Abstract
Post-transcriptional modifications of RNA constitute an emerging regulatory layer of gene expression. The demethylase fat mass- and obesity-associated protein (FTO), an eraser of N6-methyladenosine (m6A), has been shown to play a role in cancer, but its contribution to tumor progression and the underlying mechanisms remain unclear. Here, we report widespread FTO downregulation in epithelial cancers associated with increased invasion, metastasis and worse clinical outcome. Both in vitro and in vivo, FTO silencing promotes cancer growth, cell motility and invasion. In human-derived tumor xenografts (PDXs), FTO pharmacological inhibition favors tumorigenesis. Mechanistically, we demonstrate that FTO depletion elicits an epithelial-to-mesenchymal transition (EMT) program through increased m6A and altered 3′-end processing of key mRNAs along the Wnt signaling cascade. Accordingly, FTO knockdown acts via EMT to sensitize mouse xenografts to Wnt inhibition. We thus identify FTO as a key regulator, across epithelial cancers, of Wnt-triggered EMT and tumor progression and reveal a therapeutically exploitable vulnerability of FTO-low tumors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the GEO repository under accession number GSE128582. Human cancer data (bladder urothelial carcinoma, breast invasive carcinoma, cervical squamous cell carcinoma and endocervical adenocarcinoma, glioblastoma multiforme, kidney chromophobe, lung adenocarcinoma and lung squamous cell carcinoma, prostate adenocarcinoma, thyroid carcinoma and uterine corpus endometrial carcinoma) were derived from the TCGA Research Network (http://cancergenome.nih.gov/). RNA-seq data from TCGA were obtained as normalized FPKM counts from GDC (https://portal.gdc.cancer.gov/repository), and PAM50 subtype information was extracted from UCSC Xena (https://xenabrowser.net). METABRIC expression and clinical data61 were downloaded through the European Genome-Phenome Archive (EGA) (http://www.ebi.ac.uk/ega/, accession number EGAS00000000083). Previously published microarray data that were reanalyzed here are available under accession codes GSE9195, GSE10780, GSE10810, GSE12276, GSE19615, GSE20711 and GSE21653. Reannotation and processing of Affymetrix Human Genome U133 Plus 2.0 array data were performed as previously detailed76.
The hg19 reference transcriptome (version 85) was obtained from the Ensembl portal (ensembl.org). Hallmark gene sets were obtained from the MsigDB database (https://www.gsea-msigdb.org/gsea/msigdb). Human poly(A) sites were obtained from PolyA_DB2 (http://polya.umdnj.edu/PolyA_DB2).
For motility assays related to Figs. 2a,b, 3c,e and 7b and Extended Data Figs. 2a,b,d,e, 3c,f and 4d, independent repeats have been deposited on the Figshare repository (https://doi.org/10.6084/m9.figshare.14602932, https://doi.org/10.6084/m9.figshare.14602944, https://doi.org/10.6084/m9.figshare.14602953, https://doi.org/10.6084/m9.figshare.14602965, https://doi.org/10.6084/m9.figshare.14602971 and https://doi.org/10.6084/m9.figshare.14602986). Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
Code supporting this study is available at a dedicated GitHub repository (https://github.com/martinBizet/m6A_FTO_cancer).
References
Harcourt, E. M., Kietrys, A. M. & Kool, E. T. Chemical and structural effects of base modifications in messenger RNA. Nature 541, 339–346 (2017).
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Shi, H., Wei, J. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Deng, X., Su, R., Stanford, S. & Chen, J. Critical enzymatic functions of FTO in obesity and cancer. Front. Endocrinol. 9, 396 (2018).
Geula, S. et al. Stem cells. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Meyer, K. D. et al. 5′ UTR m6A promotes cap-independent translation. Cell 163, 999–1010 (2015).
Zhou, J. et al. Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Liu, N. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518, 560–564 (2015).
Yue, Y., Liu, J. & He, C. RNA N6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev. 29, 1343–1355 (2015).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Li, Z. et al. FTO plays an oncogenic role in acute myeloid leukemia as a N6-methyladenosine RNA demethylase. Cancer Cell 31, 127–141 (2017).
