Abstract
Gut dysbiosis is observed in chronic hepatobiliary diseases and is frequently associated with liver carcinogenesis; however, the extent and specific mechanisms triggered by alterations in the microbiota mediating tumorigenesis in these patients remain unclear. Here we show that Enterococcus faecalis is abundant in the microbiota of patients with hepatitis C virus-related chronic liver disease. Xenotransplantation of gut microbiota from these patients increased the number of spontaneous liver tumors in mice and enhanced susceptibility to liver carcinogens. Hepatic colonization by gelE-positive E. faecalis increased liver expression of proliferative genes in a TLR4–Myd88-dependent manner, leading to liver tumorigenesis. Moreover, decreased fecal deoxycholic acid levels were associated with colonization by E. faecalis. Overall, these data identify E. faecalis as a key promoter of liver carcinogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All sequencing data and metadata have been archived in NCBI SRA as BioProject nos. PRJNA517801 and PRJNA517981. The KEGG database v.56 utilized in HUMAnN2 is available at HUMAnN2: http://huttenhower.sph.harvard.edu/humann2. Greengene database v.13_8_99, utilized in MacQIIME, is available from MacQIIME: http://www.wernerlab.org/software/macqiime. Source data for all numerical data are provided as Source data files. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
HUMAnN2 v.0.1.9 (http://huttenhower.sph.harvard.edu/humann2), MetaPhlAn2 v.2.0.0 (http://huttenhower.sph.harvard.edu/metaphlan2), MacQIIME v..9.1 (http://www.wernerlab.org/software/macqiime), PRINSEQ v.0.20.4 (http://prinseq.sourceforge.net/manual.html), bowtie2 v.2.2.4 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml), SAMtools v.1.2 (http://www.htslib.org/), DIAMOND v.0.7.5 (https://github.com/bbuchfink/diamond) and the LEfSe web version (https://huttenhower.sph.harvard.edu/lefse/) are available for analysis of metagenomic data.
References
Moeller, A. H. et al. Cospeciation of gut microbiota with hominids. Science 353, 380–382 (2016).
Lebreton, F. et al. Tracing the Enterococci from Paleozoic origins to the hospital. Cell 169, 849–861 (2017).
Garrett, W. S. Cancer and the microbiota. Science 348, 80–86 (2015).
Dapito, D. H. et al. Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 21, 504–516 (2012).
Global Burden of Disease Cancer Collaboration. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 32 cancer groups, 1990 to 2015: a systematic analysis for the Global Burden of Disease Study. JAMA Oncol. 3, 524–548 (2017).
Baumert, T. F., Juhling, F., Ono, A. & Hoshida, Y. Hepatitis C-related hepatocellular carcinoma in the era of new generation antivirals. BMC Med. 15, 52 (2017).
Ravi, S. et al. Unusually high rates of hepatocellular carcinoma after treatment with direct-acting antiviral therapy for hepatitis C related cirrhosis. Gastroenterology 152, 911–912 (2017).
Conti, F. et al. Early occurrence and recurrence of hepatocellular carcinoma in HCV-related cirrhosis treated with direct-acting antivirals. J. Hepatol. 65, 727–733 (2016).
Li, D. K. et al. The short-term incidence of hepatocellular carcinoma is not increased after hepatitis C treatment with direct-acting antivirals: an ERCHIVES study. Hepatology 67, 2244–2253 (2018).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014).
Boursier, J. et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology 63, 764–775 (2016).
Miele, L. et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology 49, 1877–1887 (2009).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Lamont, R. J., Koo, H. & Hajishengallis, G. The oral microbiota: dynamic communities and host interactions. Nat. Rev. Microbiol. 16, 745–759 (2018).
Turnbaugh, P. J. et al. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 1, 6ra14 (2009).
Arias, C. A. & Murray, B. E. The rise of the Enterococcus: beyond vancomycin resistance. Nat. Rev. Microbiol. 10, 266–278 (2012).
Cerda-Costa, N. & Gomis-Ruth, F. X. Architecture and function of metallopeptidase catalytic domains. Protein Sci. 23, 123–144 (2014).
Barnes, A. M. T. et al. Enterococcus faecalis readily colonizes the entire gastrointestinal tract and forms biofilms in a germ-free mouse model. Virulence 8, 282–296 (2017).
