Abstract
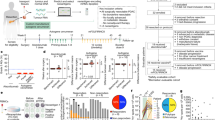
2′3′-Cyclic GMP-AMP (cGAMP) is an intracellular second messenger that is synthesized in response to cytosolic double-stranded DNA and activates the innate immune STING pathway. Our previous discovery of its extracellular hydrolase ENPP1 hinted at the existence of extracellular cGAMP. Here we detected that cGAMP is continuously exported but then efficiently cleared by ENPP1, explaining why it has previously escaped detection. By developing potent, specific and cell-impermeable ENPP1 inhibitors, we found that cancer cells continuously export cGAMP in culture at steady state and at higher levels when treated with ionizing radiation (IR). In mouse tumors, depletion of extracellular cGAMP decreased tumor-associated immune cell infiltration and abolished the curative effect of IR. Boosting extracellular cGAMP with ENPP1 inhibitors synergized with IR to delay tumor growth. In conclusion, extracellular cGAMP is an anticancer immunotransmitter that could be harnessed to treat cancers with low immunogenicity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The ENPP1 mRNA expression data were derived from the TCGA Research Network: http://cancergenome.nih.gov/. Source data for Figs. 1–5 and Extended Data Figs. 1–5, 7, 9 and 10 are provided with the paper. Independent validators for Figs. 1–5 are provided as Supplementary Figs. 1–5. All other data that support the findings of this study are available from the corresponding author upon request.
References
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Li, X.-D. et al. Pivotal roles of cGAS–cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341, 1390–1394 (2013).
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Ahn, J., Gutman, D., Saijo, S. & Barber, G. N. STING manifests self DNA-dependent inflammatory disease. Proc. Natl Acad. Sci. USA 109, 19386–19391 (2012).
Woo, S. R. et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity 41, 830–842 (2014).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Ablasser, A. et al. Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503, 530–534 (2013).
Patel, S. J., King, K. R., Casali, M. & Yarmush, M. L. DNA-triggered innate immune responses are propagated by gap junction communication. Proc. Natl Acad. Sci. USA 106, 12867–12872 (2009).
Patel, S. J. et al. Gap junction inhibition prevents drug-induced liver toxicity and fulminant hepatic failure. Nat. Biotechnol. 30, 179–183 (2012).
Chen, Q. et al. Carcinoma-astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature 533, 493–498 (2016).
Gentili, M. et al. Transmission of innate immune signaling by packaging of cGAMP in viral particles. Science 349, 1232–1236 (2015).
Bridgeman, A. et al. Viruses transfer the antiviral second messenger cGAMP between cells. Science 349, 1228–1232 (2015).
Marcus, A. et al. Tumor-derived cGAMP triggers a STING-mediated interferon response in non-tumor cells to activate the NK cell response. Immunity 49, 754–763 (2018).
Li, L. et al. Hydrolysis of 2′3′-cGAMP by ENPP1 and design of nonhydrolyzable analogs. Nat. Chem. Biol. 10, 1043–1048 (2014).
Gao, P. et al. Structure–function analysis of STING activation by c[G(2′,5′) pA(3′,5′)p] and targeting by antiviral DMXAA. Cell 154, 748–762 (2013).
Corrales, L. et al. Direct activation of STING in the tumor microenvironment leads to potent and systemic tumor regression and immunity. Cell Rep. 11, 1018–1030 (2015).
Ritchie, C., Cordova, A. F., Hess, G. T., Bassik, M. C. & Li, L. SLC19A1 is an importer of the immunotransmitter cGAMP. Mol. Cell 75, 372–381 (2019).
Luteijn, R. D. et al. SLC19A1 transports immunoreactive cyclic dinucleotides. Nature 573, 434–438 (2019).
Jønsson, K. L. et al. IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat. Commun. 8, 1–17 (2017).
Gao, D. et al. Activation of cyclic GMP-AMP synthase by self-DNA causes autoimmune diseases. Proc. Natl Acad. Sci. USA 112, E5699–E5705 (2015).
Sun, W. et al. ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc. Natl Acad. Sci. USA 106, 8653–8658 (2009).
Belli, S., van Driel, I. & Goding, J. Identification and characterization of a soluble form of the plasma cell membrane glycoprotein PC-1. Eur. J. Biochem. 217, 421–428 (1993).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Harding, S. M. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
Zhou, W. et al. Structure of the human cGAS–DNA complex reveals enhanced control of immune surveillance. Cell 174, 300–311 (2018).