Su, R. et al. R-2HG exhibits anti-tumor activity by targeting FTO/m6A/MYC/CEBPA signaling. Cell 172, 90–105 (2018).
Vu, L. P. et al. The N6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 23, 1369–1376 (2017).
Huang, Y. et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell 35, 677–691 (2019).
Su, R. et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38, 79–96 (2020).
Cui, Q. et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 18, 2622–2634 (2017).
Yang, S. et al. m6A mRNA demethylase FTO regulates melanoma tumorigenicity and response to anti-PD-1 blockade. Nat. Commun. 10, 2782 (2019).
Zhuang, C. et al. N6-methyladenosine demethylase FTO suppresses clear cell renal cell carcinoma through a novel FTO–PGC-1α signalling axis. J. Cell. Mol. Med. 23, 2163–2173 (2019).
Rong, Z. X. et al. Downregulation of fat mass and obesity associated (FTO) promotes the progression of intrahepatic cholangiocarcinoma. Front. Oncol. 9, 369 (2019).
Wen, L., Pan, X., Yu, Y. & Yang, B. Down-regulation of FTO promotes proliferation and migration, and protects bladder cancer cells from cisplatin-induced cytotoxicity. BMC Urol. 20, 39 (2020).
Niu, Y. et al. RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Mol. Cancer 18, 46 (2019).
Wu, L., Wu, D., Ning, J., Liu, W. & Zhang, D. Changes of N6-methyladenosine modulators promote breast cancer progression. BMC Cancer 19, 326 (2019).
Xu, Y. et al. The FTO/miR-181b-3p/ARL5B signaling pathway regulates cell migration and invasion in breast cancer. Cancer Commun. 40, 484–500 (2020).
Peitzsch, C., Tyutyunnykova, A., Pantel, K. & Dubrovska, A. Cancer stem cells: the root of tumor recurrence and metastases. Semin. Cancer Biol. 44, 10–24 (2017).
Moro, M. et al. Establishment of patient derived xenografts as functional testing of lung cancer aggressiveness. Sci. Rep. 7, 6689 (2017).
Huang, Y. et al. Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res. 43, 373–384 (2015).
Gil-Rendo, A. et al. Association between [18F]fluorodeoxyglucose uptake and prognostic parameters in breast cancer. Br. J. Surg. 96, 166–170 (2009).
Groheux, D., Espié, M., Giacchetti, S. & Hindié, E. Performance of FDG PET/CT in the clinical management of breast cancer. Radiology 266, 388–405 (2012).
Angeloni, V., Tiberio, P., Appierto, V. & Daidone, M. G. Implications of stemness-related signaling pathways in breast cancer response to therapy. Semin. Cancer Biol. 31, 43–51 (2015).
Velloso, F. J. et al. The crossroads of breast cancer progression: insights into the modulation of major signaling pathways. Onco. Targets Ther. 10, 5491–5524 (2017).
Zhou, S. et al. FTO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting β-catenin through mRNA demethylation. Mol. Carcinog. 57, 590–597 (2018).
Hu, B. et al. Epigenetic activation of WNT5A drives glioblastoma stem cell differentiation and invasive growth. Cell 167, 1281–1295 (2016).
Mikels, A. J. & Nusse, R. Purified Wnt5a protein activates or inhibits β-catenin–TCF signaling depending on receptor context. PLoS Biol. 4, e115 (2006).
Hung, T. H. et al. Wnt5A regulates ABCB1 expression in multidrug-resistant cancer cells through activation of the non-canonical PKA/β-catenin pathway. Oncotarget 5, 12273–12290 (2014).
Veeman, M. T., Axelrod, J. D. & Moon, R. T. A second canon: functions and mechanisms of β-catenin-independent Wnt signaling. Dev. Cell 5, 367–377 (2003).
Desmedt, C. et al. Genomic characterization of primary invasive lobular breast cancer. J. Clin. Oncol. 34, 1872–1881 (2016).