Mohamed, J. A., Huang, W., Nallapareddy, S. R., Teng, F. & Murray, B. E. Influence of origin of isolates, especially endocarditis isolates, and various genes on biofilm formation by Enterococcus faecalis. Infect. Immun. 72, 3658–3663 (2004).
Maharshak, N. et al. Enterococcus faecalis gelatinase mediates intestinal permeability via protease-activated receptor 2. Infect. Immun. 83, 2762–2770 (2015).
Steck, N. et al. Enterococcus faecalis metalloprotease compromises epithelial barrier and contributes to intestinal inflammation. Gastroenterology 141, 959–971 (2011).
Shogan, B. D. et al. Collagen degradation and MMP9 activation by Enterococcus faecalis contribute to intestinal anastomotic leak. Sci. Transl. Med. 7, 286ra268 (2015).
Enjoji, S., Ohama, T. & Sato, K. Regulation of epithelial cell tight junctions by protease-activated receptor 2. J. Vet. Med. Sci. 76, 1225–1229 (2014).
Franz, C. M., Huch, M., Abriouel, H., Holzapfel, W. & Galvez, A. Enterococci as probiotics and their implications in food safety. Int. J. Food Microbiol. 151, 125–140 (2011).
Seki, E. et al. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat. Med. 13, 1324–1332 (2007).
Yu, L. X. et al. Endotoxin accumulation prevents carcinogen-induced apoptosis and promotes liver tumorigenesis in rodents. Hepatology 52, 1322–1333 (2010).
Naugler, W. E. et al. Gender disparity in liver cancer due to sex differences in MyD88-dependent IL-6 production. Science 317, 121–124 (2007).
Singh, A. et al. NADPH oxidase 4 modulates hepatic responses to lipopolysaccharide mediated by Toll-like receptor-4. Sci. Rep. 7, 14346 (2017).
Kakiyama, G. et al. A simple and accurate HPLC method for fecal bile acid profile in healthy and cirrhotic subjects: validation by GC-MS and LC-MS. J. Lipid Res. 55, 978–990 (2014).
Kakiyama, G. et al. Modulation of the fecal bile acid profile by gut microbiota in cirrhosis. J. Hepatol. 58, 949–955 (2013).
Saito, H. E., Harp, J. R. & Fozo, E. M. Incorporation of exogenous fatty acids protects Enterococcus faecalis from membrane-damaging agents. Appl. Environ. Microbiol. 80, 6527–6538 (2014).
Seishima, J. et al. Gut-derived Enterococcus faecium from ulcerative colitis patients promotes colitis in a genetically susceptible mouse host. Genome Biol. 20, 252 (2019).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Iida, N. et al. Overuse of antianaerobic drug is associated with poor postchemotherapy prognosis of patients with hepatocellular carcinoma. Int. J. Cancer 145, 2701–2711 (2019).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Fujii, M. et al. A murine model for non-alcoholic steatohepatitis showing evidence of association between diabetes and hepatocellular carcinoma. Med. Mol. Morphol. 46, 141–152 (2013).
Iida, N. et al. Antitumor effect after radiofrequency ablation of murine hepatoma is augmented by an active variant of CC chemokine ligand 3/macrophage inflammatory protein-1alpha. Cancer Res. 70, 6556–6565 (2010).
Vankerckhoven, V. et al. Development of a multiplex PCR for the detection of asa1, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among European hospital isolates of Enterococcus faecium. J. Clin. Microbiol. 42, 4473–4479 (2004).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Nagamine, K., Hung, G. C., Li, B. & Lo, S. C. DNA sequence signatures for rapid detection of six target bacterial pathogens using PCR assays. Microbiol. Insights 8, 7–14 (2015).
Ryu, H. et al. Development of quantitative PCR assays targeting the 16S rRNA genes of Enterococcus spp. and their application to the identification of enterococcus species in environmental samples. Appl. Environ. Microbiol. 79, 196–204 (2013).
Jiang, C. et al. Intestinal farnesoid X receptor signaling promotes nonalcoholic fatty liver disease. J. Clin. Invest. 125, 386–402 (2015).
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 125, 279–284 (2001).