Zhao, Y. J., Lam, C. M. C. & Lee, H. C. The membrane-bound enzyme CD38 exists in two opposing orientations. Sci. Signal. 5, ra67 (2012).
Patel, S. D. et al. Quinazolin-4-piperidin-4-methyl sulfamide PC-1 inhibitors: alleviating hERG interactions through structure based design. Bioorganic Med. Chem. Lett. 19, 3339–3343 (2009).
Shayhidin, E. E. et al. Quinazoline-4-piperidine sulfamides are specific inhibitors of human NPP1 and prevent pathological mineralization of valve interstitial cells. Br. J. Pharmacol. 172, 4189–4199 (2015).
Hatch, E. M., Fischer, A. H., Deerinck, T. J. & Hetzer, M. W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154, 47 (2013).
Bakhoum, S. F. et al. Numerical chromosomal instability mediates susceptibility to radiation treatment. Nat. Commun. 6, 1–10 (2015).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017).
Böttcher, J. P. & Reis e Sousa, C. The role of type 1 conventional dendritic cells in cancer immunity. Trends Cancer 4, 784–792 (2018).
Rutsch, F. et al. PC-1 nucleoside triphosphate pyrophosphohydrolase deficiency in idiopathic infantile arterial calcification. Am. J. Pathol. 158, 543–554 (2001).
Deng, L. et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity 41, 543–852 (2014).
Yang, H., Wang, H., Ren, J., Chen, Q. & Chen, Z. J. cGAS is essential for cellular senescence. Proc. Natl Acad. Sci. USA 114, E4612–E4620 (2017).
Lau, W. M. et al. Enpp1: a potential facilitator of breast cancer bone metastasis. PLoS One 8, 1–5 (2013).
Takahashi, R. U. et al. Loss of microRNA-27b contributes to breast cancer stem cell generation by activating ENPP1. Nat. Commun. 6, 1–15 (2015).
Umar, A. et al. Identification of a putative protein profile associated with tamoxifen therapy resistance in breast cancer. Mol. Cell Proteomics 8, 1278–1294 (2009).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Negrini, S., Gorgoulis, V. G. & Halazonetis, T. D. Genomic instability—an evolving hallmark of cancer. Nat. Rev. Mol. Cell Biol. 11, 220–228 (2010).
Bakhoum, S. F. & Cantley, L. C. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell 174, 1347–1360 (2018).
Konno, H., Konno, K. & Barber, G. N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155, 688–698 (2013).
Xu, M. M. et al. Dendritic cells but not macrophages sense tumor mitochondrial DNA for cross-priming through signal regulatory protein A signaling. Immunity 47, 363–373 (2017).
Vanpouille-Box, C. et al. DNA exonuclease Trex1 regulates radiotherapy-induced tumour immunogenicity. Nat. Commun. 8, 15618 (2017).
Lau, L., Gray, E. E., Brunette, R. L. & Stetson, D. B. DNA tumor virus oncogenes antagonize the cGAS–STING DNA-sensing pathway. Science 350, 568–571 (2015).
Vilalta, M., Rafat, M., Giaccia, A. J. & Graves, E. E. Recruitment of circulating breast cancer cells is stimulated by radiotherapy. Cell Rep. 8, 402–409 (2014).
Kato, K. et al. Expression, purification, crystallization and preliminary X-ray crystallographic analysis of Enpp1. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 68, 778–782 (2012).
Amend, S. R., Valkenburg, K. C. & Pienta, K. J. Murine hind limb long bone dissection and bone marrow isolation. J. Vis. Exp. https://doi.org/10.3791/53936 (2016).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Kato, K. et al. Crystal structure of Enpp1, an extracellular glycoprotein involved in bone mineralization and insulin signaling. Proc. Natl Acad. Sci. USA 109, 16876–16881 (2012).