Bartosovic, M. et al. N6-methyladenosine demethylase FTO targets pre-mRNAs and regulates alternative splicing and 3′-end processing. Nucleic Acids Res. 45, 11356–11370 (2017).
Ji, Z., Lee, J. Y., Pan, Z., Jiang, B. & Tian, B. Progressive lengthening of 3′ untranslated regions of mRNAs by alternative polyadenylation during mouse embryonic development. Proc. Natl Acad. Sci. USA 106, 7028–7033 (2009).
Gonsalves, F. C. et al. An RNAi-based chemical genetic screen identifies three small-molecule inhibitors of the Wnt/wingless signaling pathway. Proc. Natl Acad. Sci. USA 108, 5954–5963 (2011).
Tan, A., Dang, Y., Chen, G. & Mo, Z. Overexpression of the fat mass and obesity associated gene (FTO) in breast cancer and its clinical implications. Int. J. Clin. Exp. Pathol. 8, 13405–13410 (2015).
Pastushenko, I. et al. Identification of the tumour transition states occurring during EMT. Nature 556, 463–468 (2018).
Zhang, Y. & Weinberg, R. A. Epithelial-to-mesenchymal transition in cancer: complexity and opportunities. Front. Med. 12, 361–373 (2018).
Lin, X. et al. RNA m6A methylation regulates the epithelial mesenchymal transition of cancer cells and translation of Snail. Nat. Commun. 10, 2065 (2019).
Rao, V. K. et al. Phosphorylation of Tet3 by cdk5 is critical for robust activation of BRN2 during neuronal differentiation. Nucleic Acids Res. 48, 1225–1238 (2020).
Shi, F. T. et al. Ten-eleven translocation 1 (Tet1) is regulated by O-linked N-acetylglucosamine transferase (Ogt) for target gene repression in mouse embryonic stem cells. J. Biol. Chem. 288, 20776–20784 (2013).
Blaschke, K. et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 500, 222–226 (2013).
Losman, J. A. & Kaelin, W. G. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 27, 836–852 (2013).
Beerling, E. et al. Plasticity between epithelial and mesenchymal states unlinks EMT from metastasis-enhancing stem cell capacity. Cell Rep. 14, 2281–2288 (2016).
Liu, J. et al. Landscape and regulation of m6A and m6Am methylome across human and mouse tissues. Mol. Cell 77, 426–440 (2020).
Clevers, H. & Nusse, R. Wnt/β-catenin signaling and disease. Cell 149, 1192–1205 (2012).
Pai, S. G. et al. Wnt/β-catenin pathway: modulating anticancer immune response. J. Hematol. Oncol. 10, 101 (2017).
Mukherjee, N. & Panda, C. K. Wnt/β-catenin signaling pathway as chemotherapeutic target in breast cancer: an update on pros and cons. Clin. Breast Cancer 20, 361–370 (2020).
Tian, T. V. et al. Identification of novel TMPRSS2:ERG mechanisms in prostate cancer metastasis: involvement of MMP9 and PLXNA2. Oncogene 33, 2204–2214 (2014).
Yuan, B. F. Liquid chromatography–mass spectrometry for analysis of RNA adenosine methylation. Methods Mol. Biol. 1562, 33–42 (2017).
Ross, R., Cao, X., Yu, N. & Limbach, P. A. Sequence mapping of transfer RNA chemical modifications by liquid chromatography tandem mass spectrometry. Methods 107, 73–78 (2016).
Lánczky, A. et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res. Treat. 160, 439–446 (2016).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Varambally, S. et al. Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 8, 393–406 (2005).
Rokavec, M., Kaller, M., Horst, D. & Hermeking, H. Pan-cancer EMT-signature identifies RBM47 down-regulation during colorectal cancer progression. Sci. Rep. 7, 4687 (2017).