Acknowledgements
We thank M. Wakabayashi and M. Nagashimada for assistance with the animal experiments; A. Katayama, N. Nishiyama and K. Hara for assistance with histology experiments; and M. Hattori, K. Ohshima, K. Hosomochi, A. Tajima and S. Hashimoto for advice on sequencing analysis. M.S.G. and F.L. were supported by grants from the US National Institutes of Health (AI072360) and the Harvard-wide Program on Antibiotic Resistance (AI083214). This work was supported by JSPS KAKENHI grant JP25893081.
Author information
Authors and Affiliations
Contributions
N.I. and E.M. designed the entire study. N.I. performed experiments and data analysis and wrote the manuscript. M.Y. and Y.F. performed bacterial cultivation. J.S., H.O., M.H., H.O. and Z.W. performed mouse and bacterial experiments. Tatsuya Yamashita and K.A. performed acquisition and analysis of clinical data. Y.M., R.A. and Y.K. performed liquid chromatography–mass spectrometry experiments. F.L. and M.S.G. provided the gelE-deleted E. faecalis strain and interpreted the data. Taro Yamashita, Y.S. and M.S.G edited the manuscript. E.M. and S.K. supervised the study and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks Tim Greten, Florencia McAllister and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of gut microbiota based on subjects-related variates.
(a) Chao1 was calculated from metagenomic taxonomy data. Boxes represent 25th-75th percentile of the distribution. The median is shown in the middle of the boxes. Whiskers represent min to max. The number of samples were 21–24 and 10 iterations were performed in the analysis. (b–h) Unweighted UniFrac distances were calculated with taxonomic data for fecal samples based on subjects-related variates including age (b), use of PPI/H2RA (c), BCAA (d), obesity (e), daily alcohol consumption (f), smoking (g) and cirrhosis (h). Distances were visualized by principal coordinates analysis. Significant differences between groups were determined by PERMANOVA, and P values and q value are shown at the bottom of the plots. (a-b) LDA was performed with relative abundance of taxa in the feces. (i) HD subjects (n=24) and CLD/HCC subjects (n=23) without treatment of PPI, H2RA or BCAA were compared by LDA. (j) Ages were matched between HD subjects (n=13) and CLD/HCC subjects (n=36) at over 55 years old and compared by LDA. LDA scores with P value < 0.05 are shown in the graphs. q, adjusted P value by false discovery rate method; PPI, proton pump inhibitors; H2RA, H2 receptor antagonists; BCAA, branched-chain amino acids. (a), Kruskal Wallis test;. *, P < 0.05; **, P < 0.01. Exact P-values were as follows: (a) HD vs. CLD, P>0.999; HD vs. HCC, P=0.0045; CLD vs. HCC, P=0.0143.
Extended Data Fig. 2 ß-diversity analysis of the mouse feces after the fecal transplantation from the human subjects.
(a) Microbiota-depleted mice received the fecal microbiota from the subjects HD2, CLD1, CLD6, HCC8 or HCC14. Unweighted UniFrac distances were calculated with taxonomy data in feces of the mice at various time points after the fecal transplantation. The results are visualized by principal coordinate analysis. d, day; M, months. (b) Chao1 was calculated from taxonomy data. Boxes represent 25th-75th percentile of the distribution. The median is shown in the middle of the boxes. Whiskers represent min to max. The number of samples were 21–24 and 10 iterations were performed in the analysis. (c to f) Germ-free or antibiotics-pretreated mice received feces of thee subjects HD50, CLD9 and HCC11 (c), HD55, CLD13 and HCC12 (d), and HD59, CLD10 and HCC13 (e). Unweighted UniFrac distances were calculated with taxonomy data in feces of the mice and visualized. (f) UniFrac distances between the subjects and feces-transplanted germ-free mice, and between those and feces-transplanted antibiotics-pretreated mice were compared. ABX, antibiotics; FT, feces-transplanted. n=9. (b), Kruskal Wallis test; (f), two-sided Mann-Whitney U test. *, P < 0.05; ***, P < 0.001. Exact P-values were as follows: (b) HD vs. CLD, P<0.0001; HD vs. HCC, P<0.0001; CLD vs. HCC, P=0.0139: (f) P=0.0244.
