Abstract
The transcriptomic classification of glioblastoma (GBM) has failed to predict survival and therapeutic vulnerabilities. A computational approach for unbiased identification of core biological traits of single cells and bulk tumors uncovered four tumor cell states and GBM subtypes distributed along neurodevelopmental and metabolic axes, classified as proliferative/progenitor, neuronal, mitochondrial and glycolytic/plurimetabolic. Each subtype was enriched with biologically coherent multiomic features. Mitochondrial GBM was associated with the most favorable clinical outcome. It relied exclusively on oxidative phosphorylation for energy production, whereas the glycolytic/plurimetabolic subtype was sustained by aerobic glycolysis and amino acid and lipid metabolism. Deletion of the glucose-proton symporter SLC45A1 was the truncal alteration most significantly associated with mitochondrial GBM, and the reintroduction of SLC45A1 in mitochondrial glioma cells induced acidification and loss of fitness. Mitochondrial, but not glycolytic/plurimetabolic, GBM exhibited marked vulnerability to inhibitors of oxidative phosphorylation. The pathway-based classification of GBM informs survival and enables precision targeting of cancer metabolism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Single-cell RNA-seq datasets 1, 2 and 3 are available from Gene Expression Omnibus (GEO, accession nos. GSE117891, GSE103224 and GSE131928, respectively). Single-cell RNA-seq read count data for the additional samples in dataset 2 (PJ053, PJ069 and PW032.706) and sequencing data for PDC samples (gene expression read counts, genetic mutations and copy number segmentation) have been submitted to Synapse (http://synapse.org, accession no. syn22314624). TCGA GBM data (Agilent chip G4502A and RNA-seq gene expression, somatic mutation calls, Affymetrix SNP 6.0 array copy number segmentation, Illumina Infinium Human Methylation 450k DNA methylation, Agilent miRNA and RPPA quantification) are available from the GDC Data Portal (https://portal.gdc.cancer.gov/). The CGGA GBM dataset is available from the CGGA data portal (http://cgga.org.cn/index.jsp). Data from ref. 69 are available from GEO (accession no. GSE13041). Data from ref. 17 are available at the Sequence Read Archive (SRA, accession no. SRP074425), European Genome-phenome Archive (EGA, accession nos. EGAS00001001033, EGAS00001000579, EGAS00001001044, EGAS00001001041 and EGAS00001001800) and the Japanese Genotype–Phenotype Archive (accession no. JGAS00000000004). The GBM longitudinal GLASS dataset is available from Synapse (http://synapse.org/glass). Source data for Figs. 5a–f,h,i, 6c, 7a–c,e–i and 8a–d,f,g,i–k and Extended Data Figs. 9b,c and 10g,h are provided as with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability
The source code used for the pathway-based classification and the code related to the figures of the study are available at GitHub: https://github.com/miccec/GBMstates.
References
Cieslik, M. & Chinnaiyan, A. M. Cancer transcriptome profiling at the juncture of clinical translation. Nat. Rev. Genet. 19, 93–109 (2018).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56 (2017).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849 (2019).
Kim, S., Kon, M. & DeLisi, C. Pathway-based classification of cancer subtypes. Biol. Direct 7, 21 (2012).
Yu, K. et al. Surveying brain tumor heterogeneity by single-cell RNA-sequencing of multi-sector biopsies. Natl Sci. Rev. 7, 1306–1318 (2020).
Yuan, J. et al. Single-cell transcriptome analysis of lineage diversity in high-grade glioma. Genome Med. 10, 57 (2018).
Leone, G., Abla, H., Gasparre, G., Porcelli, A. M. & Iommarini, L. The Oncojanus paradigm of respiratory complex I. Genes (Basel) 9, 243 (2018).
Venkataramani, V. et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature 573, 532–538 (2019).
Chen, H. et al. Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM. Nat. Commun. 10, 1903 (2019).
Phillips, H. S. et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9, 157–173 (2006).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Caruso, F. P. et al. A map of tumor–host interactions in glioma at single-cell resolution. Gigascience 9, giaa109 (2020).
