Abstract
Expanding the utility of immune-based cancer treatments is a clinical challenge due to tumor-intrinsic factors that suppress the immune response. Here we report the identification of tumoral ring finger protein 2 (RNF2), the core subunit of polycomb repressor complex 1, as a negative regulator of antitumor immunity in various human cancers, including breast cancer. In syngeneic murine models of triple-negative breast cancer, we found that deleting genes encoding the polycomb repressor complex 1 subunits Rnf2, BMI1 proto-oncogene, polycomb ring finger (Bmi1), or the downstream effector of Rnf2, remodeling and spacing factor 1 (Rsf1), was sufficient by itself to induce durable tumor rejection and establish immune memory by enhancing infiltration and activation of natural killer and CD4+ T cells, but not CD8+ T cells, into the tumor and enabled their cooperativity. These findings uncover an epigenetic reprogramming of the tumor-immune microenvironment, which fosters durable antitumor immunity and memory.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
ChIP-seq, RNA-seq and ATAC-seq data that support the findings of this study have been deposited in GEO under accession code GSE143352. Previously published ChIP-seq data that were re-analyzed here are available under accession code GSE10573 (ref. 47), GSE34520 (ref. 45) and GSE107176 (ref. 22). Human pan-cancer and BRCA data were derived from the TCGA Research Network at https://portal.gdc.cancer.gov/. For pan-cancer analysis, standardized, normalized, batch-corrected and platform-corrected datasets were downloaded from the TCGA consortium (https://gdc.cancer.gov/about-data/publications/pancanatlas). Copy-number variation data were downloaded from Broad GDAC firehose (http://gdac.broadinstitute.org/runs/analyses__latest/data/). Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
Code availability
The software and methods to generate the results are described in detail in Methods. No additional code needs to be provided.
References
Leach, D. R., Krummel, M. F. & Allison, J. P. Enhancement of antitumor immunity by CTLA-4 blockade. Science 271, 1734–1736 (1996).
Patel, S. A. & Minn, A. J. Combination cancer therapy with immune checkpoint blockade: mechanisms and strategies. Immunity 48, 417–433 (2018).
Frankel, T., Lanfranca, M. P. & Zou, W. The role of tumor microenvironment in cancer immunotherapy. Adv. Exp. Med. Biol. 1036, 51–64 (2017).
Benci, J. L. et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell 178, 933–948 (2019).
Hegde, P. S. & Chen, D. S. Top 10 challenges in cancer immunotherapy. Immunity 52, 17–35 (2020).
Yu, J. et al. Liver metastasis restrains immunotherapy efficacy via macrophage-mediated T cell elimination. Nat. Med. https://doi.org/10.1038/s41591-020-1131-x (2021).
Kalbasi, A. & Ribas, A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat. Rev. Immunol. 20, 25–39 (2020).
Bian, Y. et al. Cancer SLC43A2 alters T cell methionine metabolism and histone methylation. Nature 585, 277–282 (2020).
Sanmamed, M. F. & Chen, L. A paradigm shift in cancer immunotherapy: from enhancement to normalization. Cell 175, 313–326 (2018).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563 (2018).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Science 359, 770–775 (2018).
Adeegbe, D. O. et al. Synergistic immunostimulatory effects and therapeutic benefit of combined histone deacetylase and bromodomain inhibition in non-small cell lung cancer. Cancer Discov. 7, 852–867 (2017).
Li, F. et al. In vivo epigenetic CRISPR screen identifies Asf1a as an immunotherapeutic target in Kras-mutant lung adenocarcinoma. Cancer Discov. 10, 270–287 (2020).
Peng, D. et al. Epigenetic silencing of TH1-type chemokines shapes tumour immunity and immunotherapy. Nature 527, 249–253 (2015).
Li, J. et al. Epigenetic driver mutations in ARID1A shape cancer immune phenotype and immunotherapy. J. Clin. Invest. 130, 2712–2726 (2020).
Burr, M. L. et al. An evolutionarily conserved function of polycomb silences the MHC class I antigen presentation pathway and enables immune evasion in cancer. Cancer Cell 36, 385–401 (2019).
Su, W. et al. The polycomb repressor complex 1 drives double-negative prostate cancer metastasis by coordinating stemness and immune suppression. Cancer Cell 36, 139–155 (2019).
Connolly, R. M. et al. Combination epigenetic therapy in advanced breast cancer with 5-azacitidine and entinostat: a phase II National Cancer Institute/Stand Up to Cancer study. Clin. Cancer Res. 23, 2691–2701 (2017).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997 (2018).
Piunti, A. & Shilatifard, A. Epigenetic balance of gene expression by polycomb and COMPASS families. Science 352, aad9780 (2016).
Rai, K. et al. Dual roles of RNF2 in melanoma progression. Cancer Discov. 5, 1314–1327 (2015).
Chan, H. L. et al. Polycomb complexes associate with enhancers and promote oncogenic transcriptional programs in cancer through multiple mechanisms. Nat. Commun. 9, 3377 (2018).
Zhang, Y. et al. Estrogen induces dynamic ERα and RING1B recruitment to control gene and enhancer activities in luminal breast cancer. Sci. Adv. 6, eaaz7249 (2020).
Laugesen, A. & Helin, K. Chromatin repressive complexes in stem cells, development, and cancer. Cell Stem Cell 14, 735–751 (2014).
Piunti, A. & Shilatifard, A. The roles of polycomb repressive complexes in mammalian development and cancer. Nat. Rev. Mol. Cell Biol. 22, 326–345 (2021).
Adams, S. et al. Current landscape of immunotherapy in breast cancer: a review. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2018.7147 (2019).