Extended Data Fig. 3 The number of liver tumors and gene expression in the liver of in individual mouse that received fecal transplantation from the human subjects.
Fecal transplantation from the HD, CLD, or HCC subjects was performed in microbiota-depleted mice and followed by injection of DEN and CCl4 to induce liver carcinogenesis. (a) The mice were euthanized in 8 months after the DEN injection. The number of liver tumors per mouse is shown. (b) The mice were euthanized after injection of DEN once and CCl4 twice. mRNA expression of indicated genes was shown. These figures a and b correspond to individual plots of Fig. 2c, d, respectively.
Extended Data Fig. 4 Liver gene expression after transplantation of the subjects’ feces to germ-free mice.
Germ-free mice received fecal transplantation from the selected subjects, HD50, CLD9 and HCC11 (a and d), HD55, CLD13, HCC12 (b), HD59, CLD10, and HCC13 (c). (a–c) Then, after DEN injection once followed by CCl4 injection twice, the mice were euthanized and mRNA expression in the liver was analyzed by real-time PCR. n=4–5. (d) The mice received DEN injection twice followed by CCl4 injection and were euthanized in 8 months after the DEN injection. n=5. Means and SE are shown. GF, germ-free. (a–c), one-way ANOVA followed by Tukey’s test; (d), Kruskal-Wallis test. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P-values were as follows: (a) mki67 HD50 vs. CLD9, P=0.0073; HD50 vs. HCC11, P=0.0072; HD50 vs. GF, P=0.998; CLD9 vs. HCC11, P>0.999; CLD9 vs.GF, P=0.0054; HCC11 vs. GF, P=0.0053: ccnb1 HD50 vs. CLD9, P=0.0314; HD50 vs. HCC11, P=0.0209; HD50 vs. GF, P=0.999; CLD9 vs. HCC11, P=0.996; CLD9 vs.GF, P=0.0378; HCC11 vs. GF, P=0.0252: col1a1 HD50 vs. CLD9, P=0.433; HD50 vs. HCC11, P=0.181; HD50 vs. GF, P=0.303; CLD9 vs. HCC11, P=0.932; CLD9 vs.GF, P=0.0186; HCC11 vs. GF, P=0.0056: tnf HD50 vs. CLD9, P=0.151; HD50 vs. HCC11, P=0.0219; HD50 vs. GF, P=0.0432; CLD9 vs. HCC11, P=0.736; CLD9 vs.GF, P=0.0005; HCC11 vs. GF, P<0.0001: birc5 HD50 vs. CLD9, P=0.0053; HD50 vs. HCC11, P=0.010; HD50 vs. GF, P=0.998; CLD9 vs. HCC11, P=0.989; CLD9 vs.GF, P=0.0040; HCC11 vs. GF, P=0.0075: (b) mki67 HD55 vs. CLD13, P=0.0011; HD55 vs. HCC12, P=0.0003; CLD13 vs. HCC12, P=0.507: ccnb1 HD55 vs. CLD13, P=0.0031; HD55 vs. HCC12, P<0.0001; CLD13 vs. HCC12, P=0.045: col1a1 HD55 vs. CLD13, P=0.146; HD55 vs. HCC12, P=0.248; CLD13 vs. HCC12, P=0.929: tnf HD55 vs. CLD13, P=0.122; HD55 vs. HCC12, P=0.757; CLD13 vs. HCC12, P=0.339: birc5 HD55 vs. CLD13, P=0.0174; HD55 vs. HCC12, P<0.0001; CLD13 vs. HCC12, P=0.0047: (c) mki67 HD59 vs. CLD10, P=0.455; HD59 vs. HCC13, P=0.0205; CLD10 vs. HCC13, P=0.172: ccnb1 HD59 vs. CLD10, P=0.692; HD59 vs. HCC13, P=0.100; CLD10 vs. HCC13, P=0.354: col1a1 HD59 vs. CLD10, P=0.305; HD59 vs. HCC13, P=0.177; CLD10 vs. HCC13, P=0.929: tnf HD59 vs. CLD10, P=0.365; HD59 vs. HCC13, P=0.0403; CLD10 vs. HCC13, P=0.383: birc5 HD59 vs. CLD10, P=0.383; HD59 vs. HCC13, P=0.0467; CLD10 vs. HCC13, P=0.405: (d) HD50 vs. CLD9, P=0.303; HD50 vs. HCC11, P=0.019; CLD9 vs. HCC11, P=0.839.