Frattini, V. et al. A metabolic function of FGFR3–TACC3 gene fusions in cancer. Nature 553, 222–227 (2018).
Zhang, J. et al. The combination of neoantigen quality and T lymphocyte infiltrates identifies glioblastomas with the longest survival. Commun. Biol. 2, 135 (2019).
Venteicher, A. S. et al. Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355, eaai8478 (2017).
Wang, J. et al. Clonal evolution of glioblastoma under therapy. Nat. Genet. 48, 768–776 (2016).
D’Angelo, F. et al. The molecular landscape of glioma in patients with neurofibromatosis 1. Nat. Med. 25, 176–187 (2019).
Koh, E. H. et al. Mitochondrial activity in human white adipocytes is regulated by the ubiquitin carrier protein 9/microRNA–30a axis. J. Biol. Chem. 291, 24747–24755 (2016).
Koh, E. H. et al. miR-30a remodels subcutaneous adipose tissue inflammation to improve insulin sensitivity in obesity. Diabetes 67, 2541–2553 (2018).
Li, L. et al. miR-30a-5p suppresses breast tumor growth and metastasis through inhibition of LDHA-mediated Warburg effect. Cancer Lett. 400, 89–98 (2017).
Chan, S. Y. et al. MicroRNA-210 controls mitochondrial metabolism during hypoxia by repressing the iron-sulfur cluster assembly proteins ISCU1/2. Cell Metab. 10, 273–284 (2009).
Favaro, E. et al. MicroRNA-210 regulates mitochondrial free radical response to hypoxia and Krebs cycle in cancer cells by targeting iron sulfur cluster protein ISCU. PLoS ONE 5, e10345 (2010).
Papagiannakopoulos, T., Shapiro, A. & Kosik, K. S. MicroRNA-21 targets a network of key tumor-suppressive pathways in glioblastoma cells. Cancer Res. 68, 8164–8172 (2008).
Bobbili, M. R., Mader, R. M., Grillari, J. & Dellago, H. OncomiR-17-5p: alarm signal in cancer? Oncotarget 8, 71206–71222 (2017).
Sun, G. et al. miR-137 forms a regulatory loop with nuclear receptor TLX and LSD1 in neural stem cells. Nat. Commun. 2, 529 (2011).
Liu, Y. et al. XBP1 silencing decreases glioma cell viability and glycolysis possibly by inhibiting HK2 expression. J. Neurooncol. 126, 455–462 (2016).
Koo, J. H. & Guan, K. L. Interplay between YAP/TAZ and metabolism. Cell Metab. 28, 196–206 (2018).
Gao, Z. Y. et al. Metformin induces apoptosis via a mitochondria-mediated pathway in human breast cancer cells in vitro. Exp. Ther. Med. 11, 1700–1706 (2016).
Hirata, T. et al. Stem cell factor induces outgrowth of c-kit-positive neurites and supports the survival of c-kit-positive neurons in dorsal root ganglia of mouse embryos. Development 119, 49–56 (1993).
Strauss, B. et al. Cyclin B1 is essential for mitosis in mouse embryos, and its nuclear export sets the time for mitosis. J. Cell Biol. 217, 179–193 (2018).
Gong, A. H. et al. FoxM1 drives a feed-forward STAT3-activation signaling loop that promotes the self-renewal and tumorigenicity of glioblastoma stem-like cells. Cancer Res. 75, 2337–2348 (2015).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Cluntun, A. A., Lukey, M. J., Cerione, R. A. & Locasale, J. W. Glutamine metabolism in cancer: understanding the heterogeneity. Trends Cancer 3, 169–180 (2017).
Petan, T., Jarc, E. & Jusovic, M. Lipid droplets in cancer: guardians of fat in a stressful world. Molecules 23, 1941 (2018).
Fam, T. K., Klymchenko, A. S. & Collot, M. Recent advances in fluorescent probes for lipid droplets. Materials (Basel) 11, 1768 (2018).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Muller, F. L. et al. Passenger deletions generate therapeutic vulnerabilities in cancer. Nature 488, 337–342 (2012).
