Abstract
Glioblastomas harbor diverse cell populations, including rare glioblastoma stem cells (GSCs) that drive tumorigenesis. To characterize functional diversity within this population, we performed single-cell RNA sequencing on >69,000 GSCs cultured from the tumors of 26 patients. We observed a high degree of inter- and intra-GSC transcriptional heterogeneity that could not be fully explained by DNA somatic alterations. Instead, we found that GSCs mapped along a transcriptional gradient spanning two cellular states reminiscent of normal neural development and inflammatory wound response. Genome-wide CRISPR–Cas9 dropout screens independently recapitulated this observation, with each state characterized by unique essential genes. Further single-cell RNA sequencing of >56,000 malignant cells from primary tumors found that the majority organize along an orthogonal astrocyte maturation gradient yet retain expression of founder GSC transcriptional programs. We propose that glioblastomas grow out of a fundamental GSC-based neural wound response transcriptional program, which is a promising target for new therapy development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Bulk RNA-seq (EGAS00001003070 and EGAS00001004395), WGS (EGAS00001004395), sc and snRNA-seq (EGAS00001004656) datasets generated and analyzed in this study are available through the European Genome-Phenome Archive repository in the form of FASTQ or BAM files. Processed sc and snRNA-seq data are publicly available through the Broad Institute Single-Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP503) and CReSCENT60 (https://crescent.cloud; study ID CRES-P23). All other data supporting the findings of this study are available from the corresponding author on reasonable request. Original CSV files for Supplementary Tables 2–9 are available in the Supplementary Information. Previously published scRNA-seq data that were re-analyzed in this study are available from the following sources: Wang et al.9 (GSE138794), Bhaduri et al.10 (http://cells.ucsc.edu/?ds=gbm), Neftel et al.2 (https://singlecell.broadinstitute.org/single_cell/study/SCP393/) and Darmanis et al.8 (http://gbmseq.org/). Source data are provided with this paper.
Code availability
Code necessary to reproduce the core analyses presented in this study are available without restrictions at https://github.com/pughlab/su2c-gsc-scrna.
References
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell https://doi.org/10.1016/j.cell.2019.06.024 (2019).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Meyer, M. et al. Single cell-derived clonal analysis of human glioblastoma links functional and genomic heterogeneity. Proc. Natl Acad. Sci. USA 112, 851–856 (2015).
Verhaak, R. G. W. et al. An integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR and NF1. Cancer Cell 17, 98 (2010).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56 (2017).
Carlsson, S. K., Brothers, S. P. & Wahlestedt, C. Emerging treatment strategies for glioblastoma multiforme. EMBO Mol. Med. 6, 1359–1370 (2014).
Darmanis, S. et al. Single-cell RNA-seq analysis of infiltrating neoplastic cells at the migrating front of human glioblastoma. Cell Rep. 21, 1399–1410 (2017).
Wang, L. et al. The phenotypes of proliferating glioblastoma cells reside on a single axis of variation. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-19-0329 (2019).
Bhaduri, A. et al. Outer radial glia-like cancer stem cells contribute to heterogeneity of glioblastoma. Cell Stem Cell 26, 48–63 (2020).
Berezovsky, A. D. et al. Sox2 promotes malignancy in glioblastoma by regulating plasticity and astrocytic differentiation. Neoplasia 16, 193–206 (2014).
Lan, X. et al. Fate mapping of human glioblastoma reveals an invariant stem cell hierarchy. Nature 549, 227–232 (2017).
Natsume, A. et al. Chromatin regulator PRC2 is a key regulator of epigenetic plasticity in glioblastoma. Cancer Res. 73, 4559–4570 (2013).
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Bao, S. et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444, 756–760 (2006).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488, 522–526 (2012).
Liu, G. et al. Analysis of gene expression and chemoresistance of CD133+ cancer stem cells in glioblastoma. Mol. Cancer 5, 67 (2006).
Pollard, S. M. et al. Glioma stem cell lines expanded in adherent culture have tumor-specific phenotypes and are suitable for chemical and genetic screens. Cell Stem Cell 4, 568–580 (2009).