Extended Data Fig. 5 Abundance of Enterococcus species in feces of the subjects.
Copy number of E. faecalis (a, d, e), E. faecium (b), and E. casseliflavus (c) in feces of the subjects was determined by quantitative PCR. PPI-untreated subjects or subjects over 55 years old were analyzed in (d, e), respectively. n=21–24 (a-c), n=13–23 (d–f) Number of liver tumors in E. faecalis-bearing mice is shown according to presence of fecal gelE. n=13–50. (g–j) Relative abundance (%) of Bacillus (g), Staphylococcus (h), Pseudomonas (i), and Serratia (j) in feces of the mice transplanted with human feces was determined by 16SrRNA-based sequencing. n=21–24. (a–j) Means and SE are shown. (a–e and g–j) Kruskal-Wallis test. (f) two-sided Mann-Whitney U test. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P-values were as follows: (a) HD vs. CLD, P=0.0007; HD vs. HCC, P=0.0002; CLD vs. HCC, P>0.999: (b) HD vs. CLD, P=0.206; HD vs. HCC, P=0.229; CLD vs. HCC, P>0.999: (c) HD vs. CLD, P>0.999; HD vs. HCC, P=0.238; CLD vs. HCC, P>0.999 (d) HD vs. CLD, P=0.0068; HD vs. HCC, P=0.0031; CLD vs. HCC, P>0.999: (e) HD vs. CLD, P=0.0014; HD vs. HCC, P=0.0019; CLD vs. HCC, P>0.999: (f) P=0.049: (g) HD vs. CLD, P>0.999; HD vs. HCC, P=0.0587; CLD vs. HCC, P=0.151: (h) HD vs. CLD, P>0.999; HD vs. HCC, P>0.999; CLD vs. HCC, P>0.999: (i) HD vs. CLD, P=0.100; HD vs. HCC, P=0.318; CLD vs. HCC, P>0.999: (j) HD vs. CLD, P>0.999; HD vs. HCC, P=0.657; CLD vs. HCC, P=0.704.
Extended Data Fig. 6 Deletion of gelE results in decreased colonization in the gut.
(a) Inoculation of E. faecalis V583 wild-type (WT) or ∆gelE strains was performed orally and rectally in germ-free or mice whose microbiota were depleted by antibiotics . CFU of V583 in feces was counted 8 days after the inoculation. Means and SE are shown. n=4. Four independent experiments were combined. One-way ANOVA followed by Tukey’s test. ***, P < 0.001. (b) The indicated E. faecalis strains were cultivated on gelatin-containing agar plates and gelatinase activity was evaluated. (c) After inoculation of E. faecalis V583 WT or ∆gelE strains in germ-free mice, the mice received DEN injection twice followed by CCl4 injection and were euthanized in 8 months after the DEN injection. Number of liver tumors is shown. Kruskal-Wallis test. n=5–7. (d) Homogenized liver harvested from V583 WT- or ∆gelE–inoculated microbiota-depleted mice was cultivated and CFU count was obtained. n=5. Representative of 2 independent repeats with similar results. Means and SE are shown. Mann-Whitney U test. Exact P-values were as follows: (a) GF + V583WT vs. GF + V583ΔgelE, P=0.998; GF + V583WT vs. microbiota-depleted + V583WT, P<0.0001; GF + V583WT vs. microbiota-depleted + V583ΔgelE, P<0.0001; GF + V583ΔgelE vs. microbiota-depleted + V583WT, P<0.0001; GF + V583ΔgelE vs. microbiota-depleted + V583ΔgelE, P<0.0001; microbiota-depleted + V583WT vs. microbiota-depleted + V583ΔgelE, P=0.0021.
Extended Data Fig. 7 Effects of E. faecalis colonization on liver tumor in Myd88-/- and Tlr4-/- mice.