Trifonov, V., Pasqualucci, L., Dalla Favera, R. & Rabadan, R. MutComFocal: an integrative approach to identifying recurrent and focal genomic alterations in tumor samples. BMC Syst. Biol. 7, 25 (2013).
Sarto Basso, R., Hochbaum, D. S. & Vandin, F. Efficient algorithms to discover alterations with complementary functional association in cancer. PLoS Comput. Biol. 15, e1006802 (2019).
Duncan, C. G. et al. Integrated genomic analyses identify ERRFI1 and TACC3 as glioblastoma-targeted genes. Oncotarget 1, 265–277 (2010).
Barthel, F. P. et al. Longitudinal molecular trajectories of diffuse glioma in adults. Nature 576, 112–120 (2019).
Jolly, C. & Van Loo, P. Timing somatic events in the evolution of cancer. Genome Biol. 19, 95 (2018).
Shimokawa, N. et al. Past-A, a novel proton-associated sugar transporter, regulates glucose homeostasis in the brain. J. Neurosci. 22, 9160–9165 (2002).
Srour, M. et al. Dysfunction of the cerebral glucose transporter SLC45A1 in individuals with intellectual disability and epilepsy. Am. J. Hum. Genet. 100, 824–830 (2017).
Webb, B. A., Chimenti, M., Jacobson, M. P. & Barber, D. L. Dysregulated pH: a perfect storm for cancer progression. Nat. Rev. Cancer 11, 671–677 (2011).
Molina, J. R. et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 24, 1036–1046 (2018).
Wheaton, W. W. et al. Metformin inhibits mitochondrial complex I of cancer cells to reduce tumorigenesis. eLife 3, e02242 (2014).
Skrtic, M. et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 20, 674–688 (2011).
Criddle, D. N. et al. Menadione-induced reactive oxygen species generation via redox cycling promotes apoptosis of murine pancreatic acinar cells. J. Biol. Chem. 281, 40485–40492 (2006).
Altman, B. J., Stine, Z. E. & Dang, C. V. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat. Rev. Cancer 16, 619–634 (2016).
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1alpha, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011).
Richardson, R. B. & Harper, M. E. Mitochondrial stress controls the radiosensitivity of the oxygen effect: implications for radiotherapy. Oncotarget 7, 21469–21483 (2016).
Kim, W. et al. Cellular stress responses in radiotherapy. Cells 8, 1105 (2019).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Malta, T. M. et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173, 338–354 (2018).
Momcilovic, M. et al. In vivo imaging of mitochondrial membrane potential in non-small-cell lung cancer. Nature 575, 380–384 (2019).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Sack, L. M. et al. Profound tissue specificity in proliferation control underlies cancer drivers and aneuploidy patterns. Cell 173, 499–514 (2018).
Solimini, N. L. et al. Recurrent hemizygous deletions in cancers may optimize proliferative potential. Science 337, 104–109 (2012).
Dong, J. et al. Single-cell RNA-seq analysis unveils a prevalent epithelial/mesenchymal hybrid state during mouse organogenesis. Genome Biol. 19, 31 (2018).
Gao, S. et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 20, 721–734 (2018).
Dobin, A. & Gingeras, T. R. Mapping RNA-seq reads with STAR. Curr. Protoc Bioinformatics 51, 11.14.11–11.14.19 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 44, e71 (2016).
Risso, D., Schwartz, K., Sherlock, G. & Dudoit, S. GC-content normalization for RNA-seq data. BMC Bioinformatics 12, 480 (2011).
Zhao, Z. et al. Comprehensive RNA-seq transcriptomic profiling in the malignant progression of gliomas. Sci Data 4, 170024 (2017).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Lee, Y. et al. Gene expression analysis of glioblastomas identifies the major molecular basis for the prognostic benefit of younger age. BMC Med. Genomics 1, 52 (2008).
Hussain, S. F. et al. The role of human glioma-infiltrating microglia/macrophages in mediating antitumor immune responses. Neuro. Oncol. 8, 261–279 (2006).