Acknowledgements
This work is dedicated to T. Mitchison to celebrate his 60th birthday and his remarkable achievements in understanding biochemical mechanisms of the cell. He taught L. Li the power of definitive experiments. We thank the following people (all affiliated with Stanford) for their help: the Stanford Small Animal Imaging Facility, S. Ergun for [35S]cGAMP, C. Patel for specificity assays, F. Sunden for enzyme assays, R. Stabler for chemical synthesis, N. Weng for protein purification, and C. Walsh, D. Herschlag and all members of the Li laboratory for helpful discussions. Flow cytometry analysis for this project was performed on instruments in the Stanford Shared FACS Facility. Data were collected on an instrument in the Shared FACS Facility obtained using NIH S10 Shared Instrument Grant S10RR027431-01. This research was supported by NIH grant 5F31CA239510 (J.A.C), a Xu Family Foundation Stanford Interdisciplinary Graduate Fellowship affiliated with Stanford ChEM-H (J.A.C.), NIH grant DP2CA228044 (L.L.), Department of Defense grant W81XWH-18-1-0041 (L.L.), NSF GRFP DGE1656518 (J.A.B.), U19AI109662 (J.S.G.), R01CA197136 and S10OD018208 (E.E.G.) and K99CA201304 (M.R.).
Author information
Authors and Affiliations
Contributions
J.A.C., V.B., K.E.S., M.S. and L.L. designed the study. J.A.C., V.B., K.E.S., K.C.N., G.S., J.A.B., M.R. and L.L. performed experiments and analyzed data. R.E. performed statistical analysis. J.A.C., V.B. and L.L. wrote the manuscript. J.S.G. supervised K.C.N.; E.E.G. supervised M.R. and R.E. All authors discussed the findings and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.S. and L.L. are scientific cofounders of Angarus Therapeutics, which has exclusive licensing rights to patent PCT/US2018/50018. J.A.C., V.B., K.E.S., M.S. and L.L. are inventors on patent PCT/US2018/50018.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Measuring cGAMP in 293T cGAS cell lines by LC-MS/MS.
a, Chemical structures of cGAMP (top) and single isotopically-labeled cGAMP (bottom) used as an internal standard at a concentration of 0.5-1 μM. b-c, Full (7-8 point) standard curves and LC traces of lowest cGAMP standards in 50/50 acetonitrile/water spiked in directly (LOQ = 4 nM) (b) or after concentrating and extracting 12.5x from complete cell culture media (LOQ = 0.3 nM from original sample, 4 nM in concentrated sample) (c). IS = internal standard. R2 = coffecient of determination, determined after linear least squares regression with 1/Y2 weighting. Data are from 1 experiment, representative of (b) 64 independent experiments and (c) 13 independent experiments. d, Calibration of cell number to ATP concentration measured by LC-MS/MS. Data are from one experiment, representative of two independent experiments, 8 individual cell culture replicates are plotted. e, cGAS expression of 293T, 293T cGAS ENPP1−/−, and 293T cGAS ENPP1low cell lines analyzed by western blot (left; full scan of blot available in Source Data). ENPP1 hydrolysis activity of 32P-cGAMP in whole cell lysates from 1 million each of 293T cGAS, 293T cGAS ENPP1−/−, and 293T cGAS ENPP1low cells, measured by TLC and autoradiography (right). Data are from 1 experiment, representative of 5 independent experiments.
Extended Data Fig. 2 Development of assays to investigate the mechanism of cGAMP export.
a, Schematic of experiment for b. CD14+ PMBCs were stimulated with increasing concentrations of extracellular cGAMP for 16 h. b, IFNB1 mRNA levels were normalized to indicated gene and fold induction was calculated relative to untreated CD14+ cells. Each donor was performed as one independent experiment; 2 qPCR replicates are plotted with a bar representing the mean. c, Coomassie gel of recombinant mouse ENPP1 purified from media; elution fractions were pooled before use (left). 32P-cGAMP degradation by mouse ENPP1 analyzed by TLC (right). Data are representative of 10 independent experiments. d–e, 293T cGAS ENPP1low cells were incubated with serum-free ATP depletion media (no glucose, 6 mM 2-deoxy-D-glucose, 5 mM NaN3) or serum-free complete media for 1 hour. Levels of analytes were measured: (d) ATP by LC-MS/MS, total protein by BCA, cell death by extracellular lactate dehydrogenase activity, and (e) cGAMP by LC-MS/MS. BQL = below quantification limit. Data are from 1 experiment; 3 cell culture replicates are plotted (except for extracellular cGAMP, where 2 cell culture replicates are plotted). ATP data are representative of 3 independent validations (shown in Supplementary Fig. 5).