Inoculation of E. faecalis V583 wild-type (WT) strains was performed orally and rectally in microbiota-depleted WT or Myd88-/- mice (a, b) and Tlr4-/-, Tlr4floxLyz2cre mice (c, d). (a) the mice received DEN injection twice followed by CCl4 injection and were euthanized in 8 months later. Number of liver tumors is shown. n=7. (b, c) CFU of E. faecalis strain V583 in feces of WT, Myd88-/- (a), or Tlr4-/- (b) after inoculation was determined. n=4–6. Four independent experiments were combined. (d) After DEN injection once followed by CCl4 injection twice, the mice were euthanized and mRNA expression in the liver was analyzed by real-time PCR. n=8–9. Representative of 2 independent repeats with similar results. (a–d) Means and SE are shown. (a), two-sided Mann-Whitney U test; (b–d), one-way ANOVA followed by Tukey’s test. *, P < 0.05; **, P < 0.01. ***, P < 0.001. Exact P-values were as follows: (a) P=0.0017: (b) WT mice + V583WT vs. WT mice + V583ΔgelE, P=0.0003; WT mice + V583WT vs. Myd88-/- mice + V583WT, P=0.962; WT mice + V583WT vs. Myd88-/- mice + V583ΔgelE, P=0.002; WT mice + V583ΔgelE vs. Myd88-/- mice + V583WT, P=0.0001; WT mice + V583ΔgelE vs. Myd88-/- mice + V583ΔgelE, P=0.845; Myd88-/- mice + V583WT vs. Myd88-/- mice + V583ΔgelE, P=0.0007: (d) mki67 Tlr4fl/flLyz2wt/wt vs. Tlr4fl/flLyz2cre/wt, P=0.417; Tlr4fl/flLyz2wt/wt vs. Tlr4-/-, P=0.0091; Tlr4fl/flLyz2cre/wt vs. Tlr4-/-, P=0.151: ccnb1 Tlr4fl/flLyz2wt/wt vs. Tlr4fl/flLyz2cre/wt, P=0.836; Tlr4fl/flLyz2wt/wt vs. Tlr4-/-, P=0.0029; Tlr4fl/flLyz2cre/wt vs. Tlr4-/-, P=0.0135: col1a1 Tlr4fl/flLyz2wt/wt vs. Tlr4fl/flLyz2cre/wt, P=0.888; Tlr4fl/flLyz2wt/wt vs. Tlr4-/-, P=0.389; Tlr4fl/flLyz2cre/wt vs. Tlr4-/-, P=0.676: birc5 Tlr4fl/flLyz2wt/wt vs. Tlr4fl/flLyz2cre/wt, P=0.531; Tlr4fl/flLyz2wt/wt vs. Tlr4-/-, P=0.0057; Tlr4fl/flLyz2cre/wt vs. Tlr4-/-, P=0.0726: tnf Tlr4fl/flLyz2wt/wt vs. Tlr4fl/flLyz2cre/wt, P=0.988; Tlr4fl/flLyz2wt/wt vs. Tlr4-/-, P=0.951; Tlr4fl/flLyz2cre/wt vs. Tlr4-/-, P=0.987.
Extended Data Fig. 8 Effect of E. faecalis is not based on initiation of carcinogenesis by DNA damage.
(a, b) After transplantation of feces of HD, CLD, or HCC, or inoculation of E. faecalis strain V583, DEN was injected. One day later liver was harvested and stained with anti-phospho-H2AX antibody. (a) Representative pictures of phospho-H2AX staining from 2 independent experiments are shown. Bars, 50µm. (b) Percentages of phospho-H2AX-positive nuclei were compared between HD, CLD and HCC, or V583 WT and gelE-deleted V583. n=24 or 12, respectively. (c) After inoculation of V583, CCl4 was injected with or without DEN, and 8 months later number of liver tumors was counted. Means and SE are shown. n=12. (b) Kruskal-Wallis test for 3 groups and two-sided Mann-Whitney U test for 2 groups. (c) two-sided Mann-Whitney U test. ***, P < 0.001. Exact P-values were as follows: (b) HD vs. CLD, P>0.999; HD vs. HCC, P>0.999; CLD vs. HCC, P>0.999: V583WT vs. V583ΔgelE, P=0.932: (c) P<0.0001.
Extended Data Fig. 9 Aberrant localization of tight junction protein ZO-1 in the colon of the mice colonized with gelE-positive E. faecalis.