Quail, D. F. & Joyce, J. A. The microenvironmental landscape of brain tumors. Cancer Cell 31, 326–341 (2017).
Mall, R. et al. RGBM: regularized gradient boosting machines for identification of the transcriptional regulators of discrete glioma subtypes. Nucleic Acids Res. 46, e39 (2018).
Lambert, S. A. et al. The human transcription factors. Cell 175, 598–599 (2018).
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Wingender, E. The TRANSFAC project as an example of framework technology that supports the analysis of genomic regulation. Brief. Bioinformatics 9, 326–332 (2008).
Lee, S. B. et al. An ID2-dependent mechanism for VHL inactivation in cancer. Nature 529, 172–177 (2016).
Acknowledgements
We thank D. Bigner (Duke University Medical Center, Durham, NC, USA) for the generous gift of H423 and H502 glioma cells. This work was supported by NIH grant nos. R01CA101644, U54CA193313, R01CA131126 and R01CA239721 (to A.L.); R01CA178546, U54CA193313, R01CA179044, R01CA190891 and R01CA239698; NCI P30 Supplement GBM CARE-HOPE; The Chemotherapy Foundation (to A.I.); and Italian Association for Cancer Research (AIRC) no. IG 2018 ID. 21846 (to M.C.). S.M. is recipient of a fellowship from AIRC.
Author information
Authors and Affiliations
Contributions
A.I. and A.L. conceived and coordinated the studies and provided overall supervision. L.G. and S.M. developed and performed bioinformatics analyses of single cells and bulk tumors, with the assistance of F.D. for longitudinal and SLC45A1 studies, of F.P.C. for studies on cells of the tumor microenvironment and M.C. for clustering and quality control method benchmarking. Y.T.O. and A.K. performed cell, molecular biology and metabolic assays, with the collaboration of R.D.N. and B.F. M.L.S., P.S., K.Y., J.Y., W.Z., F.T., X.-D.S. and T.J. provided single-cell data. M.S., F.B. and A.L.D.S. assisted in the analysis of clinical data. L.G., S.M. and M.C. wrote the computational sections. A.I. and A.L. wrote, edited and revised the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.L. and A.I. are inventors of a biomarker technology that has been licensed to QIAGEN. A.I. received sponsored research funding from AstraZeneca and Taiho Pharmaceutical and has served as a paid consultant/advisor to AIMEDBIO Inc. A.L. received sponsored research funding from Celgene. All other authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The computational framework of scBiPaD.
Step 1: identification of cell sub-populations of cells in each individual tumor that share activation of similar biological functions; Step 2: determination of enriched biological pathways in each cell sub-population by defining cluster-specific ranked-lists; Step 3: identification of cell sub-populations that share coherent biological functions across multiple tumors. In Step 1-i, the ranked list for each cell in each tumor is obtained by standardizing and ranking genes. The activity matrix (NES) of all cells composing each tumor is obtained by calculating the single-sample activity of all the 5,032 biological pathways with ssMWW-GST (Step 1-ii) and used to generate the Euclidean distance between every pair of cells in each tumor (Step 1-iii). Finally, the cell sub-populations of each tumor are identified by applying the consensus clustering on the basis of the Euclidean distance of the NES (Step 1-iv). In the following step (Step 2-i), the MWW-score is used to generate a cluster-specific ranked-list of genes for each cell sub-population by comparing the expression profiles of the cells in the cluster with all other cells in the same tumor. The enriched biological pathways of each cell sub-populations are derived in Step 2-ii by using MWW-GST as in Step 1-ii. Each cell sub-population is then represented by a binary vector, with 1 indicating the enriched biological pathways (Step 3-i) and the binary matrix is used in Step 3-ii to derive the Jaccard distance. In the last step, 3-iii, cell sub-populations are clustered by Jaccard distance using consensus clustering.