Kelly, J. J. P. et al. Proliferation of human glioblastoma stem cells occurs independently of exogenous mitogens. Stem Cells 27, 1722–1733 (2009).
Florio, M. et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science 347, 1465–1470 (2015).
Zhang, C.-L., Zou, Y., He, W., Gage, F. H. & Evans, R. M. A role for adult TLX-positive neural stem cells in learning and behaviour. Nature 451, 1004–1007 (2008).
Zhu, Z. et al. Targeting self-renewal in high-grade brain tumors leads to loss of brain tumor stem cells and prolonged survival. Cell Stem Cell 15, 185–198 (2014).
Ouafik, L. et al. Neutralization of adrenomedullin inhibits the growth of human glioblastoma cell lines in vitro and suppresses tumor xenograft growth in vivo. Am. J. Pathol. 160, 1279–1292 (2002).
Park, N. I. et al. ASCL1 reorganizes chromatin to direct neuronal fate and suppress tumorigenicity of glioblastoma stem cells. Cell Stem Cell 21, 209–224.e7 (2017).
Körber, V. et al. Evolutionary trajectories of IDHWT glioblastomas reveal a common path of early tumorigenesis instigated years ahead of initial diagnosis. Cancer Cell https://doi.org/10.1016/j.ccell.2019.02.007 (2019).
Lee, J. H. et al. Human glioblastoma arises from subventricular zone cells with low-level driver mutations. Nature 560, 243–247 (2018).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Gojo, J. et al. Single-cell RNA-seq reveals cellular hierarchies and impaired developmental trajectories in pediatric ependymoma. Cancer Cell 38, 44–59 (2020).
Hovestadt, V. et al. Resolving medulloblastoma cellular architecture by single-cell genomics. Nature 572, 74–79 (2019).
Izar, B. et al. A single-cell landscape of high-grade serous ovarian cancer. Nat. Med. https://doi.org/10.1038/s41591-020-0926-0 (2020).
Ledergor, G. et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 24, 1867 (2018).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624 (2017).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330 (2018).
Kinker, G. S. et al. Pan-cancer single-cell RNA-seq identifies recurring programs of cellular heterogeneity. Nat. Genet. 52, 1208–1218 (2020).
Krieger, T. G. et al. Single-cell analysis of patient-derived PDAC organoids reveals cell state heterogeneity and a conserved developmental hierarchy. Preprint at bioRxiv https://doi.org/10.1101/2020.08.23.263160 (2020).
McFarland, J. M. et al. Multiplexed single-cell transcriptional response profiling to define cancer vulnerabilities and therapeutic mechanism of action. Nat. Commun. 11, 4296 (2020).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
John Lin, C.-C. et al. Identification of diverse astrocyte populations and their malignant analogs. Nat. Neurosci. 20, 396–405 (2017).
Chai, H. et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron 95, 531–549 (2017).
Morel, L. et al. Molecular and functional properties of regional astrocytes in the adult brain. J. Neurosci. 37, 8706–8717 (2017).
Miller, S. J. Astrocyte heterogeneity in the adult central nervous system. Front. Cell. Neurosci. 12, 401 (2018).
Haghverdi, L., Buettner, F. & Theis, F. J. Diffusion maps for high-dimensional single-cell analysis of differentiation data. Bioinformatics 31, 2989–2998 (2015).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 Genes Genom. Genet. 7, 2719–2727 (2017).
Hart, T. & Moffat, J. BAGEL: a computational framework for identifying essential genes from pooled library screens. BMC Bioinf. 17, 164 (2016).
MacLeod, G. et al. Genome-wide CRISPR-Cas9 screens expose genetic vulnerabilities and mechanisms of temozolomide sensitivity in glioblastoma stem cells. Cell Rep. 27, 971–986 (2019).
Zhou, Y. et al. Metabolic alterations in highly tumorigenic glioblastoma cells preference for hypoxia and high dependency on glycolysis. J. Biol. Chem. 286, 32843–32853 (2011).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Su, L. et al. H2A.Z.1 crosstalk with H3K56-acetylation controls gliogenesis through the transcription of folate receptor. Nucleic Acids Res. 46, 8817–8831 (2018).
