Abstract
Only a subset of recurrent glioblastoma (rGBM) responds to anti-PD-1 immunotherapy. Previously, we reported enrichment of BRAF/PTPN11 mutations in 30% of rGBM that responded to PD-1 blockade. Given that BRAF and PTPN11 promote MAPK/ERK signaling, we investigated whether activation of this pathway is associated with response to PD-1 inhibitors in rGBM, including patients that do not harbor BRAF/PTPN11 mutations. Here we show that immunohistochemistry for ERK1/2 phosphorylation (p-ERK), a marker of MAPK/ERK pathway activation, is predictive of overall survival following adjuvant PD-1 blockade in two independent rGBM patient cohorts. Single-cell RNA-sequencing and multiplex immunofluorescence analyses revealed that p-ERK was mainly localized in tumor cells and that high-p-ERK GBMs contained tumor-infiltrating myeloid cells and microglia with elevated expression of MHC class II and associated genes. These findings indicate that ERK1/2 activation in rGBM is predictive of response to PD-1 blockade and is associated with a distinct myeloid cell phenotype.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Single-cell RNA-seq data supporting the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code nos. GSE103224 and GSE141383. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
Code availability
Code is available at https://github.com/RabadanLab/GBMsinglecell.
Change history
11 February 2022
A Correction to this paper has been published: https://doi.org/10.1038/s43018-022-00343-8
References
Stupp, R. et al. Effect of tumor-treating fields plus maintenance temozolomide vs maintenance temozolomide alone on survival in patients with glioblastoma: a randomized clinical trial. JAMA 318, 2306–2316 (2017).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56 (2017).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849 (2019).
Gong, J., Chehrazi-Raffle, A., Reddi, S. & Salgia, R. Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: a comprehensive review of registration trials and future considerations. J. Immunother. Cancer 6, 8 (2018).
Reardon, D. A. et al. Effect of nivolumab vs bevacizumab in patients with recurrent glioblastoma: the CheckMate 143 Phase 3 randomized clinical trial. JAMA Oncol. 6, 1003–1010 (2020).
Nayak, L. et al. Randomized phase II and biomarker study of pembrolizumab plus bevacizumab versus pembrolizumab alone for recurrent glioblastoma patients. Clin. Cancer Res. 27, 1048–1057 (2021).
de Groot, J. et al. Window-of-opportunity clinical trial of pembrolizumab in patients with recurrent glioblastoma reveals predominance of immune-suppressive macrophages. Neuro. Oncol. 22, 539–549 (2020).
Arrieta, V. A. et al. The possibility of cancer immune editing in gliomas. A critical review. Oncoimmunology 7, e1445458 (2018).
Chongsathidkiet, P. et al. Sequestration of T cells in bone marrow in the setting of glioblastoma and other intracranial tumors. Nat. Med. 24, 1459–1468 (2018).
Jackson, C. M., Choi, J. & Lim, M. Mechanisms of immunotherapy resistance: lessons from glioblastoma. Nat. Immunol. 20, 1100–1109 (2019).
Arrieta, V. A. et al. Can patient selection and neoadjuvant administration resuscitate PD-1 inhibitors for glioblastoma? J. Neurosurg. 132, 1667–1672 (2019).
Woroniecka, K. et al. T-cell exhaustion signatures vary with tumor type and are severe in glioblastoma. Clin. Cancer Res. 24, 4175–4186 (2018).
Zhao, J. et al. Immune and genomic correlates of response to anti-PD-1 immunotherapy in glioblastoma. Nat. Med. 25, 462–469 (2019).
Cloughesy, T. F. et al. Neoadjuvant anti-PD-1 immunotherapy promotes a survival benefit with intratumoral and systemic immune responses in recurrent glioblastoma. Nat. Med. 25, 477–486 (2019).
Dhillon, A. S., Hagan, S., Rath, O. & Kolch, W. MAP kinase signalling pathways in cancer. Oncogene 26, 3279–3290 (2007).
Ruess, D. A. et al. Mutant KRAS-driven cancers depend on PTPN11/SHP2 phosphatase. Nat. Med. 24, 954–960 (2018).