Extended Data Fig. 2 Expression of subtype associated markers and mapping of marker genes on the population structure of neurodevelopmental subtype.
a, Rank order plot of changes in genes expressed in MTC cells versus the other groups. Genes are ranked from left to right in increasing expression order. Red dots indicate mitochondrial respiratory complex I genes differentially expressed in each single cell dataset [n = 2,799 cells for dataset 1, n = 9,652 cells for dataset 2, n = 4,916 cells for dataset 3; log2(FC) > 0.3 and FDR < 0.05, two-sided MWW test]. Representative genes upregulated in at least two out of three datasets are shown. Colors indicate complex I structural classes. b, Rank order plot of changes of genes expressed in NEU cells versus the other groups. Genes are ranked as in a. Red dots indicate neurotransmitter receptors differentially expressed in each single cell dataset [n = 2,799 cells for dataset 1, n = 9,652 cells for dataset 2, n = 4,916 cells for dataset 3; log2(FC) > 0.3 and FDR < 0.05, two-sided MWW test]. For each dataset, upregulated genes representing distinct neurotransmitter receptor families are indicated by different colors. c, Rank order plot of changes of genes expressed in PPR cells versus the other groups. Genes are ranked as in a, b. Red dots indicate neural progenitor marker genes differentially expressed in each single cell dataset [n = 2,799 cells for dataset 1, n = 9,652 cells for dataset 2, n = 4,916 cells for dataset 3; log2(FC) > 0.3 and FDR < 0.05, two-sided MWW test]. Representative genes differentially expressed in at least two out of three single cell datasets are indicated. d, Sankey diagram showing subtype assignment of single glioma cells according to scBiPaD classification and the described cell states4. e, Bar plot of the number of tumors and states in each of the 36 samples of the single cell cohort. f, Bar plot showing functional cell state (at least 15% of cells in the sample) composition of 36 GBM samples. g, Stream plots of proliferation markers expressed by the PPR cells at the tumor core. h, Stream plots of neural progenitor markers. Expression overlaps with proliferation markers and is excluded from the more differentiated cells at the tumor periphery. The newly born neuron marker TBR1 is expressed in a subset of cells of the neurodevelopment branch. i, Stream plots of synaptic and neurotransmitter receptor genes in non-proliferative cells at the invasive rim. Color scale indicates the log2 normalized expression of the indicated gene.
Extended Data Fig. 3 Analysis of survival-associated biological pathways in single glioma cells.
a, Consensus clustering of 103 cell sub-populations from the three single cell datasets obtained using 192 biological pathways significantly associated with patient survival. Columns and rows are cell sub-populations. Left track: red, GPM; green, MTC; blue, NEU; cyan, PPR. b, Heatmap of the biological activities of cell sub-populations in a. Each group was defined by shared activated pathways among the 5,032-pathway collection (n = 103 cell sub-populations; effect size > 0.3, FDR < 0.0001, two-sided MWW test). Columns are cell sub-populations, rows are pathway activities. Pathway activity level is color-coded. Representative pathways specifically activated in each of the four functional subtypes are indicated. Left and top tracks are as in a. c, Enrichment map network of statistically significant and not redundant GO categories [logit(NES) > 0.58 and FDR < 0.05, two-sided MWW-GST] in GPM; d, MTC; e, NEU; f, PPR medoids. Nodes are GO terms and lines their connectivity. Node size is proportional to number of genes in the GO category; line thickness indicates similarity coefficient. The right-hand side of the network in c was magnified 1.5-fold for a better visualization of the significant activities.
Extended Data Fig. 4 t-SNE plot visualization of tumors and functional cell states in single glioma cells.
a, t-SNE plot of malignant cells colored by tumor from dataset 1; b, dataset 2; c, dataset 3. d, t-SNE plot of malignant cells from dataset 1 colored according to functional states; e, t-SNE plot of malignant cells from dataset 2 colored according to functional states; f, t-SNE plot of malignant cells from dataset 3 colored according to functional states. Cells concordantly classified using 5,032 or 192 pathways are colored: red, GPM; green, MTC; blue, NEU; cyan, PPR; grey, cells not concordantly classified.