Chen, S. et al. AfterQC: automatic filtering, trimming, error removing and quality control for fastq data. BMC Bioinformatics 18, 80 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lee, J. Y., Yeh, I., Park, J. Y. & Tian, B. PolyA_DB 2: mRNA polyadenylation sites in vertebrate genes. Nucleic Acids Res. 35, D165–D168 (2007).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Veeman, M. T., Slusarski, D. C., Kaykas, A., Louie, S. H. & Moon, R. T. Zebrafish prickle, a modulator of noncanonical Wnt/Fz signaling, regulates gastrulation movements. Curr. Biol. 13, 680–685 (2003).
Metzger-Filho, O. et al. Genomic grade adds prognostic value in invasive lobular carcinoma. Ann. Oncol. 24, 377–384 (2013).
Van Grembergen, O. et al. Portraying breast cancers with long noncoding RNAs. Sci. Adv. 2, e1600220 (2016).
Acknowledgements
J.J. was supported by the Belgian ‘Fonds de la Recherche Scientifique’ (FNRS) postdoctoral fellowship, E. Collignon was supported by the L’Oréal ‘For Women In Science’ fellowship and by the Belgian FNRS and Y.J. was supported by the China Scholarship Council (CSC) postdoctoral fellowship. C.A.W., M.B., B.H. and G.D. were supported by the Belgian FNRS. N.K.S. was supported by the ULB Foundation. F.F. is a ULB Professor. F.F.’s lab was funded by grants from the FNRS and Télévie, the ‘Action de Recherche Concertée’ (ARC; AUWB-2018-2023 ULB-No 7), Wallon Region grants (U-CAN-REST, INTREPID), an FNRS Welbio grant, the ULB Foundation and the Belgian Foundation Against Cancer (FCC 2016-086 FAF-F/2016/872). P.C. and A.C. are Senior Research Associate and Research Director at the FNRS, respectively. C.S. is a research director at the FNRS and his lab is supported by grants from the FNRS. P.A.L. was supported by the National Institutes of Health (GM 058843). Trace is supported by the ‘Belgian Foundation Against Cancer’ and, as a part of the EurOPDX infrastructure, by EDIReX a Horizon 2020 grant agreement 731105. PET–CT acquisition was performed at the nuMix-CMMI, which is supported by the European Regional Development Fund (ERDF) and the Walloon Region.
Author information
Authors and Affiliations
Contributions
J.J., E. Collignon and F.F. designed experiments and interpreted data. J.J., E. Collignon, C.A.W. and Y.J. performed in vitro phenotypic assays. E. Calonne performed m6A-seq and paired-end RNA-seq experiments. C.A.W., J.J., E. Collignon, B.H., P.P., K.S., P.C. and A.C. performed RNAi and overexpression experiments as well as dot blots, western blots and RT–qPCR. S.G., C.D., Y.B., F.R., J.L.C., M.D.C., X.L., L.T., K.W.G. and C.S. provided human biopsies/data and helped with analysis of these samples. P.A.L., R.L.R. and B.F.Y. performed mass spectrometry experiments. M.K., R.M., R.D., M.G. and G.G. performed mouse xenograft experiments. Z.W. analyzed and interpreted the FDG PET experiments. L.R., E.L. and J.C.M. designed and performed PDX experiments. M.B., G.D., N.K.S., A.K. and G.M. performed bioinformatics analyses. F.F. directed the study and J.J., E. Collignon and F.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
F.F. is a cofounder of Epics Therapeutics, Belgium. The other authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 FTO downregulation promotes tumorigenesis in breast cancer.