Extended Data Fig. 3 Characterization of ENPP1 inhibitors QS1 and STF-1084.
a, Structure of QS1. b, Inhibition by QS1 (compared to STF-1084, replotted from Fig. 3e). in vitro (32P-cGAMP TLC assay, pH 7.5, purified mouse ENPP1): Ki,app = 6.4 μM, 2 independent experiments are plotted with a bar representing the mean. c, Intracellular, extracellular, and total cGAMP for 293T cGAS ENPP1−/− cells transfected with pcDNA6 (empty or containing human ENPP1) and treated ± QS1 after 24 hours. Data are from 1 experiment, representative of 2 independent validations; 2 cell culture replicates are plotted with a bar representing the mean. d, Permeability of compounds in Caco-2 assay. PA = peak area, IS = internal standard. Compounds were incubated on the apical side of a Caco-2 monolayer for 2 hours. Compound concentration on the basolateral side was monitored by LC-MS/MS. Apparent permeability rates (Papp) were calculated from the slope. Data are representative of 2 independent experiments; single points are plotted. e, Inhibitory activity of STF-1084 against alkaline phosphatase (ALPL) and ENPP2. Data are from 1 experiment, representative of 2 independent validations; 2 reaction replicates are plotted, except for ENPP2 100 uM and 32 nM, where 1 point is plotted. f, Kinome interaction map (468 kinases tested) for STF-1084 depicting kinase inhibition as a percent of control. Data visualized with the TREEspot Software Tool and reprinted with permission from KINOMEscan, a division of DiscoveRx Corporation. ©DiscoveRX Corporation 2010.
Extended Data Fig. 4 Intracellular and extracellular cGAMP from cancer cell lines.
a, cGAS expression of 4T1-luc WT and 4T1-luc shcGAS analyzed by western blot (full scan of blot available in Source Data). Intracellular cGAMP (chromatograms and quantification displayed) in 4T1-luc WT and shcGAS cells without exogenous stimulation. Concentration reported in units of molecules/cell and nM (estimated using cell volume = 4 pL). IS = internal standard. Data are from 1 experiment; 2 cell culture replicates are plotted. b, Extracellular cGAMP produced by MC38 cells over 48 hours. At time 0, cells were refreshed with media supplemented with 50 μM STF-1084. Line depicts linear regression. Data are from one experiment representative of two independent validations; 2 cell culture replicates are plotted with a bar representing the mean. c, Chromatograms for E0771, Panc02, and NMuMG cell lysate from LC-MS/MS. E0771 cell lysate was spiked with 40 nM cGAMP to determine limit of quantification. IS = internal standard. Data are representative of 2 independent validations. d, cGAS and STING expression of 4T1-luc, MC38, E0771, Panc02, and Neuro-2a cell lines analyzed by western blot (full scan of blot available in Source Data). Data are representative of 3 independent validations. e, Cell viability of cancer cell lines 4T1-luc and E0771 measured by lactate dehydrogenase extracellular activity compared to intracellular activity. At time 0, cells were left untreated or treated with IR (8 Gy or 20 Gy) and refreshed with media supplemented with 50 μM STF-1084. Data are from 1 experiment representative of two independent validations; 2 cell culture replicates are plotted with a bar representing the mean.
Extended Data Fig. 5 Validation of Cgas−/− cell lines and tools to neutralize extracellular cGAMP.
a, E0771 (left) and 4T1-luc (right) Cgas−/− cells subcloned from CRISPR knockout pools (full scan of blot available in Source Data). Data are representative of two independent validations. Six E0771 Cgas−/− subclones (1, 2, 4, 6, 8, and 9) were pooled before injection into mice. Two 4T1-luc Cgas−/− subclones were pooled before injection into mice. b, 293T cGAS ENPP1low cells were transfected with empty pcDNA6 (0.5 μg/mL) and incubated for 24 hours. Conditioned media was treated with nonbinding or neutralizing STING (1 hour pretreatment) and then incubated with CD14+ PBMCs for 16 h. Extracellular cGAMP measured by LC-MS/MS and IFNB1 expression (mRNA levels were normalized to CD14 and fold induction calculated relative to untreated CD14+ cells). Data are from 1 experiment (supported by data in Fig. 5e and Extended Data Fig. 5c,d); 2 cell culture replicates are plotted. c, Cxcl10 mRNA fold induction (normalized to Actb and untreated cells) in primary mouse bone marrow cells treated with 20 μM cGAMP in the presence of neutralizing or non-binding STING (100 μM) for 16 h. Data are from 1 experiment (supported by data in Fig. 5e and Extended Data Fig. 5b,d); cell culture replicates plotted are (from left to right) 3, 2, 2, 3, 2, 2. d, Mouse bone marrow-derived macrophages were incubated with 10 uM cGAMP and indicated concentrations of neutralizing STING protein for 2 hours. Levels of pIRF3 and IRF3 were analyzed by western blotting. Data are from 1 experiment (supported by data in Fig. 5e and Extended Data Fig. 5b,c) (full scan of blot available in Source Data).