Extended Data Fig. 5 Characterization of biological subtypes of bulk primary GBM.
a, Consensus clustering of 534 GBM on the activity of 192 survival-associated pathways (p < 0.05, log-rank test; p-value of individual pathways are reported in Supplementary Table 6b). Columns and rows are individual tumors. Left track: red, GPM; green, MTC; blue, NEU; cyan, PPR; black, unclassified. b, Heatmap of pathway activity in 304 classified GBM including 126 out of 192 survival-associated and differentially active pathways in the four GBM subtypes (effect size > 0.3 and FDR < 0.01, two-sided MWW test). Columns are individual tumors and rows are pathway activity. Pathways characteristically activated in each core subtype are indicated. Left and top tracks: red, GPM; green, MTC; blue, NEU; cyan, PPR. c, Heatmap of genes differentially expressed and upregulated in GBM subtypes (n = 304 tumors; Kruskal-Wallis analysis with post hoc correction by Nemenyi’s test for multiple comparison; FDR < 0.01 and log2(FC) > 0.5). Columns are individual tumors, rows are genes. Representative genes specifically upregulated in each GBM subtype are indicated. Tracks are as in b. d, Rank order plot of changes of genes expressed in GBM NEU. Genes are ranked from left to right in increasing expression order. Red dots indicate neurotransmitter receptors differentially expressed in NEU tumors and cells (n = 2,799 cells for dataset 1, n = 9,652 cells for dataset 2, n = 4,916 cells for dataset 3, n = 304 tumors for TCGA dataset; log2(FC) > 0.3, FDR < 0.05, two-sided MWW test). For each dataset, upregulated genes in neurotransmitter receptor families are indicated by colors. e, Rank order plot of changes of genes expressed in GBM PPR. Genes are ranked as in d. Red dots indicate neural progenitor genes differentially expressed in each dataset (n = 2,799 cells for dataset 1, n = 9,652 cells for dataset 2, n = 4,916 cells for dataset 3, n = 304 tumors for TCGA dataset; log2(FC) > 0.3, FDR < 0.05, two-sided MWW test). Representative genes differentially expressed in at least three datasets are indicated. f, Heatmap showing the 50 highest scoring genes of the four GBM subtypes-specific signatures. Rows are genes and columns are tumors (n = 304 tumors). Track are as in b, c. g, Two-dimensional representation of GBM subtype enrichment scores (n = 304 tumors). Quadrant are GBM subtypes, the position of dots (tumors) reflects the relative subtype-specific score of each tumor as indicated by x- and y-axes, and their color the subtype simplicity score. Gray, tumors that do not fall in the respective subtype quadrant.
Extended Data Fig. 6 Validation of the biological classification of GBM and comparison with established classifiers.
Subtype-specific gene signatures were used to classify GBM from independent cohorts. a, Heatmap of GBM from the TCGA cohort profiled by RNA-seq (n = 129 tumors). b, Heatmap of GBM from the CGGA cohort (n = 94 tumors). c, Heatmap of GBM69 (n = 158 tumors). d, Kaplan-Meier of patients in a (128 out of 129 patients with survival data available). e, Kaplan-Meier of patients in b (90 out of 94 patients with survival data available). f, Kaplan-Meier of patients in c (156 out of 158 patients with survival data available). Patients were stratified according to the four biological subtypes; survival differences were assessed using the log-rank test. g, Kaplan-Meier of patients with GBM from the TCGA cohort profiled by Agilent microarray (n = 302 patients, log-rank test) and h, Patients with GBM from the TCGA cohort profiled by RNA-seq (n = 145 patients, log-rank test) classified according to mesenchymal, proneural and classical subtype. i, Kaplan-Meier of GBM patients as in g and j, patients as in h classified according to mesenchymal, proneural and proliferative subtype.