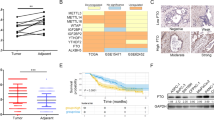
a, FTO expression in the Affymetrix data meta-analysis cohort (left, 171 normal vs. 812 tumor breast tissues, p = 5e-50). RT-qPCR quantification of FTO in an in-house cohort (right, 9 normal vs. 47 tumor breast tissues, p = 0.005), normalized to ACTB and SDHA. The box defines the IQR split by the median, with whiskers extending to the most extreme values within 1.5×IQR beyond the box. P values calculated by two-tailed t-test. b, Quantification of m6A levels by mass spectrometry in paired human normal and tumor breast samples (n = 6 per group, pairs connected with line, red dots represent the group medians, two-tailed paired t-test, p = 0.046). c, Validation of FTO depletion by two shRNAs in SKBR3 cells by RT-qPCR (upper panel, mean + SD, pRNAi#1 = 6e-10; pRNAi#2 = 0.0045, nRNAi#1 = 7; nRNAi#2 = 3), normalized to ACTB and SDHA) and western blotting (lower panel, representative of n = 3 independent replicates with histone H3 as loading control). P values calculated by two-tailed paired t-test. d, Light microscopy imaging of mammospheres formed by RNAi Ctrl and RNAi FTO SKBR3 cells with scale bar of 100 µm (representative of n = 3 independent experiments). e, Effects of FTO knockdown on colony-forming capacity (left, P = 0.001) and mammosphere-forming capacity (right, P = 0.02) in SKBR3 cells with a second shRNA (RNAi FTO #2). Data from n = 3 technical replicates (mean + SD) within a single experiment, representative of 3 biologically independent experiments provided as Source Data. P values calculated by two-tailed t-test. f, Representative western blot of doxycycline-induced overexpression of wild-type FTO (FTO-WT) and of a catalytically inactive FTO mutant (FTO-MUT) in SKBR3 cells (upper panel) with β-actin as loading control (n = 3). Effect of FTO-WT and FTO-MUT overexpression on colony formation and mammosphere formation in SKBR3 cells (bottom). Data from n = 3 technical replicates (mean + SD) within a single experiment, representative of 3 and 2 biologically independent experiments, respectively, provided as Source Data. P values calculated by two-tailed t-test, as indicated. g, FTO depletion by shRNA in breast cancer MCF7 cells (n = 6) by RT-qPCR. Data as mean + SD, normalized to ACTB and SDHA. P values calculated by two-tailed paired t-test (p = 4e-6).
Extended Data Fig. 2 Depletion of FTO promotes breast cancer cell motility and metastases.
a-b, Real-time chemotactic invasion (a) and migration (b) of RNAi FTO (red, shRNA #1 and #2, respectively) and RNAi control SKBR3 cells (green). c, Wound scratch assay performed with RNAi Ctrl and RNAi FTO SKBR3 cells. Representative pictures (left) with quantification of wound closure (as percentage of the width at 0 h, right). Data from a single experiment, representative of n = 2 biologically independent experiments provided as Source Data. d, Cell viability by MTT assay (left, mean + SD, n = 5, two-tailed paired t-test, p = 0.37) and real-time proliferation (right) of FTO-depleted and control SKBR3 cells. e, Western blotting of FTO depletion and rescue by overexpression of wild-type FTO (FTO-WT) or a catalytically inactive FTO mutant (FTO-MUT) in SKBR3 cells (upper panel, n = 3 with β-actin as loading control) and real-time chemotactic migration (lower panel). f, Expression of FTO in PDX shown by immunochemistry. Representative image of tissue sections with scale bar of 250 µm (n = 3). g, Metabolism imaging with 18F-FDG PET of subiliac lymph nodes xenograft mice bearing RNAi Ctrl (black, n = 4) or RNAi FTO (red, n = 5) SKBR3 tumors. Representative images (9 weeks after inoculation) are shown (pathological uptake in LN encircled; physiological uptake in bladder (Bl)). The box defines the IQR split by the median, with whiskers extending to the most extreme values within 1.5×IQR beyond the box. P-value calculated by two-tailed t-test (P = 0.0061). h, Kaplan-Meier based on FTO expression and disease-specific survival in the breast cancer METABRIC cohort (n(high FTO) = 660, n(low FTO) = 661, logrank test, P = 7e-4). i, Forest plot showing association between FTO expression and distant-metastasis-free survival in breast cancer. Data shown as hazard ratio (log2), with 95% confidence interval, for both the aggregate KM Plot data set (‘overall’, presented in Fig. 2f) and individual cohorts. The number of samples (n) and P-values (calculated by two-sided logrank tests, in bold if P < 0.05) are indicated for each cohort. Data for (a), (b), (d), (e) are from n = 3 technical replicates (mean + SD) within a single experiment, representative of 3 biologically independent experiments (except for (e): 2 replicates). Independent repeats have been deposited on the Figshare repository (DOI: 10.6084/m9.figshare.14602965).