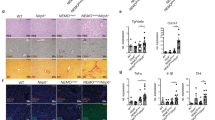
Inoculation of E. faecalis V583 wild-type (wt) or ∆gelE strains (a–e, and g) or fecal transplantation from HD, CLD and HCC (f) was performed orally and rectally in microbiota-depleted WT or F2rl1-/- mice. After DEN injection once followed by CCl4 injection twice, colon was harvested. (a–d) Colon tissues were stained with anti-ZO-1 antibody (green) and DAPI (blue). Bar, 100µm. Arrows indicate internalization of ZO-1 in the epithelial cells. Representative pictures from 2 independent experiments are shown. (e) Immunoblotting of colonic protein was performed with anti-ZO-1 antibody or anti-GAPDH antibody. Representative pictures from 2 independent experiments are shown. (f, g) mRNA expression of colon tissues were determined by PCR. n=16, 15, 16 and 6 for HD, CLD, HCC and ABX in (f), biologically independent samples. n=9 mice per group in (g) and two independent experiments were combined. (h) CFU of E. faecalis strain V583 in feces of WT and F2rl1-/- mice after inoculation was determined. n=4–6. Four independent experiments were combined. (f to h) Means and SE are shown. (f and h), one-way ANOVA followed by Tukey’s test; (g), two-sided Mann-Whitney U test. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P-values were as follows: (f) tnf HD vs. CLD, P=0.221; HD vs. HCC, P=0.0025; HD vs. ABX, P=0.997; CLD vs. HCC, P=0.309; CLD vs. ABX, P=0.365; HCC vs. ABX, P=0.023: il1b HD vs. CLD, P=0.213; HD vs. HCC, P=0.0099; HD vs. ABX, P=0.966; CLD vs. HCC, P=0.592; CLD vs. ABX, P=0.230; HCC vs. ABX, P=0.0283: il6 HD vs. CLD, P=0.472; HD vs. HCC, P=0.0201; HD vs. ABX, P=0.994; CLD vs. HCC, P=0.434; CLD vs. ABX, P=0.546; HCC vs. ABX, P=0.0749: (g) tnf V583WT vs. V583ΔgelE, P=0.0002: il1b V583WT vs. V583ΔgelE, P=0.0028: il6 V583WT vs. V583ΔgelE, P=0.297. ABX, antibiotics; GF, germ-free.
Extended Data Fig. 10 Fibrosis, gene expression and tumor number in the liver of E. faecalis-inoculated NASH mouse model.
Streptozotocin was subcutaneously injected on 2 days after birth and high-fat diet was started in 4 week-old of age. Fecal transplantation from the HD, CLD, or HCC subjects (a, b) or inoculation of E. faecalis strain V583 (b–d) was performed after depletion of microbiota in 6 week-old of age. The mice were euthanized in 17 week-old of age. (a) Representative images of the murine liver after Sirius Red staining. Bar, 100µm. (b) Area of fibrosis was measured as Sirius-red-positive area and shown. n=11–12 or 6. (c) The number of liver tumors per mouse is plotted. n=6. (d) mRNA expression in the liver was analyzed by real-time PCR. n=6. Representative of 2 independent repeats with similar results. (e) Plasma LPS level was shown. n=6. Means and SE are shown. (b), left, Kruskal-Wallis test. Right, two-sided Mann-Whitney U test; (c–e), two-sided Mann-Whitney U test. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P-values were as follows: (b) HD vs. CLD, P=0.0304; HD vs. HCC, P<0.0001; CLD vs. HCC, P=0.176: V583WT vs. V583ΔgelE, P=0.0260: (c) V583WT vs. V583ΔgelE, P=0.0286: (d) mki67, P=0.0022: ccnb1, P=0.0152: col1a1, P=0.132: birc5, P=0.0411: tnf, P=0.0260: ccl2, P=0.0411: (e) P=0.0022.
Supplementary information
Supplementary Information
Supplementary Tables 1–9 and Methods.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed immunoblots and/or gels.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Iida, N., Mizukoshi, E., Yamashita, T. et al. Chronic liver disease enables gut Enterococcus faecalis colonization to promote liver carcinogenesis. Nat Cancer 2, 1039–1054 (2021). https://doi.org/10.1038/s43018-021-00251-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00251-3