
Extended Data Fig. 7 Single-cell transcriptomic analysis confirms the enrichment of lipids-related gene signatures in AraC-specific residual cell clusters.
(a,b) Cumulative total cell tumor burden of human viable CD45+CD33+CD44+ AML cells for PDX TUH07 (A) and TUH69 (B) was assessed in bone marrow and spleen at day 0 (D0) and at day 8 (D8) for the three different treatments by flow cytometry. Data are assessed post in vivo treatment with venetoclax (100 mg/kg) and aracytine (30 mg/kg). Data are displayed as mean ± s.e.m. TUH07, D0 n=4 mice, VEN n=6, AraC n=5 mice, VEN+AraC n=6 mice; TUH69, D0 n=5 mice, VEN n=5, AraC n=5 mice, VEN+AraC n=8 mice. P values: TUH07, D0/VEN **P=0.0095, D0/AraC *P=0.0159, D0/VEN+AraC **P=0.0095, AraC/VEN+AraC **P=0.0043; TUH69, D0/VEN P=0.1508, D0/AraC P=0.3095, D0/VEN+AraC **P=0.0016, AraC/VEN+AraC **P=0.0451. Data analysis in A and B is by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance. *P < 0.05, **P < 0.01, ****P < 0.0001, ns:non-significant. (c) Unsupervised hierarchical clustering was done by principal component analysis (PCA) in PDX TUH07 and TUH69. 9 different clusters were identified. (d) UMAP visualization of single cell from PDX TUH07 and TUH69 colored per patient (n=31,604 cells). (e) Visualization of the repartition of the total cell count per cluster with distinction of cells coming from each PDX. (f) Visualization of the repartition of the total cell count per cluster with the different conditions highlighted (Day0; Day8 following VEN; AraC; VEN plus AraC). (g) Visualization of single cell scores for CD36 gene enrichment on violin plots. The center line represents the median (Q2) and the box spans the 25th (Q1) to 75th percentiles (Q3). Minimum (Q0) and maximum (Q4) values are outside the box. D0 (Q0: −4.2108; Q1: −0.9785; Q2: 0.347; Q3: 2.3348; Q4: 5.1356), VEN (Q0: −2.7858; Q1: 0.669; Q2: 2.4531; Q3: 3.0013; Q4: 5.2046), AraC (Q0: −2.9885; Q1: −0.454; Q2: 1.8428; Q3: 3.0775; Q4: 5.019), VEN+AraC (Q0: −1.5777; Q1: 1.0628; Q2: 2.2119; Q3: 2.8247; Q4: 4.7953). (h,i) Visualization on the UMAP plot of single cell scores for OxPHOS gene signature enrichment. (H) and fatty acid metabolism gene signature enrichment (I) using Single-Cell Signature Explorer.
Extended Data Fig. 8 VEN plus AraC specific residual cell clusters are phenotypically heterogeneous in vivo.
(a–c) Visualization on the UMAP plot of hematopoietic cell signatures that best match the three VEN+AraC clusters (#6, #7, #8) using Single Cell Explorer. Cluster #6 is monocyte-like based on gene signature created from Wu et al. (Journal of Hematology & Oncology, 2020). (A); Cluster #7 is HSC like based on Van Galen gene signature (Van Galen et al. Cell, 2019). (B) and cluster #8 is Myelo-Erythroid Progenitor (MEP) like based on gene signature created from Velten et al. NCB, 2017 (C). (d) UMAP plot displaying SCENIC clusters identified based on Gene Regulatory Network inference. Colors indicate SCENIC clusters. (e) Heatmap displaying binary activity of regulons identified by SCENIC Gene regulatory network analysis.
Extended Data Fig. 9 VEN plus AraC specific residual cell clusters are transcriptionally (and not genetically) heterogeneous in vivo.
(a–c) Volcano plots showing active regulons identified by SCENIC gene regulatory network analysis for the Seurat clusters #6 (A), #7 (B), #8 (C). (D,E) UMAP plot displaying binary activity of top regulons MITF (d) and TP53 (e). (f) Repartition of clonal prevalence in PDX TUH69 per condition (Day0; Day8 following VEN; AraC; VEN+AraC) using single-cell mutational analysis via Mission Bio Tapestri.