Svensson, S. et al. ERK phosphorylation is linked to VEGFR2 expression and Ets-2 phosphorylation in breast cancer and is associated with tamoxifen treatment resistance and small tumours with good prognosis. Oncogene 24, 4370–4379 (2005).
Roth, P., Valavanis, A. & Weller, M. Long-term control and partial remission after initial pseudoprogression of glioblastoma by anti-PD-1 treatment with nivolumab. Neuro. Oncol. 19, 454–456 (2017).
Okada, H. et al. Immunotherapy response assessment in neuro-oncology: a report of the RANO working group. Lancet Oncol. 16, e534–e542 (2015).
Vassilakopoulou, M. et al. Preanalytical variables and phosphoepitope expression in FFPE tissue: quantitative epitope assessment after variable cold ischemic time. Lab. Invest. 95, 334–341 (2015).
Nagasawa-Masuda, A. & Terai, K. ERK activation in endothelial cells is a novel marker during neovasculogenesis. Genes Cells 21, 1164–1175 (2016).
Pombo Antunes, A. R. et al. Single-cell profiling of myeloid cells in glioblastoma across species and disease stage reveals macrophage competition and specialization. Nat. Neurosci. 24, 595–610 (2021).
Hara, T. et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell 39, 779–792 (2021).
Yuan, J. et al. Single-cell transcriptome analysis of lineage diversity in high-grade glioma. Genome Med. 10, 57 (2018).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl Acad. Sci. USA 113, E1738–E1746 (2016).
Sankowski, R. et al. Mapping microglia states in the human brain through the integration of high-dimensional techniques. Nat. Neurosci. 22, 2098–2110 (2019).
Friebel, E. et al. Single-cell mapping of human brain cancer reveals tumor-specific instruction of tissue-invading leukocytes. Cell 181, 1626–1642 (2020).
Klemm, F. et al. Interrogation of the microenvironmental landscape in brain tumors reveals disease-specific alterations of immune cells. Cell 181, 1643–1660 (2020).
Saha, D., Martuza, R. L. & Rabkin, S. D. Macrophage polarization contributes to glioblastoma eradication by combination immunovirotherapy and immune checkpoint blockade. Cancer Cell 32, 253–267 (2017).
Goswami, S. et al. Immune profiling of human tumors identifies CD73 as a combinatorial target in glioblastoma. Nat. Med. 26, 39–46 (2020).
Rao, G. et al. Anti-PD-1 induces M1 polarization in the glioma microenvironment and exerts therapeutic efficacy in the absence of CD8 cytotoxic T cells. Clin. Cancer Res. 26, 4699–4712 (2020).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Schalper, K. A. et al. Neoadjuvant nivolumab modifies the tumor immune microenvironment in resectable glioblastoma. Nat. Med. 25, 470–476 (2019).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Hellmann, M. D. et al. Nivolumab plus Ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 378, 2093–2104 (2018).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Johanns, T. M. et al. Immunogenomics of hypermutated glioblastoma: a patient with germline POLE deficiency treated with checkpoint blockade immunotherapy. Cancer Discov. 6, 1230–1236 (2016).
Lukas, R. V. et al. Clinical activity and safety of atezolizumab in patients with recurrent glioblastoma. J. Neurooncol. 140, 317–328 (2018).
Bouffet, E. et al. Immune checkpoint inhibition for hypermutant glioblastoma multiforme resulting from germline biallelic mismatch repair deficiency. J. Clin. Oncol. 34, 2206–2211 (2016).
Ahmad, H., Fadul, C. E., Schiff, D. & Purow, B. Checkpoint inhibitor failure in hypermutated and mismatch repair-mutated recurrent high-grade gliomas. Neurooncol. Pract. 6, 424–427 (2019).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Touat, M. et al. Mechanisms and therapeutic implications of hypermutation in gliomas. Nature 580, 517–523 (2020).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Wick, W. et al. Lomustine and bevacizumab in progressive glioblastoma. N. Engl. J. Med. 377, 1954–1963 (2017).