Philip, B. et al. Mutant IDH1 promotes glioma formation in vivo. Cell Rep. 23, 1553–1564 (2018).
Bhat, K. P. L. et al. Mesenchymal differentiation mediated by NF-κB promotes radiation resistance in glioblastoma. Cancer Cell 24, 331–346 (2013).
Xie, Y. et al. The human glioblastoma cell culture resource: validated cell models representing all molecular subtypes. EBioMedicine 2, 1351–1363 (2015).
Sirko, S. et al. Reactive glia in the injured brain acquire stem cell properties in response to sonic hedgehog. Cell Stem Cell 12, 426–439 (2013); erratum 12, 629 (2013).
Robel, S., Berninger, B. & Götz, M. The stem cell potential of glia: lessons from reactive gliosis. Nat. Rev. Neurosci. 12, 88–104 (2011).
Mohanraj, S. et al. CReSCENT: CanceR single cell ExpressioN toolkit. Nucleic Acids Res. 48, W372–W379 (2020).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Alles, J. et al. Cell fixation and preservation for droplet-based single-cell transcriptomics. BMC Biol. 15, 44 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Innes, B. T. & Bader, G. D. scClustViz – single-cell RNAseq cluster assessment and visualization. F1000Research 7, 1522 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0591-3 (2020).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS J. Integr. Biol. 16, 284–287 (2012).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinf. 14, 7 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Acknowledgements
Research was supported by Stand Up To Cancer (SU2C) Canada Cancer Stem Cell Dream Team Research Funding (SU2C-AACR-DT-19-15) provided by the Government of Canada through Genome Canada and the Canadian Institute of Health Research, with supplemental support from the Ontario Institute for Cancer Research, through funding provided by the Government of Ontario. SU2C Canada is a Canadian Registered Charity (reg. no. 80550 6730 RR0001). Research Funding is administered by the American Association for Cancer Research International – Canada, the Scientific Partner of SU2C Canada. L.M.R. was supported by an Ontario Graduate Scholarship and the Frank Fletcher Memorial Fund from the University of Toronto. O.K.N.W. was supported by funding from the Canadian Institute for Health Research, the Cecil Yip Doctoral Research Award and the David Stephen Cant Graduate Scholarship in Stem Cell Research from the University of Toronto. Funding to H.A.L. and S.W. was from Canadian Institute for Health Research. T.J.P. holds the Canada Research Chair in Translational Genomics and is additionally supported by the Princess Margaret Cancer Foundation, a Senior Investigator Award from the Ontario Institute for Cancer Research and the Government of Canada through Genome Canada and the Ontario Genomics Institute (OGI-167). Additional infrastructure support came from the Canada Foundation for Innovation, Leaders Opportunity Fund (CFI no. 32383); Ontario Ministry of Research and Innovation, Ontario Research Fund Small Infrastructure Program; Ontario Institute for Cancer Research; the Chan Zuckerberg Initiative; and the Princess Margaret Cancer Foundation. P.B.D. is additionally supported by the Canadian Institutes for Health Research, the Ontario Institute for Cancer Research, the Terry Fox Research Institute, the Hospital for Sick Children Foundation, the Bresler family, Jessica’s Footprint Foundation, the Hopeful Minds Foundation and B.R.A.I.N. Child. P.B.D. holds a Garron Family Chair in Childhood Cancer Research at The Hospital for Sick Children. G.D.B. was supported by NRNB (US National Institutes of Health, National Center for Research Resources grant no. P41 GM103504). T.J.P., P.B.D. and G.D.B. are supported by a Canadian Cancer Society Impact Grant. We thank R. Hassam, O. Cseh and I. Restall (University of Calgary) for technical assistance and the Calgary Brain Tumor and Tissue Bank for providing patient samples for the establishment of the BT cell line. We thank the staff of the Princess Margaret Genomics Centre (www.pmgenomics.ca), Bioinformatics and High-Performance Computing Core and the BC Cancer Agency Genome Sciences Centre (www.bcgsc.ca) for their expertise in generating the sequencing data used in this study.