Zhang, Z. et al. Role of remodeling and spacing factor 1 in histone H2A ubiquitination-mediated gene silencing. Proc. Natl Acad. Sci. USA 114, E7949–E7958 (2017).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Cai, G., Yang, Q. & Sun, W. RSF1 in cancer: interactions and functions. Cancer Cell Int. 21, 315 (2021).
Wu, J. et al. The oncogenic impact of RNF2 on cell proliferation, invasion and migration through EMT on mammary carcinoma. Pathol. Res. Pract. 215, 152523 (2019).
Barger, C. J., Branick, C., Chee, L. & Karpf, A. R. Pan-cancer analyses reveal genomic features of FOXM1 overexpression in cancer. Cancers https://doi.org/10.3390/cancers11020251 (2019).
Shabason, J. E. & Minn, A. J. Radiation and immune checkpoint blockade: from bench to clinic. Semin. Radiat. Oncol. 27, 289–298 (2017).
Wang, W. et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569, 270–274 (2019).
Babic, M. et al. NK cell receptor NKG2D enforces proinflammatory features and pathogenicity of TH1 and TH17 cells. J. Exp. Med. https://doi.org/10.1084/jem.20190133 (2020).
Borst, J., Ahrends, T., Babala, N., Melief, C. J. M. & Kastenmuller, W. CD4(+) T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Minn, A. J. Interferons and the immunogenic effects of cancer therapy. Trends Immunol. 36, 725–737 (2015).
Feng, M. et al. Phagocytosis checkpoints as new targets for cancer immunotherapy. Nat. Rev. Cancer 19, 568–586 (2019).
Ross, S. H. & Cantrell, D. A. Signaling and Function of interleukin-2 in T lymphocytes. Annu. Rev. Immunol. 36, 411–433 (2018).
Xia, J., Gill, E. E. & Hancock, R. E. NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat. Protoc. 10, 823–844 (2015).
Klusmann, I. et al. Chromatin modifiers Mdm2 and RNF2 prevent RNA:DNA hybrids that impair DNA replication. Proc. Natl Acad. Sci. USA 115, E11311–E11320 (2018).
Lee, H. S. et al. The chromatin remodeler RSF1 controls centromeric histone modifications to coordinate chromosome segregation. Nat. Commun. 9, 3848 (2018).
Forero, A. et al. Expression of the MHC class II pathway in triple-negative breast cancer tumor cells is associated with a good prognosis and infiltrating lymphocytes. Cancer Immunol. Res. 4, 390–399 (2016).
Morey, L. & Helin, K. Polycomb group protein-mediated repression of transcription. Trends Biochem. Sci. 35, 323–332 (2010).
Brookes, E. et al. Polycomb associates genome-wide with a specific RNA polymerase II variant, and regulates metabolic genes in ESCs. Cell Stem Cell 10, 157–170 (2012).
Cohen, I. et al. PRC1 preserves epidermal tissue integrity independently of PRC2. Genes Dev. 33, 55–60 (2019).
Endoh, M. et al. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 135, 1513–1524 (2008).
Wang, H. et al. Role of histone H2A ubiquitination in polycomb silencing. Nature 431, 873–878 (2004).
Yasinska, I. M. et al. The Tim-3-galectin-9 pathway and its regulatory mechanisms in human breast cancer. Front. Immunol. 10, 1594 (2019).
Li, X. et al. Immune profiling of pre- and post-treatment breast cancer tissues from the SWOG S0800 neoadjuvant trial. J. Immunother. Cancer 7, 88 (2019).
Toniolo, P. A. et al. Deregulation of SOCS5 suppresses dendritic cell function in chronic lymphocytic leukemia. Oncotarget 7, 46301–46314 (2016).
Li, Q. et al. Adoptive transfer of tumor reactive B cells confers host T-cell immunity and tumor regression. Clin. Cancer Res. 17, 4987–4995 (2011).
Alspach, E. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019).
Ribas, A. et al. Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. JAMA 315, 1600–1609 (2016).
Hirschhorn-Cymerman, D. et al. Induction of tumoricidal function in CD4+ T cells is associated with concomitant memory and terminally differentiated phenotype. J. Exp. Med. 209, 2113–2126 (2012).
Kerdiles, Y., Ugolini, S. & Vivier, E. T cell regulation of natural killer cells. J. Exp. Med. 210, 1065–1068 (2013).
Waggoner, S. N., Cornberg, M., Selin, L. K. & Welsh, R. M. Natural killer cells act as rheostats modulating antiviral T cells. Nature 481, 394–398 (2011).
Jiao, L. et al. A partially disordered region connects gene repression and activation functions of EZH2. Proc. Natl Acad. Sci. USA 117, 16992–17002 (2020).
Boyle, S. et al. A central role for canonical PRC1 in shaping the 3D nuclear landscape. Genes Dev. 34, 931–949 (2020).
Illingworth, R. S. et al. The E3 ubiquitin ligase activity of RING1B is not essential for early mouse development. Genes Dev. 29, 1897–1902 (2015).
Hanna, A. et al. Inhibition of Hedgehog signaling reprograms the dysfunctional immune microenvironment in breast cancer. Oncoimmunology 8, 1548241 (2019).
Zabala, M. et al. Optimization of the Tet-on system to regulate interleukin 12 expression in the liver for the treatment of hepatic tumors. Cancer Res. 64, 2799–2804 (2004).
Li, C. et al. Novel HDAd/EBV reprogramming vector and highly efficient Ad/CRISPR-Cas sickle cell disease gene correction. Sci. Rep. 6, 30422 (2016).