Azoulay, M. et al. Benefit of re-operation and salvage therapies for recurrent glioblastoma multiforme: results from a single institution. J. Neurooncol. 132, 419–426 (2017).
Robert, K. J. et al. CD8+ T-cell-mediated immunoediting influences genomic evolution and immune evasion in murine gliomas. Clin. Cancer Res. 26, 4390–4401 (2020).
Chen, P. et al. Symbiotic macrophage–glioma cell interactions reveal synthetic lethality in PTEN-null glioma. Cancer Cell 35, 868–884 (2019).
Hutter, G. et al. Microglia are effector cells of CD47-SIRPα antiphagocytic axis disruption against glioblastoma. Proc. Natl Acad. Sci. USA 116, 997–1006 (2019).
Strauss, L. et al. Targeted deletion of PD-1 in myeloid cells induces antitumor immunity. Sci. Immunol. 5, eaay1863 (2020).
Gromeier, M. et al. Very low mutation burden is a feature of inflamed recurrent glioblastomas responsive to cancer immunotherapy. Nat. Commun. 12, 352 (2021).
Liu, D. et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat. Med. 25, 1916–1927 (2019).
Alspach, E. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395 (2018).
Mundt, S. et al. Conventional DCs sample and present myelin antigens in the healthy CNS and allow parenchymal T cell entry to initiate neuroinflammation. Sci. Immunol. 4, eaau8380 (2019).
Qian, J. et al. TLR2 promotes glioma immune evasion by downregulating MHC class II molecules in microglia. Cancer Immunol. Res. 6, 1220–1233 (2018).
Schartner, J. M. et al. Impaired capacity for upregulation of MHC class II in tumor-associated microglia. Glia 51, 279–285 (2005).
Sarkar, S. et al. Therapeutic activation of macrophages and microglia to suppress brain tumor-initiating cells. Nat. Neurosci. 17, 46–55 (2014).
Sarkar, S. et al. Microglia induces Gas1 expression in human brain tumor-initiating cells to reduce tumorigenecity. Sci. Rep. 8, 15286 (2018).
Zhang, J. et al. Compartmental analysis of T-cell clonal dynamics as a function of pathologic response to neoadjuvant PD-1 blockade in resectable non-small cell lung cancer. Clin. Cancer Res. 26, 1327–1337 (2020).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Hothorn, T., Hornik, K. & Zeileis, A. Unbiased recursive partitioning: a conditional inference framework. J. Comput. Graph. Stat. 15, 651–674 (2006).
Hothorn, T. & Zeileis, A. partykit: A modular toolkit for recursive partytioning in R. https://www2.uibk.ac.at/downloads/c4041030/wpaper/2014-10.pdf (2015).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 3, 861 (2018).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Acknowledgements
This work was supported by NIH grant nos. 1R01NS110703-01A1 and 5DP5OD021356-05 (both A.M.S.); no. P50CA221747 SPORE for Translational Approaches to Brain Cancer (principal investigator M. S. Lesniak, with support to A.M.S.); developmental funds from the Robert H. Lurie Cancer Center Support Grant (no. P30CA060553, A.M.S.); Vagelos Precision Medicine Award (F.M.I., R.R. and J.Z.), U54CA193313 (R.R.), U54CA209997 (R.R.), R35CA253126 (R.R.); Keep Punching (F.M.I.); The William Rhodes and Louise Tilzer-Rhodes Center for Glioblastoma at New York-Presbyterian Hospital (F.M.I.); NIAID nos. 1R01AI099195 (U.B.) and R01AI134988 (U.B.). V.A.A. is financially supported by the Mexican government through the Mexican National Council for Science and Technology and the Plan of Combined Studies in Medicine of the National Autonomous University of Mexico. A.X.C. is funded by the Medical Scientist Training Program (no. T32GM007367). We thank T. Sudhakar for sample collection at Columbia University; K. McCortney, R. Javier and J. Walshon from the Nervous System Tumor Bank supported by the P50CA221747 SPORE for Translational Approaches to Brain Cancer; and B. Shmaltsuyeva for immunohistochemistry at the Northwestern University Pathology Core Facility funded by Cancer Center Support Grant (no. NCI CA060553). We thank L. Kai for technical support on multiplex staining performed at the Immunotherapy Assessment Core at Northwestern University and in the Flow Cytometry & Cellular Imaging Core Facility, which is supported in part by NIH through MD Anderson´s Cancer Center Support Grant no. CA016672, the NCI´s Research Specialist 1 (no. R50 CA23707, J.K.B.) and NIH grant nos. R01-CA120813 and 1R01-CA237418 (both A.B.H.).