Extended Data Fig. 10 Mitochondrial inhibitors enhance anti-AML effects of VEN plus AraC in vitro.
(a,b) Percent of loss of mitochondrial membrane potential in MOLM14 (A) and U937 (B) AML cell lines following 24 h of treatment with venetoclax plus aracytine (0.5 μM; 0.5 μM) in combination or not with an electron transfer chain (ETC) complex I inhibitors (IACS-010759, 1 μM; ONC-212, 1 μM; CPI-613, 200 μM). Loss of mitochondrial membrane potential was assessed by flow cytometry using fluorescent TMRE probe staining. Data are displayed as mean ± s.e.m (n=4 independent experiments). P values: MOLM14, CTL/VEN *P=0.0327, CTL/AraC P=0.2002, CTL/VEN+AraC **P=0.0017, AraC/VEN+AraC **P=0.0016, IACS/VEN+IACS **P=0.0015, IACS/AraC+IACS **P=0.0022, IACS/VEN+AraC+IACS ****P < 0.0001, AraC+IACS/VEN+AraC+IACS ***P=0.0003, ONC/VEN+ONC ***P=0.0004, ONC/AraC+ONC *P=0.0155, ONC/VEN+AraC+ONC ***P=0.0001, AraC+ONC/VEN+AraC=ONC ****P < 0.0001, CPI/VEN+CPI P=0.0543, CPI/AraC+CPI **P=0.0047, CPI/VEN+AraC+CPI ***P=0.0002, AraC+CPI/VEN+AraC=CPI **P=0.0020; U937, CTL/VEN P=0.2558, CTL/AraC P=0.8183, CTL/VEN+AraC *P=0.0375, AraC/VEN+AraC *P=0.0472, IACS/VEN+IACS *P=0.0138, IACS/AraC+IACS P=0.1499, IACS/VEN+AraC+IACS **P=0.0081, AraC+IACS/VEN+AraC=IACS *P=0.0187, ONC/VEN+ONC ****P < 0.0001, ONC/AraC+ONC P=0.1076, ONC/VEN+AraC+ONC **P=0.0020, AraC+ONC/VEN+AraC=ONC **P=0.0021, CPI/VEN+CPI P=0.1345, CPI/AraC+CPI *P=0.0320, CPI/VEN+AraC+CPI *P=0.0136, AraC+CPI/VEN+AraC=CPI P=0.0953. Data analysis in A and B is by two-tailed unpaired Student’s t-test with Welch’s correction for unequal variance. *P < 0.05, **P < 0.01, ****P < 0.0001, ns:non-significant. (c) Gating strategy for flow cytometry analysis. Cells from the bone marrow compartment of NSG mice are selected according to the forward scatter (FSC) and side scatter (SSC) area (A) parameters. Then, doublet exclusion is performed using height (H) versus area (A) parameters of FSC and live/dead discrimination is applied using annexinV dye. The AML blast population is CD45+CD33+. The tumor burden is determined using the beads and blasts true count.
Supplementary information
Supplementary Tables
Supplementary Tables 1–7.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Bosc, C., Saland, E., Bousard, A. et al. Mitochondrial inhibitors circumvent adaptive resistance to venetoclax and cytarabine combination therapy in acute myeloid leukemia. Nat Cancer 2, 1204–1223 (2021). https://doi.org/10.1038/s43018-021-00264-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00264-y
This article is cited by
-
Decoding leukemia at the single-cell level: clonal architecture, classification, microenvironment, and drug resistance
Experimental Hematology & Oncology (2024)
-
Chronic lymphocytic leukemia patient-derived xenografts recapitulate clonal evolution to Richter transformation
Leukemia (2024)
-
ACOX1-mediated peroxisomal fatty acid oxidation contributes to metabolic reprogramming and survival in chronic lymphocytic leukemia
Leukemia (2024)
-
Mitochondrial fusion is a therapeutic vulnerability of acute myeloid leukemia
Leukemia (2023)
-
To metabolomics and beyond: a technological portfolio to investigate cancer metabolism
Signal Transduction and Targeted Therapy (2023)