Zhang, Z. et al. USP49 deubiquitinates histone H2B and regulates cotranscriptional pre-mRNA splicing. Genes Dev. 27, 1581–1595 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramirez, F. et al. epTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 44, e71 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Faust, G. G. & Hall, I. M. SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Fabregat, A. et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Acknowledgements
We thank L. Z. Shi and H. Wang for helpful discussion on this project. We are grateful to H. Shen and S. Bugide for technical assistance. E.S.Y. is a ROAR Southeast Cancer Foundation Endowed Chair. This work was supported by grants from Autotech LLC (to E.S.Y.), from Breast Cancer Research Foundation of Alabama (to E.S.Y.), from American Association for Cancer Research/Triple Negative Breast Cancer Foundation (15-20-43-YANG) (to E.S.Y.), and from start-up funds from UAB (to J.W.L.). J.W.L. is also supported by the DoD PRCRP Career Award (W81XWH-18-1-0315) and National Institutes of Health (NIH) R01AI148711. N.W. is supported by the DoD grant W81XWH-19-1-0084. We also thank V. S. Hanumanthu from the Comprehensive Flow Cytometry Core as well as the UAB Small Animal Imaging Shared Facility (supported by the O'Neal Comprehensive Cancer Center (NIH P30 CA013148) and funding to Rheumatic Diseases Core Center/Center for AIDS Research (NIH P30 AR048311/NIH P30 AI27667).
Author information
Authors and Affiliations
Contributions
Z.Z., J.W.L. and E.S.Y. conceived the concept. Z.Z., J.W.L., E.S.Y. and L.L. designed studies, interpreted data and wrote the manuscript. Z.Z., L.L., J.W.L., X.C., M.L., L.Z., C.L. and S.G. performed experiments with assistance from D.D.M. P.X., Y.C., Z.C. and Z.Z. performed bioinformatic analyses. N.W., T.T., W.J.B. and B.P.S. contributed to data interpretation and study design. J.W.L. and E.S.Y. collected funding support. E.S.Y. critically reviewed the manuscript and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
E.S.Y. is a consultant for AstraZeneca, Eli Lilly and Company and is on the Advisory Board for Bayer Pharmaceuticals, AstraZeneca, Clovis Oncology and Strata Oncology. The research in the laboratory of E.S.Y. is supported by funding from Novartis International AG, Eli Lilly and Company, Clovis Oncology and the American Society of Clinical Oncology. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks Lluis Morey and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 RNF2 marks the signature of immunologically cold tumors and is negatively associated with cytotoxicity of immune cells.
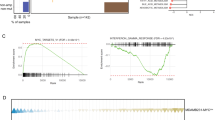
a, GSEA analysis of the gene signature of cold tumor (noted as T cell exclusion) revealed in single cell RNAseq of tumor cells from melanoma patients using oncogenic gene sets (C6). Gene sets related to epigenetic pathways are shown. n = 7,186 single tumor cells from 33 human melanoma tumors (from 31 patients). b-c, The ranking of 248 (b) or 524 (c) epigenetic regulators based on their median expression levels in relation to the high and low expression of ‘cold’ gene signature, extracted from the dataset in panel a via the Single Cell Portal. Each value at the y-axis for each gene is defined as its median expression level in the single tumor cells with high expression of cold gene signature minus that in the tumor cells with low expression of cold gene signature. These values were used to sort and plot genes on a two-dimensional plane. The x-axis denotes the rank of these genes. RNF2 is ranked as the top 4th/3rd gene. The number (n) of single tumor cells included in this single cell RNAseq is noted. d-f, Analysis of genes extracted from TCGA datasets. Heatmaps (Left) with the top parts zoomed in (Right) show correlations between expression levels of 248 epigenetic genes with the immune cell cytotoxicity gene signature, GZMA and PRF1, in diverse human cancers. The expression levels of these genes are extracted from TCGA. The coefficients (r) (d), and p values of these correlations (e) are shown. The zoomed in images of the top parts of the graph are shown in the right. f, The volcano plot shows the correlation of RNF2 to GZMA and PRF1 in human cancers. The y-axis and x-axis denote -Log10 (FDR) and coefficients (r) of correlations, respectively. The case numbers (n) are noted. n = 11,160 patients.
Extended Data Fig. 2 RNF2/CBX2/CBX8/RSF1 is amplified/overexpressed in human breast cancer patients, and associated with shorter survival time.
a, The box plot shows the expression levels of RNF2 in human normal breast tissue, invasive breast cancer tissues (Left) and triple negative breast cancer (TNBC) tissues (Right). Log2 fold change (Log2Fold) of cancer tissues relative to normal tissues and significance (p) of the correlations are displayed. The lower and upper bound of box plot represent the first quartile (Q1, 25% of data) and third quartile (Q3, 75% of data) of the data. Center line within the box represents the median value (also the second quartile). The whisker marks 1.5*IQR (Inter Quartile Range, the distance between Q1 and Q3) at both side of the box. Dots are the outliers, which are the values outside the whiskers (> Q3 + 1.5*IQR or < Q1 - 1.5*IQR). Normal controls for breast cancer patients: n = 125; breast cancer patients: n = 1,097; normal controls for TNBC patients: n = 112; TNBC patients: n = 123. b, The percentages of primary invasive breast cancer (BRCA), primary TNBC and metastatic breast cancer patients with RNF2 amplification. c, Images with patient IDs from The Human Protein Atlas (http://www.proteinatlas.org) display the expression of RNF2 protein in normal breast tissues and human breast cancer tissues by immunohistochemistry staining. Scale bars: 100 (Upper)/50 (Bottom) µm. The website link for each image is provided in the Resources Table. d, The percentages of invasive breast cancer patients (n = 1,981) with the gain and amplification of RSF1, obtained from the METABRIC dataset via cBioPortal. e, Images with patient IDs from The Human Protein Atlas (http://www.proteinatlas.org) display the expression of RSF1 protein in normal breast tissues and human breast cancer tissues by immunohistochemistry staining. Scale bars: 100 (Upper)/50 (Bottom) µm. The website link for each image is provided in the Resources Table. f, The percentages of breast cancer patients with the amplification of CBX2 and CBX8 obtained from METABRIC dataset via cBioPortal. g-i, The correlations of amplification of RNF2 (g) or RSF1 (h) or CBX2/CBX8 (i) to the survival of invasive breast cancer patients (PanCancer Atlas) (g), or breast cancer patients (METABRIC dataset) (h, i) obtained from cBioPortal. P value is generated using two-tailed LogRank Test.