Author information
Authors and Affiliations
Contributions
L.M.R., O.K.N.W., P.B.D., G.D.B. and T.J.P. conceived the project, designed the study and interpreted results. N.S., M.R., T.K., Z.X. and L.M.R. generated sc and snRNA-seq data. L.M.R. and O.K.N.W. performed scRNA-seq analysis. F.J.C., F.M.G.C. and P.G. generated and pre-processed bulk RNA-seq or WGS data. O.K.N.W. and F.M.G.C. performed bulk RNA-seq analysis. L.M.R. and F.M.G.C. performed WGS analysis. F.J.C., M.K., N.R., L.L., C.C., H.A.L. and J.E.J. derived GSC cultures used in the study and performed LDAs, xenografts and cytokine assays. G.M., M.A., D.A.B., J.E.J., N.L., E.L., N.I.P., J.K.B. and M.K. performed genome-wide CRISPR–Cas9 screens. G.M. and L.M.R. analyzed screen data. K.Y., J.S., S.D., M.B. and M.D.C. provided tumor tissue. F.J.C., D.C.C., M.K., M.L., B.H.K., H.A.L., S.W., M.A.M., R.A.M. and S.A. provided experimental and analytical support. L.M.R., G.D.B., P.B.D. and T.J.P. wrote the manuscript with feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cancer thanks Aaron Diaz, Benjamin Deneen, Justin Lathia, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Visualization and benchmarking of intra-GSC clustering.
a, t-SNE representation of intra-GSC heterogeneity across 29 patient-derived GSCs. Cells are colored by transcriptional cluster. Samples ordered by number of clusters. b, Comparison of cluster number (top), marker genes per cluster (middle) and average silhouette width per cluster (bottom) between our original GSC smart local moving (SLM) clustering algorithm (blue), Louvain (yellow), Louvain with multilevel refinement (green), k-means (salmon) and spectral (pink). across 29 GSCs. The number of data points in the boxplots (middle, bottom) corresponds to the number of clusters in the matched histogram (top). Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value.
Extended Data Fig. 2 Defining intra-GSC transcriptional heterogeneity.
a, Heat map of Jaccard Index (more similar = blue, less similar = white) between marker gene lists across 86 intra-GSC clusters. A subset of 14 clusters, from 13 samples, display increased similarity (labelled as Cluster 1). b, Enriched pathways from 358 genes common to all 14 clusters defined in Extended Data Fig. 2a. c, Spearman correlation between inferred proportion G2M cells from scRNA-seq data vs. survival in an orthotopic xenograft model (left; n = 18 independent GSC xenograft models) and doubling time in vitro (n = 15 GSC cultures) in adherent (green) or neurosphere (orange) GSCs. Red line represents a linear regression line. Shaded grey area represents 95% confidence interval. d, 14 intra-GSC clusters share increased marker gene overlap and define a core proliferation module shared across 13 patients. Expression of select marker genes common across all clusters. Columns separated by intra-GSC cluster, bolded labels represent clusters with upregulation of the proliferation module. e, Relative expression of top 5 significant marker genes (based on logFC, one-sided Wilcoxon rank-sum test, FDR < 0.05) for clusters C1 and C2 within G549_L (left). UMAP visualization of select marker genes of C2 (right). f, Relative expression of top 5 significant marker genes (based on logFC, one-sided Wilcoxon rank-sum test, FDR < 0.05) for clusters C1-C5 within G837_L (left). UMAP visualization of select marker gene of C5 (right).
Extended Data Fig. 3 Validation of inferred single cell CNV profiles and impact on marker gene expression.