Extended Data Fig. 3 FTO depletion in prostate, cervical and lung cancers.
a, FTO depletion by shRNA in PC3 prostate cancer cells by RT-qPCR (left). Mean + SD from 3 independent experiments, normalized to ACTB and SDHA. P value calculated by two-tailed paired t-test (P = 0.004). Representative western blot (right) of FTO depletion in PC3 cells (n = 3) with β-actin as loading control. b, Nude mice were injected subcutaneously with PC3 prostate cells and tumors was monitored every two days (n = 6 tumors per group; mean ± SEM, p = 0.0002, two-way ANOVA). c, Real-time chemotactic invasion of RNAi FTO (red) and RNAi control PC3 cells (green). d, FTO depletion by shRNA in Ca Ski cervical cancer cells (left, n = 3, P = 0.009) and H1650 lung cancer cells (right, n = 3, p = 0.0004) by RT-qPCR. Data as mean + SD, normalized to ACTB and SDHA. P values calculated by two-tailed paired t-test. e, Effects of FTO knockdown on tumorsphere-forming capacity in Ca Ski (left, P = 0.02) and H1650 cells (right, p = 0.008). Data from n = 3 technical replicates (mean + SD) within a single experiment, representative of 2 biologically independent experiments provided as Source Data. P values calculated by two-tailed t-test. f, Real-time chemotactic migration of FTO-depleted (red) and control (green) H1650 cells. g-h, Kaplan-Meier curves showing the overall survival of patients with either high-FTO or low-FTO lung tumors (n(high FTO) = 333, n(low FTO) = 326, logrank test, P = 0.0073) (g) or uterine tumors (n(high FTO) = 310, n(low FTO) = 232, logrank test, P = 0.048) (h). Motility data in (c), (f) from n = 3 technical replicates (mean + SD) within a single experiment, representative of 2 and 3 biologically independent experiments, respectively. Independent repeats have been deposited on the Figshare repository (DOI: 10.6084/m9.figshare.14602971).
Extended Data Fig. 4 FTO downregulation promotes EMT in several cancers.
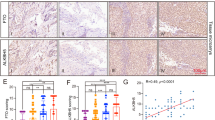
a, Representative light microscopy imaging of RNAi Ctrl or RNAi FTO SKBR3 cells (n = 3 independent experiments). Cells displaying elongated morphology and extended pseudopodia are indicated with arrows. Scale bars represent 50μm. b, Representative western blot of mesenchymal EMT markers in FTO-depleted (RNAi FTO) and control (RNAi Ctrl) SKBR3 cells (n = 3) with HDAC1 as loading control. c, Immunochemistry staining of EMT markers in SKBR3 xenografted tumors (from Fig. 1e at endpoint). Percent VIM and FN1 positive cells (n = 3 fields × 3 tumors per group; mean + SEM). P-values calculated by two-tailed t-test (pVIM = 0.0002; pFN1 < 0.0001). d, Real-time chemotactic migration of FTO-depleted and control SKBR3 cells in the absence and presence of TGFβ. Data from n = 3 technical replicates (mean + SD) within a single experiment, representative of 3 biologically independent experiments. Independent repeats have been deposited on the Figshare repository (DOI: 10.6084/m9.figshare.14602986). e, Relative gene expression of mesenchymal (VIM, MMP2) and epithelial (FSTL3) EMT marker genes, as estimated by RT-qPCR in SKBR3 cells following RNAi of FTO, treated or not with TGFβ. Mean + SD from n = 3 independent experiments, normalized to ACTB and SDHA. P values calculated by two-tailed paired t-test as indicated. f, Representative western blot of mesenchymal EMT markers FN1 and SLUG in FTO-depleted and control PC3 cells (n = 3) with β-actin and histone H3 as loading controls. g, Relative gene expression of mesenchymal EMT marker genes, as estimated by RT-qPCR in MCF7 (left, n = 4 except for nFN1 = 3) and H1650 (right, n = 3 except for nSLUG = 4) cells following RNAi of FTO. Mean + SD, normalized to ACTB and SDHA. P values calculated by two-tailed t-test. Representative western blot of mesenchymal EMT markers FN1, CDH1 and SLUG in FTO-depleted and control MCF7 (left) and H1650 (right) cells (n = 3) with HSP90 and HDAC1 as loading controls. h, Immunochemistry staining of VIM in PC3 xenografted tumors (from Fig. 3c at endpoint). Percent VIM positive cells (n = 3 fields × 3 tumors per group; mean + SEM). P-value calculated by two-tailed t-test (p = 0.0008). Scale bars represent 250μm.