Extended Data Fig. 3 Targeting Rnf2/Rsf1 results in tumor rejection in syngeneic murine breast cancer models.
a-c, a, Immunoblots of indicated proteins in 4T1 cells transduced with the scrambled shRNA or two independent shRNAs targeting Rnf2 gene, which represents two independent experiments with similar results. The knockdown efficiency has been independently confirmed. b-c, The volumes (b) and representative BLIs at day 28 after inoculation (c) of tumors transduced with shRNAs as described in panel a (mean ± SEM, n = 5 mice/group) in mice. d, Immunoblots of indicated proteins in Ctrl and Rnf2 KO (two independent gRNAs, g1 and g2) EMT6 cells. The knockout efficiency was independently confirmed at least one more time. e-h. e, Treating regimen. f, Tumor volumes of doxycycline inducible Rnf2 KO tumors at the indicated days after implantation in BALB/c mice. Tumor volumes were calculated by length x width2 / 2. g, h, The weights (g) and images (h) of the tumors at the end of the study. n = 4 mice for - Dox, n = 5 mice for + Dox, each mouse harboring one tumor. i-j, Immunoblots show the selective subunits of PRC1 complex interacting with Rnf2 that was immunoprecipitated from chromatin of 4T1 cells (i) or Rnf2 interacting with Cbx4 that was immunoprecipitated from whole cell lysates of 4T1 cells (j). Ft, Flowthrough, E, Elute, which represents two independent experiments with similar results. k, Immunoblots of Rsf1/β-actin in Ctrl and Rsf1 KO (two independent gRNAs, g1 and g2) 4T1 cells. The knockout efficiency was independently confirmed at least one more time. l-q, l, m, The tumor volumes (mean ± SEM) in mice implanted with control or Rsf1 KO 4T1 tumor cells (guide 1 (n = 4 mice/group) (l) or guide 2 (n = 5 mice in control group, n = 4 mice in Rsf1 KO group) (each mouse harboring one tumor). (m). n-q, Representative tumor BLIs of tumors at day 22/47 after inoculation (n, o) or at longer time points (p, q). X, mice were sacrificed because of the big tumor burdens at the day when images were taken. r, The proliferation (mean of quadruplicates, n = 4 for technical replicates) (fold changes in cell numbers) of control or Rnf2 KO (g1 and g2) or Rsf1 KO (g1 and g2) 4T1 cells in vitro. Cell numbers at day 0 are set as 1. s-u, s. The volumes of control or Rnf2 KO 4T1 tumors (g1 and g2) implanted into the 4th mammary pads of the immuno-compromised NOD-Prkdcem26Cd52Il2rgem26Cd22/NjuCrl (NCG) mice. Mean ± SEM (n = 5 mice/group, each mouse harboring one tumor). t-u, The volumes of control tumors or Rnf2 KO 4T1 tumors ((g1) (t) or (g2) (u)) implanted into the immuno-compromised NCG mice (as in panel s) or immune competent BALB/c mice (as in Fig. 1b-c). Mean ± SEM (n = 5 mice/group, each mouse harboring one tumor). v, Cell numbers of 4T1 tumors (Ctrl, Rnf2 KO and Rsf1 KO) and of intratumoral immune cells displayed in Fig. 2a-b. n = 6 mice for Ctrl and Rnf2 KO; n = 5 mice for Rsf1 KO, each mouse harboring one tumor.Symbols, individual mouse (bars, mean ± SEM). Two-way ANOVA with Tukey’s test in b, s, t, u; unpaired two tailed Student’s t test in f, g, r; two-way ANOVA with Sidak’s test in l, m.
Extended Data Fig. 4 Rnf2/Rsf1 is negatively correlated to infiltration/activation of NK and CD4+ T cells.