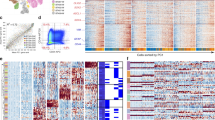
a, Spearman correlation between inferred scRNA-seq CNV score from averaged intra-GSC clusters (left; n = 56 clusters from 20 GSC cultures) or averaged samples (right; n = 20 GSC cultures) and log2 ratios from matched genes from WGS of GSC samples (n = 20 GSC cultures). Each point represents a gene within a given sample. b, Distribution of InferCNV scores for genes labelled as deletion (<0; n = 11,617 genes), neutral (0; n = 100,426 genes) or amplified (>0; n = 12,777 genes across) by GISTIC from corresponding WGS data. Gene counts per GISTIC CNV state represent a cumulative number of genes across 20 GSCs. Median scores for deletions (-0.15) and gains (0.17) used as cut offs to classify InferCNV scores as at least single copy gains or losses. Box plots within the violin plot represent the median, upper and lower quartiles of the distribution and whiskers represent 1.5-times interquartile range. Tips of the violin plot extend to the minimum and maximum values of the distribution. c, Visualization of single cell CNV calls averaged by intra-GSC cluster (denoted “_C#”), averaged by sample (“SampleAverage”) or results of matched WGS (“_WGS”). Samples (rows separated by solid lines) ordered by increasing cluster number. WGS CNV track below dashed line. Sample average above dashed line and cluster transcriptional profiles represent remaining rows. d, Binary heat map depicting chromosome arms (y-axis; sorted by genomic position) that are gained (red), deleted (blue) or copy-neutral (white) across intra-GSC clusters (x-axis; ordered alphabetically; n = 86 clusters from 29 GSC cultures). e, Proportion of cluster marker genes located within a variable CNV loci (y-axis) across intra-GSC clusters (x-axis; n = 69 clusters) from samples with variable cluster CNV profiles (n = 22 GSC cultures) as determined in Extended Data Fig. 3d. Clusters with significant (Fisher’s Exact Test p < 0.05) enrichment of marker genes within variable CNV loci are colored dark blue.
Extended Data Fig. 4 Defining global inter-GSC cluster relationships and evaluation of batch correction methods.
a, UMAP projection of 69,393 GSC cells from 29 patients reveals patient-specific clustering patterns (left panel, cells colored by patient). Unbiased clustering reveals 61 transcriptional clusters (right panel, cells colored by transcriptional cluster). GSCs derived from different regions of the same tumor underlined with red (G945-I,J,K) and black (G946-J,K) bars. b, Transcriptional clusters from the same sample and patient are more similar to each other compared to cells from other samples. Dendrogram of average gene expression profiles of transcriptional clusters defined in Extended Data Fig. 4a based on distance (1-Spearman correlation)(top). Sample composition of transcriptional clusters (bottom). Vertical bars colored by sample. Labels at bottom depict sample identifier and proportion of sample for up to the top three samples/cluster. c, UMAP visualizations of global GSC clustering results with CONOS batch correction (top row), with Liger batch correction (middle row) and fastMNN batch correction (bottom row). Cells are colored by sample ID (left column) and transcriptional cluster (right column) (n = 69,393 cells from 29 GSC cultures). d, Proportion of cells (y-axis) corresponding to a given sample across transcriptional clusters (x-axis) across original and batch corrected datasets. e, Number of transcriptional clusters in original clustering pipeline vs. post-batch correction. f, Box plots representing the number of samples with >10 cells per transcriptional cluster across original and batch corrected clustering results (Original=61 clusters; Conos=12 clusters; Liger=78 clusters; fastMNN=39 clusters). Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value. Outliers displayed as circles.
Extended Data Fig. 5 Characterization and interpretation of GSC transcriptional gradient.
a, PCA plot of 69,393 cells from 29 GSC cultures. Plot colored by cell density (left). PCA plot with cells belonging to outlier sample G800_L, colored red. Remainder of cells colored grey (middle). Quantification of deviation from the mean of PC2 (y-axis) across samples. G800_L (red) represents an outlier with >95% of cells within the sample greater than two standard deviations from the mean. Horizontal dashed red line represents threshold of two standard deviations to determine outliers (right). Box plots represent the median, upper and lower quartiles of the distribution and whiskers represent 1.5-times interquartile range or the most extreme value. Outliers represented as circles. b, Correlation of cell type gene signature scores from PC1 cell embeddings (n = 65,655 cells from 28 GSC cultures; outlier G800_L removed as in Fig. 2a). Only correlations with Spearman correlation coefficient greater than |0.5| shown. Bars colored by gene signature source. c, Enriched MSigDB gene sets (FDR < 0.01) for top 100 and bottom 100 genes for PC1. (n = 65,655 cells from 28 GSC cultures; outlier G800_L removed as in Fig. 2a). d, Gene Set Enrichment Analysis (GSEA) on PC1 loadings (gene associations with PC1) visualized using EnrichmentMap (n = 65,655 cells from 28 GSC cultures; outlier G800_L removed as in Fig. 2a). Similar pathways (circles) are grouped into labeled clusters (larger bubbles). Blue circles denote positively associated pathways (Injury Response associated) and red circles denote negatively associated pathways (Developmental associated). Edges (lines) denote overlap between pathways.