Extended Data Fig. 7 Analysis of the tumor microenvironment and GBM driver alterations in the biological GBM subtypes.
a, Box plots of GBM subtypes tumor purity scores computed by ABSOLUTE; p-values: Kruskal-Wallis test with Nemenyi post hoc correction for multiple comparison (n = 282 tumors). Box plots span the first to third quartiles and whiskers show the 1.5× interquartile range. b, Correlation analysis of nontumor cell fraction in relationship with GBM cell state fraction (n = 36 tumors; Spearman’s correlation; p = 0.089 GPM versus macrophages; p = 0.067 GPM versus neutrophils; p = 0.017 GPM versus oligodendrocytes; p = 0.092 MTC versus macrophages; p = 0.026 PPR versus oligodendrocytes; *p < 0.10; **p < 0.05). Rows are GBM cell states. Columns are non-tumor cell types. Blue to red scale indicates negative to positive correlation. c, Heatmap of the expression of the top 25 microglia- and macrophage-specific genes in nontumor cells from two GPM and two MTC GBM from single cell dataset 1. Cells are ordered by gene expression fold-change of macrophage- versus microglia-specific genes. The top horizontal track shows in red and green nontumor cells from GBM whose tumor cells have a dominant GPM (S4_D1, n = 67 cells, and S12_D1, n = 246 cells) or MTC state (S1_D1, n = 65 cells, and S5_D1, n = 29 cells), respectively. Representative microglia and macrophages marker genes are indicated. d, Bar plots showing the frequency distribution of GBM driver genes grouped by signaling pathways across GBM subtypes. Asterisks indicate the statistical significance (n = 496 tumors; two-sided Fisher’s exact test).
Extended Data Fig. 8 Characterization of GBM biological states by multi-omics data analysis.
a, Heatmap of the M-values of the 100 probes most differentially methylated between GBM subtypes (n = 59 tumors; two-sided MWW test, p < 0.01 and absolute methylation log2(fold-change) > 0.58). b, Volcano plots of differentially expressed miRNA. Upregulated miRNAs in each GBM subtype are indicated in red [n = 294 tumors; log2(FC) ≥ 0 and p-value < 0.0005, two-sided MWW test]. Vertical and horizontal gray lines demarcate log2(FC) and p-value cutoff, respectively. Representative miRNAs upregulated in each functional subtype are indicated. c, Representative miRNA-gene targets networks significantly upregulated in PPR and d, NEU GBM subtypes [n = 294 tumors; log2(FC) > 0 and p < 0.0005, two-sided MWW test]. miRNA targets whose expression was anti-correlated with miRNA expression are listed (n = 294 tumors; Spearman’s correlation, ρ < 0 and p < 0.05) and biological pathways regulated by miRNA-target activity are indicated. Red nodes indicate miRNA targets of interest for the biology of the specific GBM subtype. p-values of individual genes in a-d are reported in Supplementary Table 14a, c, d, respectively. e-h, Box plots showing the expression of selected proteins or phosphoproteins significantly up-regulated (n = 103 tumors; two-sided MWW test) by RPPA in e, GPM; f, MTC; g, NEU; h, PPR GBM. Box plots span the first to third quartiles and whiskers show the 1.5× interquartile range.
Extended Data Fig. 9 Genomic and metabolic characterization of GBM PDCs.
a, Classification of PDCs by random forest. Upper panel, bar plot showing mean ± s.d. of NES of subtype-specific biological activity in each PDC subgroup. Middle panel, representative biological pathways exhibiting differential activity among subtypes [n = 79 PDCs; logit(NES) > 0.3 and FDR < 0.05, two-sided MWW test]. Bottom panel, representative genes differentially expressed in PDC subtypes (n = 79 PDCs; log2(FC) > 0.3 and FDR < 0.05, two-sided MWW test). Red, green, blue, and cyan indicate significant pathway activation/gene up-regulation in PDCs classified as GPM, MTC, NEU or PPR, respectively; gray, pathway activation/gene up-regulation in any other subtype; white, lack of activation or up-regulation. b, OCR kinetics in 2 MTC PDCs each derived from an independent patient and 2 GPM PDCs each derived from an independent patient shows elevated OCR in MTC PDCs. Data are mean ± s.d. from one representative experiment for each PDC including n ≥9 replicates (see Source Data Extended Data Fig. 9). c, ECAR kinetics in 2 MTC PDCs each derived from an independent patient and 2 GPM PDCs each derived from an independent patient shows elevated glycolysis in GPM PDCs. Data are mean ± s.d. from one representative experiment for each PDC including n ≥7 replicates (see Source Data Extended Data Fig. 9). mpH, milli pH unit. Experiments were repeated two times with similar results. d, Box plots showing the expression of SLC1A5 in GPM (n = 67 tumors) and MTC (n = 108 tumors) primary GBM. e, Box plots showing the expression of SLC1A5 in GPM and MTC PDCs (n = 21 GPM PDCs each derived from an independent patient and n = 25 MTC PDCs each derived from an independent patient); p-value: two-sided MWW test. Box plots span the first to third quartiles and whiskers show the 1.5× interquartile range.