a-b, BALB/c mice were inoculated with control 4T1 tumor into the left 4th mammary fat pads or simultaneously with control tumor into the left 4th mammary fat pads and Rnf2 KO 4T1 tumors (Rnf2 KO g2) into the right 4th mammary pads. a, Tumor volumes (mean ± SEM, n = 5 mice in the group of 4T1 tumor injected into the left 4th mammary fat pads (Control Tumor), each mouse harboring one tumor; n=4 mice in the group of simultaneously injected with control tumor into the left 4th mammary fat pads (Control Tumor-Abscopal) and Rnf2 KO 4T1 tumors into the right 4th mammary pads (Rnf2 KO-Abscopal), each mouse harboring two tumors: one is control tumor injected in the left mammary fat pad, the other one is Rnf2 KO tumor injected into the right mammary fat pad). p < 0.0001 (control tumors injected alone (Control Tumor) vs. Rnf2 KO tumors injected into the right mammary pads of the mice in which control tumors were injected simultaneously into the left mammary pads (Rnf2 KO-Abscopal)); p<0.0001 (control tumor injected into the left mammary pads of the mice in which Rnf2 KO tumors were injected simultaneously into the right mammary pads (Control Tumor-Abscopal) vs. Rnf2 KO tumors injected into the right mammary pads of the mice in which control tumors were injected simultaneously into the left mammary pads (Rnf2 KO-Abscopal)); not statistically significant (control tumors injected alone (Control Tumor) vs. control tumor injected into the left mammary pads of the mice in which Rnf2 KO tumors were injected simultaneously into the right mammary pads (Control Tumor-Abscopal)) (two-way ANOVA with Tukey’s test). b, Representative BLIs of tumors at indicated days. c-d, The counts of genes encoding granzymes (GZMB, GZMC, GZMD, GZME, GZMF) and mast cell proteases (MCPT1, MCP2, MCPT8), revealed in RNAseq of FACS-sorted NK cells from control 4T1 tumors compared to those in Rnf2 KO tumors (Upper) or Rsf1 KO tumors (Bottom), at day 7 after implantation. The RNAseq was performed in control tumors (n=2), Rnf2 KO tumors (n=2, Rnf2 KO g1 and g2), Rsf1 KO tumors (n=2, Rsf1 KO g1 and g2). The full list of p values can be found in the source data for this figure (two-tailed Wald test). e, The volumes of 4T1 control tumors in BALB/c mice treated with/without α-asialo GM1 at days 2, 5, 10 post-implantation. Mean ± SEM (n=5 mice/group, each mouse harboring one tumor.). Two-way ANOVA with Sidak’s test. f, The correlation of expression of NKG2D with the overall survival of invasive breast cancer patients (TCGA dataset) via cBioPortal. High, expression level > 1.5 SD above the mean. P value is generated using two-tailed LogRank test. g, The correlation of RNF2 expression to the published NK cell signature in TNBC was analyzed by Pearson correlation (two-tailed, no adjustment for multiple comparisons because of one correlation test for a gene pair). The expression levels of these genes are extracted from the TCGA dataset. The values of the coefficients (r) and significance (p) are indicated. Shaded area, 95% confidential interval. h-j, The expression levels of RNF2 and indicated genes from the published NK cell gene signature are extracted from TCGA RNAseq datasets of various cancer types (n = 11,160 patients). The correlations of RNF2 expression to the levels of these genes as a whole (h), or individually (r in i; p values in j) are shown. k, The volumes of control 4T1 tumors in BALB/c mice treated with α-CD4 (GK1.5) or its isotype control antibody at days 2, 5 post-implantation. Mean ± SEM (n = 5 mice/group, each mouse harboring one tumor.). Two-way ANOVA with Sidak’s test. l, The correlation of RNF2 to CD4 expression level in TNBC was analyzed by Pearson correlation (two-tailed, no adjustment for multiple comparisons because of one correlation test for a gene pair). The expression levels of these genes are extracted from the TCGA dataset. The values of the coefficients (r) and significance (p) of the correlations are displayed. Shaded area, 95% confidential interval. m, The correlation of CD4 expression with the overall survival of invasive breast cancer patients (TCGA dataset) via cBioPortal. P value is generated using LogRank test. High, the expression level > 2 SD above the mean. P value is generated using two-tailed LogRank test. n-p, The expression levels of RNF2 and genes encoding T cell markers and MHCII are extracted from TCGA RNAseq datasets of various cancer types (n = 11,160 patients). The correlations of RNF2 expression to the levels of these genes as a whole (n) or individually (r in o; p values in p) are shown. q, Frequencies of peripheral blood CD8+ T cells in mice bearing control or Rnf2 KO tumors at day 10 after injection of anti-CD8 (2.43) or control antibody. Symbols depict individual mouse (bars, mean ± SEM). n = 5 mice/group, each mouse harboring one tumor. One-way ANOVA with Tukey’s test.
Extended Data Fig. 5 Tumoral NKG2DL and NK/CD4+ T cells expressed IFNγR are required for these immune cells activation stimulated by Rnf2 KO tumor.
a, The frequencies of NKG2DL+CD45− tumor cells isolated from Ctrl/Rnf2 KO/Rsf1 KO 4T1 tumors (Left, n = 5 mice for Ctrl, n = 4 mice for Rnf2 KO and Rsf1 KO, each mouse harboring one tumor) or EMT6 tumors (Right, n = 5 mice for Ctrl and Rnf2 KO g1, n = 4 mice for Rnf2 KO g2, each mouse harboring one tumor) at day 7 post-implantation. Symbols, individual mouse (bars, mean ± SEM). b, Tumor cells (CD45–) were isolated and enriched from indicated tumors, co-cultured with NK/CD4+ T cells/both. Anti-NKG2DL/control antibody was added into the co-culture. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+NK cells (Left) or IFNγ+CD4+ T-cells (Right) are shown as mean ± SEM of triplicates. c, Frequencies of intratumoral NKG2D+ cells of CD4+ T effector cells of control, Rnf2 KO and Rsf1 KO 4T1 tumors at day 7 post-implantation. Symbols, individual mouse (bars, mean ± SEM). n = 5 mice/group, each mouse harboring one tumor. d, In vitro co-culture experiment was set up as that shown in panel b, except the addition of anti-NKG2DL antibody. Instead, CD4+ T-cells were pre-incubated with anti-NKG2D antibody for 30 min before being added into the co-culture. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+CD4+ T-cells (Left) or IFNγ+NK cells (Right) are shown as mean ± SEM of triplicates. e, Tumor cells (GFP+) were isolated and enriched by FACS for GFP positive populations from indicated 4T1 tumors in mice. They were co-cultured with NK/CD4+ T cells/both. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+NK cells (Left) or IFNγ+CD4+ T cells (Right) are shown as mean ± SEM of triplicates. f, Tumor cells (CD45–) were enriched from indicated 4T1 tumors in mice and co-cultured with pre-activated NK/CD4+ T-cells/both. CD4+ T-cells were pre-incubated with anti-NKG2D antibody for 30 min before being added into the co-culture. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). The percent tumor killing is shown as mean ± SEM of triplicates. g-h, The frequencies of MHCI+CD45− tumor cells and/or the MFI of MHCI expression on CD45– tumor cells isolated from control (Ctrl)/Rnf2 KO cells of 4T1 tumors (g, n = 4 mice for Ctrl, n = 5 mice for Rnf2 KO, each mouse harboring one tumor) or EMT6 tumors (h, n = 5/group) at day 7 post-implantation. Symbols depict individual mouse (bars, mean ± SEM). i-j, Tumor cells (CD45–) were enriched from indicated 4T1 tumors in mice, co-cultured with NK/CD4+ T-cells/both. CD4+ T-cells (i) or NK cells (j) were pre-incubated with anti-IFNγ receptor (IFNγR) antibody (GR-20) for 30 min before being added into the co-culture. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+CD4+ T-cells or IFNγ+NK cells are shown as mean ± SEM of triplicates. k-m, Expression levels of IFNγR1 and IFNγR2 in Rnf2 KO plus IFNγR1 KO (DKO 1) (k), Rnf2 KO plus IFNγR2 KO (DKO2) (l), and Rnf2 KO plus IFNγR1 KO and IFNγR2 KO (TKO) (m) 4T1 cells, measured by Flowcytometry (Histogram), which represents two independent experiments. n-o, The volumes (n) and representative BLIs at day 49 (o) of indicated 4T1 tumors in BALB/c mice. Mean ± SEM (n = 5 mice/group). DKO 1: Rnf2 KO + IFNγR1 KO; DKO 2: Rnf2 KO + IFNγR2 KO; TKO: Rnf2 KO + IFNγR1 KO + IFNγR2 KO. ****, control tumor vs. Rnf2 KO + IFNγR1 KO tumor; ****, control tumor vs. Rnf2 KO + IFNγR2 KO tumor; ****, control tumor vs. Rnf2 KO + IFNγR1 KO + IFNγR2 KO tumor; ****, control tumor vs. Rnf2 KO tumor. p, Tumor cells (CD45–) were enriched from indicated tumors in mice, co-cultured with NK/CD4+ T-cells/both. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+CD4+ T-cells (Left) or IFNγ+NK cells (Right) are shown as mean ± SEM of triplicates. q-r, q, The volumes of control 4T1 tumors in BALB/c mice treated with control/anti-IFNγ antibody at day 2, 5 post-implantation. Mean ± SEM (n = 5 mice/group, each mouse harboring one tumor). r, The relative ratio of the fold changes in the tumor volumes (=Average of Fold Changes in tumor volumes of the group of anti-IFNγ divided by Average of Fold Changes in tumor volumes of the group of Control antibody) in control 4T1 tumors (panel q) or Rnf2 KO tumors (shown in Fig. 3h) at the end of the study. n = 5 mice for Ctrl Tumor Ctrl Antibody; n=4 mice for Ctrl Tumor IFNγ Antibody; n=4 mice for Rnf2 KO Ctrl Antibody; n=5 mice for Rnf2 KO IFNγ Antibody. Each mouse harbored one tumor. s, Tumor cells (CD45–) were enriched from indicated 4T1 tumors in mice, co-cultured with NK/CD4+ T-cells/both. Control/anti-IL-2 antibody (JES6-1A12) was supplemented in the co-culture. Tumor cells (Ctrl/Rnf2 KO) in each replicate were pooled from 2-3 tumors from 2-3 mice (each mouse harboring one tumor). Frequencies of IFNγ+CD4+ T-cells (Left) or IFNγ+NK cells (Right) are shown as mean ± SEM of triplicates. *p<0.05, **p<0.01, ***p<0.001,****p<0.0001, n.s., not statistically significant (unpaired two-tailed Student’s t test in a, b, c, f, g, h, p; one-way ANOVA with Tukey’s test in d, e, i, j, s; two-way ANOVA with Tukey’s test in n; two-way ANOVA with Sidak’s test in q). The full list of p values can be found in the source data for this figure.
Extended Data Fig. 6 Rnf2/Rsf1 KO tumors upregulate genes related to immunity.
a, Log2FC (fold change) (Left) and adjusted p values (padj) (Right) of the upregulated immune cell markers expressed in Rnf2 KO 4T1 tumor cells (n = 2, Rnf2 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice, each mouse harboring one tumor) compared to those in control tumor cells (n = 2, two groups of cells, each group of cells pooled from 25 mice, each mouse harboring one tumor) isolated and sorted from in vivo tumors at day 7 post-implantation followed by RNAseq analysis, as in Fig. 4a. b-e, Analysis of differentially expressed genes revealed in RNAseq of FACS-sorted Rnf2 KO 4T1 tumor cells (n = 2, Rnf2 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice) (b), or Rsf1 KO 4T1 tumor cells (n = 2, Rsf1 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice) (c) compared to control 4T1 tumors cells (n = 2, two groups of cells, each group of cells pooled from 25 mice) isolated from in vivo tumors at day 7 post-tumor implantation, using NetworkAnalyst. The representative enriched gene sets with FDR q values are shown (b, c). Heatmaps display these DEGs and representative genes related to immunity are noted (d, e). f, GSEA analysis of differentially expressed genes (DEGs) revealed in RNAseq of FACS-sorted Rsf1 KO 4T1 tumor cells (n = 2, Rsf1 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice) compared to control 4T1 tumors cells (n = 2, two groups of cells, each group of cells pooled from 25 mice) isolated from corresponding tumors implanted in mice at day 7 post-tumor implantation. The representative enriched gene sets with FDR q values are shown. g, Representative GSEA Enrichment plots (score curves) of DEGs in Rnf2 KO compared to control tumors. GSEA analysis of DEGs revealed in RNAseq of 4T1 tumors is performed as in Fig. 4a. The representative Enrichment plots of positively or negatively regulated genes in Rnf2 KO tumor cells compared to control tumor cells are shown.