Extended Data Fig. 6 Diffusion Map and bulk RNA-sequencing of 72 GSCs confirms Developmental and Injury Response transcriptional states.
a, Spearman correlation between diffusion component 1 (DM1; x-axis) and principal component 1 (PC1; y-axis) cell embeddings for a subset of 14,000 GSCs (500 cells/sample). b, Diffusion Map of 14,000 GSCs. Cells coloured by PC1 cell embeddings (left; Related to Fig. 2a), scaled Developmental transcriptional program score (middle) and scaled Injury Response transcriptional program score (right). c, Spectral clustering determined GSCs (n = 72 GSC cultures) profiled with bulk RNA-sequencing separated into two stable clusters. For each cluster number (x-axis), boxplots depict 200 pairwise similarities (y-axis) (adjusted Rand index, ARI) between the solution obtained for the full dataset and random subsets of data containing 80% of samples. Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value. Outliers displayed as circles. d, PCA plot of GSCs profiled with bulk RNA-sequencing colored by GSVA score for Developmental signature (n = 72 GSC cultures). Circles denote GSCs from the Developmental cluster, while triangles denote GSCs from the Injury Response Cluster. e, GSEA on differentially expressed genes between Developmental and Injury Response clusters as determined by bulk RNA-sequencing, visualized with EnrichmentMap. Similar pathways (circles) are grouped into labeled clusters (larger bubbles). Blue circles denote Injury Response associated pathways and red circles denote Developmental associated pathways. Edges (lines) denote overlap between pathways. f, Spearman correlation at the individual cell (n = 65,655) level between PC1 cell embeddings from scRNA-seq and Developmental and Injury Response gene signature scores derived from bulk RNA-sequencing.
Extended Data Fig. 7 Continuous transcriptional gradient of Developmental and Injury Response cell states across patients.
a, Distribution of AUC gene signature scores for Developmental (left) and Injury Response (right) programs across all GSC cells (n = 65,655 cells from 28 GSC cultures). Red line marks classification threshold to determine if a given program is active or not. b, Proportion of cells across samples categorized as being resembling Developmental or Injury Response states, as well as intermediate hybrid states. c, Position of cells on the Developmental (x-axis) and Injury Response (y-axis) gradient across all samples (n = 65,655 cells from 28 GSC cultures). Cells are colored by relative expression of the Developmental (red) and Injury Response (black) expression programs. GSC cultures with intermediate scores either contain subpopulations of both subtypes or middling scores for both states. Samples ordered as presented in Fig. 2d. d, Violin plots depicting the distribution of Developmental (red) and Injury Response (black) programs post-fastMNN correction for cells within samples. Samples sorted by increasing median Injury Response program score. e, Pearson correlation of median Developmental (top panel) and Injury Response (bottom panel) between transcriptional program scores derived from the original expression matrix (x-axis) and expression matrix post-fastMNN batch correction (y-axis). Blue line represents linear regression line, shaded grey area represents 95% confidence interval and each dot represents the median raw AUC score per GSC. f, Ridge plots depicting distribution of the difference in Developmental (red) and Injury Response (black) scores (x-axis) across cells within samples (y-axis) (n = 65,655 cells from 28 GSC cultures). Samples ordered as presented in Fig. 2d. Vertical black line represents the median.