Author information
Authors and Affiliations
Contributions
V.A.A., A.X.C. and J.R.K. performed the majority of experiments and analyses. V.A.A., J.R.K., R.R., F.M.I. and A.M.S. conceptualized and designed the study. V.A.A. and L.A.D.C. quantified IHC images. V.A.A., A.X.C. and A.M.S. wrote the manuscript. R.S., P.U., J.Z., R.V.L., C.D., D.C., X.L., A.G., S.J.K., J.S., D.Z., J.N.B. and J.T.Y. compiled the clinical data for analysis. T.F.C. and R.P. provided tumor samples and clinical data for the validation cohort. C.A. and L.C. provided administrative support. V.A.A., A.X.C., K.B.B. and H.Z. performed survival and statistical analyses. P.C. and J.N.B. are responsible for the Tumor Bank at Columbia University. C.H. is responsible for the Tumor Bank at Northwestern University. P.U. acquired tumor specimens. C.H., M.M., D.J.B. and P.C. scored the tumor slides. J.Y., W.Z. and P.S. provided scRNA-seq data. A.X.C., J.Z. and R.R. performed scRNA-seq data analysis. D.J., B.Z., C.K., J.K.B., X.L. and A.B.H. performed Opal multiplex tissue staining. V.A.A., A.X.C. and C.L.-C. analyzed multiplex immunofluorescence images. V.A.A., C.L.-C., B.L., G.R. and U.B. acquired and analyzed flow cytometry data. V.A.A. and S.J.K. performed immunoblots and the peptide competition assay. A.M.S., R.R. and F.M.I. supervised the entire study.
Corresponding authors
Ethics declarations
Competing interests
F.M.I. is a consultant for Merck, Novocure, Regeneron, Abbvie, Alexion, Guidepoint and Tocagen. He has received clinical or research support (including equipment or material) from Bristol Myers Squibb and Merck. A.M.S., F.M.I. and R.R. are co-authors for the following patent filed by Columbia University: Systems and methods for predicting clinical responses to immunotherapies. A.M.S., V.A.A, F.M.I. and R.R. are co-authors of the following patent filed by Northwestern University: 5. Methods for treating glioblastoma. A.M.S. has received a consulting honorarium from Abbvie. R.R. is a member of the SAB of AimedBio and a founder of Genotwin. R.V.L. has received support from Roche-Genentech for meeting travel to present study results; honoraria for advisory boards for AstraZeneca, Abbvie and Ziopharm; and honoraria for medical editing for EBSCO publishing, Medlink Neurology and American Physician Institute. He has also received honoraria for consultation with Eisai and Abbvie, and honoraria for creating and presenting CME Board review material for the American Physician Institute. R.V.L. received drug support (but no additional support) from BMS for an investigator-initiated trial. A.B.H. is a consultant for Caris Life Sciences and WCG Oncology Advisory Board, receives royalties and milestone payments from Celldex Therapeutics and DNAtrix and receives clinical or research support (including equipment and materials) from Celularity, Codiak Biosciences, Moleculin and Carthera. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks Robert Manguso and Ignacio Melero for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of the staining technique and quantification of p-ERK.
a, Titration of the p-ERK antibody (clone: D13.14.4E) using different dilutions performed in GBM samples. We show the same region of a GBM sample stained with the indicated dilutions of the p-ERK antibody with a low and high magnification image for each dilution. b, (left) Microarray containing breast cancer tissues stained with p-ERK antibody (1:500 dilution) representing a positive control. (right) Magnification of one the breast cancer tissues showing specific staining in the endothelium (red rectangle). c, (left) Nontumoral brain tissue stained with p-ERK antibody (1:500 dilution) representing a negative control. (right) Magnification of the white matter showing p-ERK staining with minimal background. Dilution titration and staining of positive and negative controls were performed as a single experiment in one standardized run. d, Workflow used for the software-based quantification of p-ERK+ cells.