Extended Data Fig. 7 Rsf1 regulates expression of an overlapping group of immune-related genes with Rnf2.
a, Representative GSEA Enrichment plots (score curves) of DEGs in Rsf1 KO compared to control tumors. GSEA analysis of DEGs revealed in RNAseq of 4T1 tumors is performed as in Extended Fig. 6f. The representative Enrichment plots are shown. b, Analysis of DEGs revealed in published RNAseq of control siRNA- compared to RNF2 siRNA-treated human TNBC cell line MDA-MB-231 (log2FC≥ 1 and log2FC≤ -1) using Network Analyst. The representative enriched gene sets with FDR q values are shown. c, The overlap of DEGs in Rnf2 KO and Rsf1 KO 4T1 tumor cells compared to control 4T1 tumors cells, which were isolated by FACS from in vivo tumors. DEGs are determined as in Fig. 4a and Extended Fig. 6f. P value of overlap is calculated using web tool SSOTGNB (http://nemates.org/MA/progs/overlap_stats.html). d, The log2FC (Fold Change) of overlapping DEGs in Rnf2 KO tumors and Rsf1 KO tumors. e. g:Profiler analysis of the 134 overlapped DEGs in panel c. The adjust p value (padj) is indicated (one-tailed hypergeometric test with adjustment for multiple comparisons). f, A floating bars graph (min, max and line at mean) depicts the counts of genes encoding chemokines revealed in RNAseq of FACS-sorted tumors cells, as in Fig. 4a and Extended Data Fig. 6f. These Rnf2 KO (n = 2, Rnf2 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice, each mouse harboring one tumor), Rsf1 KO (n = 2, Rsf1 KO g1 and g2, two groups of cells, each group of cells pooled from 25 mice, each mouse harboring one tumor) and control (n = 2, two groups of cells, each group of cells pooled from 25 mice, each mouse harboring one tumor) 4T1 tumors cells were isolated by FACS from corresponding in vivo tumors. The adjust p values (padj) are determined by the two-tailed Wald test and can be found in the source data for this figure. g, The correlations of expressions of RNF2, RSF1 or EZH2 to the levels of human chemokine genes were analyzed using the RNAseq data extracted from invasive breast cancer TCGA dataset. Heatmaps show the correlation coefficients (Left) and q values of each correlation (Right). h, The correlations of the chemokine gene CCL5 or CXCL10 to the overall survival of invasive breast cancer patients, extracted from the TCGA dataset via cBioPortal. High expression is defined as the expression levels greater than 2 SD above the mean. P values are generated using two-tailed LogRank test.
Extended Data Fig. 8 Rnf2 binds to immune related genes in mouse and human breast cancer cells.
a-b, The overlap of Rnf2 target genes in cultured 4T1 cells determined by Rnf2 CHIPseq with genes displaying significantly more/less accessible chromatin sites in Rnf2 KO 4T1 tumors determined by ATACseq as in Fig. 5a (a) or with DEGs (determined by RNAseq shown in Fig. 4a) in Rnf2 KO 4T1 tumors (compared to control 4T1 tumors) (b). P values of the overlap are calculated using web tool SSOTGNB (http://nemates.org/MA/progs/overlap_stats.html). c-d, Screenshots of genes H2-Ab1 and H2-Eb1 in both control and Rnf2 KO 4T1 tumor cells obtained from RNAseq and ATACseq (c) or of gene H2-Ab1 in mouse embryonic stem cells (mES) obtained by reanalyzing published datasets of Rnf2 CHIPseq (d). e, Expression level of gene H2-Ab1 in control and Rnf2 KO mES cells obtained by reanalyzing published dataset of RNA microarray. f, Genes occupied by RNF2 were identified by CHIPseq analysis of human TNBC cell line MDA-MB-231 using anti-RNF2 antibody compared to control antibody. g, Integrative genomics viewer (IGV) screenshots of control IgG or RNF2 ChIPseq (as in panel f) tracks (scale bar, 40) of HLA-DPA1, HLA-DPB1, CCL20 and CXCL8. h, GSEA analysis of genes occupied by RNF2 revealed in published CHIPseq of human TNBC cell line MDA-MB-231 using anti-RNF2 antibody compared to control antibody (as in panel f). The representative enriched pathways with FDR q values are shown. i, The correlations of RNF2 expression in BRCA patients to the levels of its bound genes that were determined as in panel f. The expression levels of these RNF2 bound genes and of RNF2 were extracted from the invasive breast cancer TCGA dataset. The coefficient (yellow) and q value (purple) of each correlation are indicated at the y-axis (two-tailed Spearman correlation analysis). The number (n) of RNF2 bound genes with FDR q values (indicative of the significance of the correlations of these genes to RNF2 expression) < 0.05, and the percentages of these genes (q < 0.05) among the total RNF2 bound genes are noted in blue. The blue vertical bars mark the q value at 0.05. n = 1,070 patients.