Extended Data Fig. 5 FTO loss upregulates Wnt/β-catenin signaling in breast and prostate cancers.
a, Numbers of m6A peaks and related transcripts identified by m6A-seq in RNAi Ctrl and RNAi FTO SKBR3 cells (upper panel, two independent biological replicates). Scatterplot displaying differentially m6A-methylated peaks (lower panel, hyper and hypo = peaks with increased and decreased m6A in FTO-depleted cells, respectively). b-d, Visualization of the IP m6A signal in RNAi FTO and RNAi Ctrl SKBR3 cells at the DVL3, FZD1, SOX11, MARK2, CSNK1G2 (b), CTNNB1 (c), and WNT5A (d) transcripts. e, Representative western blot of β-catenin in RNAi FTO #2 and RNAi Ctrl #2 SKBR3 cells with β-actin as loading control (n = 3). f, Representative western blot of cytoplasmic and nuclear β-catenin in RNAi FTO and RNAi Ctrl SKBR3 cells (n = 3). Loading controls: β-Tubulin (cytoplasmic) and histone H3 (nuclear). Wnt3a treatment as positive control for nuclear translocation of β-catenin. g, Representative immunofluorescence staining of β-catenin (red) and DAPI (blue) in RNAi Ctrl and RNAi FTO SKBR3 (n = 3). Scale bars represent 20μm. h, Relative gene expression of Wnt/β-catenin targets measured by RT-qPCR in SKBR3 cells. Mean + SD from n = 6 independent replicates (except for n(TCF1) = 5), normalized to ACTB and SDHA. Two-tailed paired t-tests: PTCF1 < 0.0001; PAXIN2 = 0.0001; PNMYC=0.001. i, Western blot of total β-catenin in RNAi FTO vs. RNAi Ctrl from SKBR3 xenografted tumors in mice (n = 4 per group) with β-actin as loading control. j, Numbers of m6A peaks and related transcripts identified in three human breast cancer biopsies by m6A-seq (upper left panel). The m6A motif (upper right) retrieved from sample BC1 includes the consensus motif DRACH. Bar graph displaying the distribution of m6A peaks relatively to transcriptomic regions (lower panel, representative of sample BC1). k, m6A IP signal at transcripts related to the Wnt/β-catenin signaling (CSNK1G2, MARK2, AXIN1 and FZD1), in three human breast cancer biopsies.
Extended Data Fig. 6 Regulation of FTO targets.
a, Numbers of m6A peaks and related transcripts identified by m6A-seq in RNAi Ctrl and RNAi FTO PC3 cells (upper panel, two independent biological replicates). Schematic representation of m6A/FTO mRNA targets (red asterisks) identified within the Wnt/β-catenin signaling pathway (lower panel). b, Visualization of the IP m6A signal in RNAi FTO and RNAi Ctrl PC3 cells at the WNT3A and GSK3B transcripts. c, Representative immunochemistry staining of β-catenin in PC3 xenografted tumors with scale bar at 250 µm. Percent β-catenin positive cells (right, n = 3 fields × 3 tumors per group; mean + SD). Two-tailed t-test, P = 0.005. d, Relative quantification of the CSNK1D levels in RNAi FTO and RNAi Ctrl SKBR3 cells displayed in the western blot shown in Fig. 6a. e, Western blot showing the level of CSNK1D in an independent biological replicate of RNAi FTO and RNAi Ctrl SKBR3 cells (left, with HDAC1 as loading control, related to Fig. 6a). Relative quantification of CSNK1D levels (right). Western blot data in (d-e) are representative of n = 3 biologically independent experiments and normalized to HDAC1.