Extended Data Fig. 2 Quantification and cut point optimization of p-ERK+ cell density in tumoral regions.
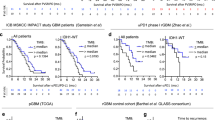
a, Dot plot showing the distribution of p-ERK quantification of all GBM samples treated and nontreated with PD-1 blockade N = 62 tumors). b, From top to bottom, micrographs showing one high p-ERK tumor sample and two low p-ERK tumor samples with positive staining in the endothelial cells (red arrows). In the dot plot, the magenta dot represents CU100 patient, the green dot represents NU01688 patient, and the red dot represents CU110 patient. IHC images are representative of 62 independent GBM samples. c, Conditional inference trees analysis for cut-point optimization in the GBM cohort treated with PD-1 blockade reveals a cut-point value similar to the median of all tumor samples. d, Forest plot representing the univariable analysis using a Cox regression model evaluating the clinical and molecular prognostic factors that might confound the association between survival p-ERK and presented as Hazard ratio (95% CI). N = 29 GBM patients. P value by two-sided Wald test. e, Kaplan-Meier curve comparing OS of recurrent GBM patients scored as either high or low p-ERK by assessment of a neuropathologist counting from initiation of PD-1 blockade (anti-PD-1 therapy group, N = 29 GBM patients) and from surgery at recurrence (no-immunotherapy group, N = 33 GBM patients). p-ERK scores in tumor regions were designated as follows: 0-1 were considered as low, and 2–3 as high; P value by two-sided log-rank test.
Extended Data Fig. 3 Preservation of the p-ERK epitope and peptide competition assay neutralizing the p-ERK1/2 antibody tested in FFPE GBM samples.
a, Protein extraction from FFPE GBM tissues for assessment of selected phosphoproteins. b, Western blot targeting p-ERK, ERK1, ERK2, p-AKT, AKT, and β-actin in a subset of GBM samples used for survival analysis. Western blotting was done as a single experiment in 12 independent GBM samples and 2 GBM cell lines. c, Peptide competition assay in which p-ERK1/2 antibody was neutralized with a blocking peptide employing extracted proteins obtained from GBM samples. The peptide competition assay was assessed by western blot. One western blot was incubated with the neutralized p-ERK antibody and the other with the free p-ERK antibody. d, Peptide competition assay employing IHC using the same GBM samples used to perform western blot employing the neutralized and free p-ERK antibody to perform the staining. The experiments were done in 4 independent GBM samples and 2 GBM cell lines as a single experiment.
Extended Data Fig. 4 Evaluation of the ischemic time on p-ERK degradation by IHC and western blot, and comparison to the samples used in this study.
a, Representatives images of the analysis conducted to evaluate p-ERK degradation in endothelial cells of GBM samples at different periods of ischemic time. For this, 3 human tumor specimens were obtained during surgery, and immediately divided into similar size portions, which then were subjected to different ischemic times before processing. Specific endothelial cells subjected to analysis are labeled with colors assigned by the software. b, Blue bars represent p-ERK+ cells mm2 in tumor regions, and dots represent p-ERK intensity on individual endothelial cells within the same samples used to evaluate the effect of ischemic time on p-ERK degradation, and tumor samples used for survival analysis (PD-1 immunotherapy cohort and no immunotherapy cohort). Each dot represents one ROI analyzing one endothelial cell. Green dots (N = 24, 20, 13 endothelial cells from NU02608, NU02617, and NU02609, respectively) represent a statistically significant group compared to the group of 0 hrs. of ischemic time represented as gray dots (N = 18, 19, 14 endothelial cells from NU02608, NU02617, and NU02609, respectively). All samples were normalized to the average of values of the three 0 hrs. groups. P values by two-sided Kruskal Wallis test with post hoc Dunn’s multiple comparison test. c, Western blot showing p-ERK and other phosphoproteins in samples subjected to different ischemic times. Densitometry analysis for p-ERK western blot was performed using ERK1 and ERK2 staining. For this densitometry, every patient had density normalized by 0 minutes of ischemic time. N = 3 GBM samples. Error bars represent SEM. Western blot was done as a single experiment in 3 independent GBM samples.