Extended Data Fig. 8 Genome-Wide CRISPR-Cas9 screens in GSCs.
a, Box and whisker plots of TKOv3 gRNA library complexity in T0 populations for 70,948 individual gRNAs from a single independent screen per GSC (n = 11 screens in 11 GSC cultures). Box plots represent the median, first and third quartiles of the distribution and whiskers represent 1.5-times the interquartile range. Outliers displayed as circles. b, Precision-recall curves for 11 GSC CRISPR-Cas9 screen produced with BAGEL pipeline and v2 reference for essential/non-essential genes. c, Barplot depicting the number of shared fitness genes across GSC screens. d, Heatmap of quantile normalized gene fitness Bayes factor (qBF) scores for the 1,484 most variable genes across 11 GSC screens. Samples (columns) annotated with GSVA score for Developmental and Injury Response gene signature scores from bulk RNA-sequencing. e, GSEA on differentially essential genes between Developmental and Injury Response GSCs, visualized with EnrichmentMap. Similar pathways (circles) are grouped into labeled clusters (larger bubbles). Blue circles denote pathways more essential in Injury Response GSCs and red circles denote pathways more essential in Developmental GSCs. Edges (lines) denote overlap between pathways.
Extended Data Fig. 9 Characterization of axes of variation in glioblastoma and single nuclei RNA-sequencing of 53,853 nuclei from 10 patient tumors.
a, Spearman correlation of cell type gene signature scores to PC1 and PC2 cell embeddings for combined PCA of GSC and tumor cells (n = 65,655 cells from 28 GSC cultures and 14,207 malignant cells from 7 tumors). Only correlations with Spearman correlation coefficient greater than |0.4| shown. Bars colored by gene signature source. b, Projection of GSCs (top row; n = 65,655 cells) and =patient tumor cells (bottom row; n = 14,207 cells) onto GBM cell state map: astrocyte-like (AC; bottom left quadrant), oligodendrocyte precursor cell-like (OPC; upper left quadrant), neural progenitor cell-like (NPC, upper right quadrant) and mesenchymal-like (MES; bottom right quadrant). Cells are colored by density (left panels) and Developmental - Injury Response gradient program scores (right panels). c, Proportion of cells across samples that map to each of the 4 GBM cell states. d, UMAP visualization of 53,853 nuclei from 10 patient tumors colored by transcriptional cluster (left), patient (middle) and cell type (right). e, Pearson correlation between average transcriptional cluster expression (left). Proportion patient cells per transcriptional cluster (middle), as colored in panel B. Box plots detailing expression of cell type marker genes per cluster (right). Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value. Outliers are removed. f, Proportion of cell types across tumors (as colored in the right panel of Extended Data Fig. 9d). Numbers in brackets represent the total number of nuclei per tumor.
Extended Data Fig. 10 Validation of GSC-state CNVs in patient tumors and identification of GSC-like tumor cells.
a, Genome-wide inferred CNV profiles for 14,207 malignant cells from 7 patient tumors. Columns represent genomic regions, ordered by genome position across all chromosomes. Rows represent CNVs for individual cells, annotated by sample. b, Developmental (left) and Injury Response (right) program scores across quartiles. Numbers underneath quartile labels depict the number of cells harbouring respective Developmental or Injury Response CNVs. Enrichment of CNVs between upper and lower quartiles was determined using a Chi-squared test. Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value. Outliers are displayed as circles. c, Train and test accuracy for logistic regression model, 30 random 80:20 train test splits (left). Distributions of model coefficients corresponding to the 30 trained models (right). Model coefficients are weights by which the logistic regression model describes class likelihood as a function of PC1 and PC2. Box plots represent the median, first and third quartiles of the distribution and whiskers represent either 1.5-times interquartile range or most extreme value. Outliers displayed as circles. d, Proportion of cells in GSCs correctly classified as being GSCs (blue) or misclassified representing tumor-like GSCs (white). Proportion of tumor cells correctly classified as being tumor (black) or misclassified as being GSC-like (grey). e,f, PCA plot of all GSCs and tumor cells as in Fig. 5a. Black line represents contour encompassing 99% of tumor cells. Blue line represents contour encompassing 99% of GSCs. Grey dots represent tumor cells classified as being GSC-like. White dots with blue outline represent GSC cells classified as being tumor-like. g, Differential gene expression analysis between tumor cells and GSC-like tumor cells. Each dot represents a gene (x-axis) ordered by average log2 fold change (y-axis). Red dashed line represents a log2 fold change of double between groups. h, Differential gene expression analysis between GSCs and tumor-like tumor cells. Each dot represents a gene (x-axis) ordered by average log2 fold change (y-axis). Red dashed line represents a log2 fold change of double between groups. i, Expression of mature and young astrocyte gene signatures between tumor cells (black; n = 12,145 cells) and GSC-like tumor cells (grey; n = 2,062 cells). j, Expression of mature and young astrocyte gene signatures between GSCs (blue; n = 64,502 cells) and tumor-like GSCs (white; n = 1,153 cells).