Extended Data Fig. 9 Ablation of Rnf2/Rsf1 induces antitumor memory response.
a, BALB/c mice were inoculated with Rnf2 KO 4T1 tumors (Rnf2 KO g2) or Rsf1 KO 4T1 tumors (Rsf1 KO g1) in the left 4th mammary pads. At day 45 after the primary tumors were rejected, wildtype 4T1 tumors were implanted into the right 4th mammary pads of these mice or of naïve mice. n=5 mice /group, each mouse harboring one tumor. Representative BLIs of wildtype 4T1 tumors at days 97, 131, 182 and 307 after the 2nd challenge are shown. X, mice were sacrificed because of the big tumor burdens at the day when images were taken. b, BALB/c mice were inoculated with Rnf2 KO 4T1 tumors (Rnf2 KO g2) in the left 4th mammary pads. At day 45, TSA tumors were implanted into the right 4th mammary pads of these mice or of naïve mice. The tumor volumes were measured by caliper (mean ± SEM, n = 5 mice for naïve, n=4 mice for Rnf2 KO, each mouse harboring one tumor). c, Frequencies of intratumoral indicated immune cells in wildtype 4T1 tumors at day 7 after the 2nd challenge. d-e, Mice were inoculated with Rnf2 KO 4T1 tumors and re-challenged with wildtype 4T1 tumors on the contralateral, as in panel a and Fig. 8a. α-asialo GM1 antibody was injected at days −1, +1, +4 relative to the 2nd challenge. d, Volumes (Left, mean ± SEM, n = 5 mice/group, each mouse harboring one tumor ; Right, individual mouse) of wildtype 4T1 tumors. e,. Representative BLIs of wildtype 4T1 tumors at indicated days after the 2 challenge. f, Frequencies of peripheral blood CD8+ T-cells in Rnf2 KO mice at day 10 after injection of anti-CD8 (2.43) or control antibody. Symbols, individual mouse (bars, mean ± SEM). n = 5 mice/group, each mouse harboring one tumor. g-h, g, Mice were inoculated with Rnf2 KO 4T1 tumors (Rnf2 KO g2) and re-challenged with wildtype 4T1 tumors on the contralateral, as in panel a and Fig. 8a. Anti-CD8 or control antibody was injected at days -1 and +1 relative to the 2nd challenge. g, Volumes (Left, mean ± SEM, n = 5 mice/group, each mouse harboring one tumor ; Right, individual mouse) of wildtype 4T1 tumors. h, Representative BLIs of wildtype 4T1 tumors at day 26 or day 96 after the 2nd challenge. i, Frequencies of peripheral blood CD4+ T-cells in Rnf2 KO mice at day 10 after injection of anti-CD4 (GK1.5) or control antibody. Symbols, individual mouse (bars, mean ± SEM). n = 5 mice/group, each mouse harboring one tumor. Unpaired two-tailed Student’s t test in c, f, i; two-way ANOVA with Sidak’s test in b, d, g.
Extended Data Fig. 10 Gating Strategy used for flow cytometry.
a, Gating Strategy used for analysis of immune cells from tumors isolated from tumor-bearing mice presented on Fig. 2a–e, Fig. 3a, b, Fig. 4c, Fig. 7e, Fig. 8e, Extended Data Fig. 3v, 5a, c, g, h, Extended Data Fig. 9c. b, Gating Strategy used for analysis of each subset of CD4+ T-cells in a (5-2). c, Gating Strategy used for analysis of each subset or marker of NK cells in a (5-3). d, Gating Strategy used for analysis of each subset or marker of CD45− tumor cells in a. e, Gating Strategy used for analysis of CD4+ T-cells or NK cells from the in vitro co-culture assays presented on Fig. 3c, d, f, g, Fig. 4j, k, Extended Data Fig. 5b, d, e, Extended Data Fig. 5i–j, p, s. f, Gating Strategy used for analysis of dead tumors in the in vitro killing assays presented on Fig. 3e, Fig. 4l, Extended Data Fig. 5f. g, Gating Strategy used for sorting of tumor cells (CD45−) and NK cells (CD45+NKp46+CD3−) for RNA-Seq and ATAC-Seq presented on Fig. 4a, b, Fig. 5a–f, Fig. 6, Fig. 7f-j, Extended Data Figs.4c,d, 6, 7a, c–f, 8a–c.
Supplementary information
Supplementary Tables
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1
Numerical data for Fig. 1.
Source Data Fig. 1
Unprocessed blots for Fig. 1.
Source Data Fig. 2
Numerical data for Fig. 2.
Source Data Fig. 3
Numerical data for Fig. 3.
Source Data Fig. 4
Numerical data for Fig. 4.
Source Data Fig. 5
Numerical data for Fig. 5.
Source Data Fig. 6
Numerical data for Fig. 6.
Source Data Fig. 7
Numerical data for Fig. 7.
Source Data Fig. 7
Unprocessed blots for Fig. 7.
Source Data Fig. 8
Numerical data for Fig. 8.
Source Data Extended Data Fig. 1
Numerical data for Extended Fig. 1.
Source Data Extended Data Fig. 2
Numerical data for Extended Fig. 2.
Source Data Extended Data Fig. 3
Numerical data for Extended Fig. 3.
Source Data Extended Data Fig. 3
Unprocessed blots for Extended Fig. 3.
Source Data Extended Data Fig. 4
Numerical data for Extended Fig. 4.
Source Data Extended Data Fig. 5
Numerical data for Extended Fig. 5.
Source Data Extended Data Fig. 6
Numerical data for Extended Fig. 6.
Source data Extended Data Fig. 7
Numerical Data for Extended Fig. 7
Source Data Extended Data Fig. 8
Numerical data for Extended Fig. 8.
Source Data Extended Data Fig. 9
Numerical data for Extended Fig. 9.
Rights and permissions
About this article
Cite this article
Zhang, Z., Luo, L., Xing, C. et al. RNF2 ablation reprograms the tumor-immune microenvironment and stimulates durable NK and CD4+ T-cell-dependent antitumor immunity. Nat Cancer 2, 1018–1038 (2021). https://doi.org/10.1038/s43018-021-00263-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00263-z
This article is cited by
-
RNF2 inhibits E-Cadherin transcription to promote hepatocellular carcinoma metastasis via inducing histone mono-ubiquitination
Cell Death & Disease (2023)
-
CD4+ T cells in cancer
Nature Cancer (2023)
-
Development and validation of an integrative pan-solid tumor predictor of PD-1/PD-L1 blockade benefit
Communications Medicine (2023)