Extended Data Fig. 6 FACS analysis and tumor growth following extracellular cGAMP depletion in untreated tumors.
a, FACS gating scheme for experiments in Fig. 5f–i, Fig. 7b, c and Extended Data Fig. 6b. b, WT or Cgas−/− E0771 cells (1x106) were orthotopically injected into WT, Cgas−/− or Sting1−/− C57BL/6 J mice on day 0. Neutralizing STING or non-binding STING was intratumorally injected on day 2. Sample sizes of n mice, from left to right (non-binding STING, neutralizing STING): n = (5, 5); (4, 5); (5, 5); (5, 4). Mean ± SD, unpaired two-tailed t test with Welch’s correction. c, Established E0771 tumors (100 ± 20 mm3) were injected with non-binding (n = 8 mice) or neutralizing STING (n = 9 mice) every other day for the duration of the experiment. Tumor volumes for individual mice are shown. P value for tumor volume determined by pairwise comparisons using post hoc tests with a Tukey adjustment and for Kaplan Meier curve determined using the Log-rank Mantel-Cox test. d, ENPP1 activity in 4T1-luc, E0771, and MDA-MB-231 cells using the 32P-cGAMP degradation assay. Data from 1 experiment, representative of 3 independent validations
Extended Data Fig. 7 Cytokine production in cancer cell lines treated with ionizing radiation.
CXCL10 production by cancer cell lines 4T1-luc and E0771 measured by ELISA. At time 0, cells were left untreated or treated with IR (8 Gy or 20 Gy) and refreshed with media supplemented with 50 μM STF-1084. Data are from 1 experiment, representative of 2 independent validations; 2 cell culture replicates are plotted with a bar representing the mean.
Extended Data Fig. 8 FACS analysis of tumors treated with ionizing radiation.
a, FACS gating scheme for live dead analysis in established tumors. E0771 cells (5x104) were orthotopically injected into WT C57BL/6 J mice. The tumors were treated with IR (8 Gy) when they reached 100 ± 20 mm3 and harvested and analyzed by FACS 24 h after IR. For caspase activity, a single cell suspension was incubated for 1 h with the FAM-FLICA Poly Caspase substrate before FACS stain and analysis. Mean ± SD, unpaired two-tailed t test with Welch’s correction. b, FACS gating scheme for experiments in Fig. 6c,d
Extended Data Fig. 9 4T1-luc Enpp1−/− tumor growth.
a, Validating Enpp1−/− 4T1-luc clones (11 clones were pooled) using the 32P-cGAMP degradation assay (3 day incubation). Lysates were normalized by protein concentrations. Data are from 1 experiment, representative of three independent validations; 2 technical protein concentration replicates are plotted with a bar representing the mean. b, Established 4T1-luc WT (harboring scrambled sgRNA) (n = 10 mice) or Enpp1−/− tumors (n = 10 mice) (100 ± 20 mm3) were monitored without treatment. Tumor volumes for individual mice are shown. c, Established 4T1-luc WT (harboring scrambled sgRNA) (n = 10 mice) or Enpp1−/− tumors (n = 10 mice) (100 ± 20 mm3) were treated with IR (20 Gy) and monitored. Tumor volumes for individual mice are shown. d, Established 4T1-luc WT (harboring scrambled sgRNA) (n = 9 mice) or Enpp1−/− tumors (n = 9 mice) (100 ± 20 mm3) were treated with three intratumoral injections of 10 µg cGAMP on day 2, 4, and 7 and monitored. Tumor volumes for individual mice are shown.