Extended Data Fig. 5 Progression-free survival of the validation cohort from the Cloughesy T et al.14 clinical trial.
a, b, Kaplan-Meier showing progression-free survival following PD-1 blockade based on p-ERK high vs low for pre-study (a) and on-study (b) tumor samples. N = 13 GBM patients. P values by two-sided log rank test.
Extended Data Fig. 6 Multiplex immunofluorescence staining of recurrent GBM samples employing GFAP marker.
a, Bar plot showing the comparison of GFAP+ p-ERK+ cells and other cells expressing p-ERK+. N = 6 tumor samples. P value by two-sided Mann Whitney U test. Data is presented as mean ± s.d. b, Representative images of three different tumor samples derived form results in a. From top to bottom: a BRAFV600E GBM sample having high p-ERK staining, a wild-type BRAF/PTPN11 GBM having high p-ERK staining, and a wild-type BRAF/PTPN11 GBM displaying low p-ERK staining. For the three tumor samples: (left) H&E and p-ERK IHC images of the same tumor region. (middle), Multiplex immunofluorescence images showing the markers for GFAP, p-ERK, and DAPI. (right) Multiplex immunofluorescence images showing the markers for GFAP, CD163, and DAPI. Experiment was done using a tumor sample in one standardized run per patient.
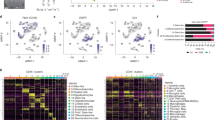
Extended Data Fig. 7 Single-cell RNA seq of GBM patients with high and low p-ERK IHC staining.
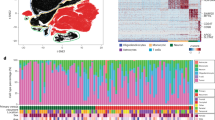
UMAP representation of 28,194 individual cells from 10 GBM patients measured with scRNA-seq (left). UMAP graph showing the representation of 3,153 myeloid cells derived from the 10 GBM patients (right). Each dot represents an individual cell.
Supplementary information
Supplementary Tables
Table 1. Clinical characteristics of our GBM cohort treated with anti-PD-1 therapy. Table 2. Clinical characteristics of our GBM cohort that did not receive immunotherapy. Table 3. Clinical characteristics of the validation cohort treated with adjuvant PD-1 blockade. Table 4. GO term enrichment in myeloid cells from high-p-ERK GBMs versus myeloid cells from low-p-ERK GBMs.
Supplementary Data 1
Flow cytometry gating for GBM patients treated with PD-1 blockade.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed immunoblots.
Source Data Extended Data Fig. 4
Unprocessed immunoblots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Arrieta, V.A., Chen, A.X., Kane, J.R. et al. ERK1/2 phosphorylation predicts survival following anti-PD-1 immunotherapy in recurrent glioblastoma. Nat Cancer 2, 1372–1386 (2021). https://doi.org/10.1038/s43018-021-00260-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00260-2
This article is cited by
-
Radio-pathomic maps of glioblastoma identify phenotypes of non-enhancing tumor infiltration associated with bevacizumab treatment response
Journal of Neuro-Oncology (2024)
-
Disclosing a metabolic signature of cisplatin resistance in MDA-MB-231 triple-negative breast cancer cells by NMR metabolomics
Cancer Cell International (2023)
-
Kunitz-type protease inhibitor TFPI2 remodels stemness and immunosuppressive tumor microenvironment in glioblastoma
Nature Immunology (2023)
-
IFN-γ Triggered IFITM2 Expression to Induce Malignant Phenotype in Elderly GBM
Journal of Molecular Neuroscience (2023)
-
Molecular Profiling and Targeted Therapies in Gliomas
Current Neurology and Neuroscience Reports (2023)