Extended Data Fig. 7 FTO-low tumors are sensitive to WNT inhibitor therapy.
a, Relative gene expression of Wnt/β-catenin target genes AXIN2 and TCF1, as estimated by RT-qPCR, in SKBR3 cells following RNAi of FTO, treated or not with iCRT3. Mean + SD from 3 independent experiments, normalized to ACTB and SDHA. P values calculated by two-tailed t-test, as indicated. b, Wnt/β-catenin transcriptional activity, as measured by TOPFlash β-catenin/TCF-LEF reporter assay performed on SKBR3 cells following RNAi of FTO, in the absence and presence of iCRT3 (mean + SD, n = 5, iCRT3 doses as indicated). P values calculated by two-tailed t-test, as indicated. c, The migration capacity, as shown in Fig. 7b, was quantified by the slope of the migration curves and corrected for cell viability (as measured by MTT assay in Fig. 7c, from 2 biologically independent experiments). P-values calculated by two-tailed t-test (p(RNAi Ctrl)=0.58, p(RNAi FTO) = 0.018). d, Levels of the mesenchymal EMT marker proteins FN1, VIM and SNAIL in RNAi Ctrl and RNAi FTO SKBR3 cells in the absence and presence of iCRT3 (n = 3) with histone H3 and β-actin as loading controls. e, Levels of FN1 protein in FTO-depleted and control tumors derived from SKBR3 xenografts treated with iCRT3 or vehicle (DMSO), with β-actin as loading control (n = 3 independent experiments). f, Nude mice growing tumors from RNAi Ctrl (left) or RNAi FTO (right) SKBR3 cells were treated with LGK974 (n = 5 tumors per group, mean ± SEM). P values calculated by two-way ANOVA (p(RNAi Ctrl)=0,69; p(RNAi FTO) = 0.48). g, Nude mice growing tumors from RNAi Ctrl (left) or RNAi FTO (right) PC3 cells were treated with or without Trastuzumab (n = 5 tumors per group, mean ± SEM). P values calculated by two-way ANOVA, (p(RNAi Ctrl)<0.0001; p(RNAi FTO) < 0.0001).
Supplementary information
Source data
Source Data Fig. 1
Numerical source data.
Source Data Fig. 2
Numerical source data.
Source Data Fig. 3
Numerical source data.
Source Data Fig. 4
Numerical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Numerical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Numerical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Numerical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 8
Numerical source data.
Source Data Extended Data Fig. 1
Numerical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Numerical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Numerical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Numerical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4
Numerical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Numerical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Numerical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Jeschke, J., Collignon, E., Al Wardi, C. et al. Downregulation of the FTO m6A RNA demethylase promotes EMT-mediated progression of epithelial tumors and sensitivity to Wnt inhibitors. Nat Cancer 2, 611–628 (2021). https://doi.org/10.1038/s43018-021-00223-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00223-7
This article is cited by
-
The role of RNA-modifying proteins in renal cell carcinoma
Cell Death & Disease (2024)
-
Transcriptome-wide profiling identifies colon cancer-associated m6A transcripts and potential RNA methyl modifiers
Molecular Biology Reports (2024)
-
Methylation across the central dogma in health and diseases: new therapeutic strategies
Signal Transduction and Targeted Therapy (2023)
-
m6A RNA methylation orchestrates transcriptional dormancy during paused pluripotency
Nature Cell Biology (2023)
-
Regulation of m6A modification on ferroptosis and its potential significance in radiosensitization
Cell Death Discovery (2023)