Extended Data Fig. 10 A systemic ENPP1 inhibitor delays Panc02 tumor growth as a single agent and synergizes with ionizing radiation.
a, Structure of ENPP1 inhibitor STF-1623. b, Inhibition by STF-1623. In vitro (32P-cGAMP TLC assay, pH 7.5, purified mouse ENPP1: Ki,app = 16 nM. 3 independent experiments are plotted. In cells (cGAMP export assay, human ENPP1 transfected into 293 T cGAS ENPP1−/− cells): IC50 = 68 nM. Data are from 1 experiment; 2 cell culture replicates are plotted. c, Mean apparent permeability (Papp) for STF-1623 and controls. For PAMPA and MDCK, mean was calculated from 2 cell culture replicates, 1 experiment. For Caco-2, mean was calculated from 2 independent experiments (data for atenolol and propranolol are reproduced from Fig. 3f for comparison). d, Permeability of compounds in Caco-2 assay. PA = peak area, IS = internal standard. Compounds were incubated on the apical side of a Caco-2 monolayer for 2 hours. Compound concentration on the basolateral side was monitored by LC-MS/MS. Apparent permeability rates (Papp) were calculated from the slope. Data are representative of 2 independent experiments; single points are plotted. e, PBMCs were incubated with STF-1623 for 16 h. Data are from one experiment; 2 cell culture replicates are plotted. f, Kinome interaction map (468 kinases tested) for STF-1623 depicting kinase inhibition as a percent of control. Data visualized with the TREEspot Software Tool and reprinted with permission from KINOMEscan, a division of DiscoveRx Corporation. ©DiscoveRX Corporation 2010. g, Intracellular and extracellular cGAMP concentrations for 293T cGAS ENPP1−/− cells transfected with pcDNA6 (empty or containing human ENPP1) and treated ± 2 μM STF-1623 after 24 hours. Data are from 1 experiment; cell culture replicates plotted from left to right are (intracellular) 2, 3, 2, 5 and (extracellular) 2, 3, 2, 3. h, PBMCs were electroporated ± 200 nM cGAMP and incubated ± 2 μM STF-1623 for 16 h. IFNB1 and CXCL10 mRNA levels were normalized to ACTB and fold induction calculated relative to untreated cells. Data are from 1 experiment; 2 cell culture replicates are plotted. i, Mice were injected subcutaneously with 300 mg/kg STF-1623 at time 0. At indicated times, the mouse was sacrificed, blood was drawn by cardiac puncture, and serum isolated after clotting. STF-1623 concentrations were measure by LC-MS/MS. Data are from 1 experiment; 2 mice per time point are plotted except for 8 hours, where 3 mice are plotted. j, Mice bearing established subcutaneous Panc02 tumors (100 ± 20 mm3) were implanted with a subcutaneous pump containing STF-1623 (50 mg/kg/day) on day 0 and left untreated or treated with IR (20 Gy) on day 1. No IR: n = 10 mice, no IR + STF-1623: n = 10 mice, IR (20 Gy): n = 10 mice, IR (20 Gy) + STF-1623: n = 15 mice. Pumps were removed on day 8. Tumor volumes for individual mice are shown. P value determined by pairwise comparisons using post hoc tests with a Tukey adjustment. In b and g, cGAMP is measured by LC-MS/MS. BQL = below quantification limit. In b,e,g,h, and i, replicates are plotted individually with a bar representing the mean.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 1
Numerical source data.
Source Data Fig. 2
Numerical source data.
Source Data Fig. 3
Numerical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Numerical source data.
Source Data Fig. 5
Numerical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 1
Numerical source data.
Source Data Extended Data Fig. 2
Numerical source data.
Source Data Extended Data Fig. 3
Numerical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4
Numerical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 5
Numerical source data.
Source Data Extended Data Fig. 7
Numerical source data.
Source Data Extended Data Fig. 9
Numerical source data.
Source Data Extended Data Fig. 10
Numerical source data.
Rights and permissions
About this article
Cite this article
Carozza, J.A., Böhnert, V., Nguyen, K.C. et al. Extracellular cGAMP is a cancer-cell-produced immunotransmitter involved in radiation-induced anticancer immunity. Nat Cancer 1, 184–196 (2020). https://doi.org/10.1038/s43018-020-0028-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-0028-4
This article is cited by
-
Nanomaterial-encapsulated STING agonists for immune modulation in cancer therapy
Biomarker Research (2024)
-
Second messenger 2'3'-cyclic GMP-AMP (2'3'-cGAMP): the cell autonomous and non-autonomous roles in cancer progression
Acta Pharmacologica Sinica (2024)
-
Discovery of VH domains that allosterically inhibit ENPP1
Nature Chemical Biology (2024)
-
CD4+ T cells produce IFN-I to license cDC1s for induction of cytotoxic T-cell activity in human tumors
Cellular & Molecular Immunology (2024)
-
Cholesterol-binding motifs in STING that control endoplasmic reticulum retention mediate anti-tumoral activity of cholesterol-lowering compounds
Nature Communications (2024)