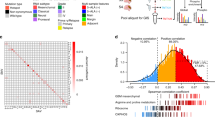
Extended Data Fig. 10 SLC45A1 is the target of chromosome 1p36.23 deletion in MTC GBM.
a, Schematics of chromosome location peak deletions in MTC GBM (n = 153 tumors) identified using GISTIC2 (Benjamini Hochberg FDR q-value < 0.01; q-value of chromosome bands are reported in Supplementary Table 16b). b, The matrix of homozygous deleted genes identified by UNCOVER as associated with MTC NES in primary GBM (n = 487 tumors; p = 0.034, permutation test). Top row, blue to yellow: higher to lower NES values for samples (columns). Deletions in each sample are in dark blue; samples not deleted are in yellow. The last row shows the alteration profile from the entire analysis. The bar plot on the right side indicates the gene weight for each alteration. c, Association of homozygous deletions in each GBM subtype. Circles are color-coded and their dimension reflects the -log10(p-value) of the enrichment (n = 487 tumors; p-value, two-sided Fisher’s exact test; see Supplementary Table 16g). Blue to red scale indicates positive to negative association. d, Frequency of genetic alterations of GBM driver genes in SLC45A1-deleted (n = 20 tumors) compared to SLC45A1 wild-type GBM (n = 705 tumors). The bottom track indicates the dataset (green, TCGA; blue, GLASS). Asterisk, p = 2.33e-03, two-sided Fisher’s Exact test (n = 725 tumors). e, Sample density plot depicting the relative frequency distribution of CCF estimated for the genetic alterations occurring in SLC45A1-deleted GBM (n = 20 tumors). Blue dot, CCF of SLC45A1 deletion. f, Evolutionary trees of genetic alterations in primary and recurrent SLC45A1-deleted GBM (n = 8 matched primary and recurrent tumor pairs); yellow, red and black branches are truncal, primary private and recurrent private alterations, respectively; the length of branches is proportional to the number of genetic alterations. GBM driver genes are indicated. g, PCR amplification of genomic DNA shows deletion of SLC45A1 in PDC-002 and PDC-064. h, Immunoblot of FLAG-SLC45A1 in PDC-002, PDC-064 (harboring SLC45A1 deletion) and PDC-078 (SLC45A1 wild type). Experiments in g, h were repeated two times with similar results. See Source Data Extended Data Fig. 10.
Supplementary information
Supplementary Information
Supplementary Computational Methods.
Source data
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed microphotographs.
Source Data Fig. 6
Unprocessed gels.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed blots and microphotographs.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed gels and blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Garofano, L., Migliozzi, S., Oh, Y.T. et al. Pathway-based classification of glioblastoma uncovers a mitochondrial subtype with therapeutic vulnerabilities. Nat Cancer 2, 141–156 (2021). https://doi.org/10.1038/s43018-020-00159-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00159-4
This article is cited by
-
Integrative multi-omics characterization reveals sex differences in glioblastoma
Biology of Sex Differences (2024)
-
Mammalian target of rapamycin inhibition protects glioma cells from temozolomide-induced cell death
Cell Death Discovery (2024)
-
Imaging cancer metabolism using magnetic resonance
npj Imaging (2024)
-
IDHwt glioblastomas can be stratified by their transcriptional response to standard treatment, with implications for targeted therapy
Genome Biology (2024)
-
Genome-wide CRISPR screens identify novel regulators of wild-type and mutant p53 stability
Molecular Systems Biology (2024)