Supplementary information
Supplementary Note 1
Supplementary Methods. Detailed description of analyses performed in this study.
Supplementary Table 1
Sample and patient information of whole genome, bulk RNA-seq, scRNA-seq, CRISPR and functional assays. Clinical characteristics and functional data for GSC lines and patient tumor samples, including age, sex, tumor stage, sphere formation and xenograft survival.
Supplementary Table 2
sc and snRNA-seq metrics. Sequencing metrics containing the number of barcodes associated with cell/nuclei as determined by Dropbead, sequencing depth per library, mitochondrial content, and the median number of genes and UMIs detected per cell/nuclei after quality control.
Supplementary Table 3
Top 50 differentially expressed marker genes for intra-GSC clusters. Differential gene expression as implemented in scClustViz (one-sided Wilcoxon rank-sum test, FDR < 0.05) was used to generate marker genes for transcriptional clusters within each GSC sample. The top 50 marker genes per cluster were selected based on log fold change (logFC).
Supplementary Table 4
Summary of CNV inferred from scRNA-seq data. CNV calls inferred from scRNA-seq data with InferCNV. Averaged at sample level and by intra-sample transcriptional cluster (denoted ‘C#’).
Supplementary Table 5
Summary of CNV data from WGS. Gene-level CNVs represented as log2 ratios (columns with prefix ‘log2’) and discrete copy number states from Sequenza and GISTIC analysis of WGS data (columns with prefix ‘GISTIC’).
Supplementary Table 6
Curated neural cell and GBM gene signatures. Curated list of publicly available brain and GBM cell-type gene signatures. Gene signatures were used to score scRNA-seq data with AUCell. Gene signatures are labeled as ‘FirstAuthor_Journal_PublicationYear_SignatureName’.
Supplementary Table 7
Developmental and Injury Response GSC signatures from bulk RNA-seq. Output of differential gene expression analysis (DESeq2; FDR < 0.05, two-sided Wald test) between the two clusters identified from spectral clustering of GSCs profiled with bulk RNA-seq (n = 72). Marker genes of clusters correspond to Developmental and Injury Response gene signatures.
Supplementary Table 8
Raw BF scores for 11 CRISPR screens. Output of BAGEL pipeline used to calculate BF for each gene representing a confidence score that gene knockout produces a fitness defect.
Supplementary Table 9
Normalized qBFs from CRISPR screens and z score differences between Developmental and Injury Response GSCs. qBF scores to allow for comparison across screens. List of differentially essential genes between Developmental and Injury Response GSCs.
Supplementary Table 10
Metadata for public sc and snRNA-seq datasets. A summary of public datasets used in this study.
Supplementary Table 11
Oligonucleotide sequence information. A summary of sequences for RT–qPCR primers and CRISPR gRNAs.
Supplementary Data 1
Zipped original csv files of Supplementary Tables 2–9, provided at request of author to ensure that gene names are not converted into dates in Excel files.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Richards, L.M., Whitley, O.K.N., MacLeod, G. et al. Gradient of Developmental and Injury Response transcriptional states defines functional vulnerabilities underpinning glioblastoma heterogeneity. Nat Cancer 2, 157–173 (2021). https://doi.org/10.1038/s43018-020-00154-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00154-9
This article is cited by
-
Integration of scRNA-seq data by disentangled representation learning with condition domain adaptation
BMC Bioinformatics (2024)
-
Research progress of N1-methyladenosine RNA modification in cancer
Cell Communication and Signaling (2024)
-
Mesenchymal glioma stem cells trigger vasectasia—distinct neovascularization process stimulated by extracellular vesicles carrying EGFR
Nature Communications (2024)
-
Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming
Nature Cancer (2024)
-
Integrated transcriptomics uncovers an enhanced association between the prion protein gene expression and vesicle dynamics signatures in glioblastomas
BMC Cancer (2024)