Abstract
We report a systematic analysis of the DNA methylation variability in 1,595 samples of normal cell subpopulations and 14 tumor subtypes spanning the entire human B-cell lineage. Differential methylation among tumor entities relates to differences in cellular origin and to de novo epigenetic alterations, which allowed us to build an accurate machine learning-based diagnostic algorithm. We identify extensive individual-specific methylation variability in silenced chromatin associated with the proliferative history of normal and neoplastic B cells. Mitotic activity generally leaves both hyper- and hypomethylation imprints, but some B-cell neoplasms preferentially gain or lose DNA methylation. We construct a DNA-methylation-based mitotic clock, called epiCMIT, whose lapse magnitude represents a strong independent prognostic variable in B-cell tumors and is associated with particular driver genetic alterations. Our findings reveal DNA methylation as a holistic tracer of B-cell tumor developmental history, with implications in differential diagnosis and the prediction of clinical outcome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
DNA methylation and gene expression data that support the findings of this study have been deposited at the European Genome-phenome Archive (EGA) under accession number EGAS00001004640. Previously published DNA methylation data that were reanalyzed in this study can be found under the following accession codes: B cells, EGAS00001001196; ALL, GSE16368, GSE47051, GSE7658515 and GSE6922916; MCL, EGAS00001001637 and EGAS00001004165; CLL, EGAD00010000871 and EGAD00010000948; MM, EGAS00001000841; in vitro B-cell differentiation model of NBCs from human primary samples, GSE72498. Normalized DNA methylation matrices used for the analyses in this study are available at http://resources.idibaps.org/paper/the-proliferative-history-shapes-the-DNA-methylome-of-B-cell-tumors-and-predicts-clinical-outcome. Published gene expression datasets can be found under the following accession codes: B cells, EGAS00001001197; ALL, GSE47051; MCL, GSE36000; CLL, EGAS00000000092 and EGAD00010000254; MM, GSE19784; in vitro B-cell differentiation model of NBCs from human primary samples, GSE72498. ChIP–seq datasets that were reanalyzed in this study can be found under the following accession codes: GSE109377 (NALM6 ALL cell line, n = 1) and EGAS00001000326 (15 normal B cell donors and 5 individuals with MCL, 7 individuals with CLL and 4 individuals with MM) available from Blueprint (https://www.blueprint-epigenome.eu/). All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
The source code for the DNA methylation classifier of B-cell tumor entities and subtypes and for the calculation of the epiCMIT mitotic clock can be found at https://github.com/Duran-FerrerM/Pan-B-cell-methylome. All other source code supporting the findings of this study is available from the corresponding author on reasonable request.
References
Roy, N. & Hebrok, M. Regulation of cellular identity in cancer. Dev. Cell 35, 674–684 (2015).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304 (2018).
Swerdlow, S. H. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues 4th edn, Vol. 2 (International Agency for Research on Cancer (IARC), 2017).
Luo, C., Hajkova, P. & Ecker, J. R. Dynamic DNA methylation: in the right place at the right time. Science 361, 1336–1340 (2018).
Kulis, M. et al. Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat. Genet. 47, 746–756 (2015).
Nordlund, J. et al. Genome-wide signatures of differential DNA methylation in pediatric acute lymphoblastic leukemia. Genome Biol. 14, r105 (2013).
Lee, S.-T. et al. Epigenetic remodeling in B-cell acute lymphoblastic leukemia occurs in two tracks and employs embryonic stem cell-like signatures. Nucleic Acids Res. 43, 2590–2602 (2015).
Queirós, A. C. et al. Decoding the DNA methylome of mantle cell lymphoma in the light of the entire B cell lineage. Cancer Cell 30, 806–821 (2016).
Nadeu, F. et al. Genomic and epigenomic insights into the origin, pathogenesis, and clinical behavior of mantle cell lymphoma subtypes. Blood 136, 1419–1432 (2020).
Kulis, M. et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat. Genet. 44, 1236–1242 (2012).
Oakes, C. C. et al. DNA methylation dynamics during B cell maturation underlie a continuum of disease phenotypes in chronic lymphocytic leukemia. Nat. Genet. 48, 253–264 (2016).
Shaknovich, R. et al. DNA methylation signatures define molecular subtypes of diffuse large B-cell lymphoma. Blood 116, e81–e89 (2010).
Agirre, X. et al. Whole-epigenome analysis in multiple myeloma reveals DNA hypermethylation of B cell-specific enhancers. Genome Res. 25, 478–487 (2015).
Kaiser, M. F. et al. Global methylation analysis identifies prognostically important epigenetically inactivated tumor suppressor genes in multiple myeloma. Blood 122, 219–226 (2013).
Oakes, C. C. & Martin-Subero, J. I. Insight into origins, mechanisms, and utility of DNA methylation in B cell malignancies. Blood 132, 999–1006 (2018).
Ziller, M. J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013).
Puente, X. S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Karube, K. et al. Integrating genomic alterations in diffuse large B-cell lymphoma identifies new relevant pathways and potential therapeutic targets. Leukemia 32, 675–684 (2018).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Somasundaram, R., Prasad, M. A. J., Ungerbäck, J. & Sigvardsson, M. Transcription factor networks in B-cell differentiation link development to acute lymphoid leukemia. Blood 126, 144–152 (2015).
Sánchez-Tilló, E. et al. The EMT activator ZEB1 promotes tumor growth and determines differential response to chemotherapy in mantle cell lymphoma. Cell Death Differ. 21, 247–257 (2014).
Wolf, C. et al. NFATC1 activation by DNA hypomethylation in chronic lymphocytic leukemia correlates with clinical staging and can be inhibited by ibrutinib. Int. J. Cancer 142, 322–333 (2018).
Blonska, M. et al. Jun-regulated genes promote interaction of diffuse large B-cell lymphoma with the microenvironment. Blood 125, 981–991 (2015).
Huerta-Yepez, S. et al. Overexpression of Yin Yang 1 in bone marrow-derived human multiple myeloma and its clinical significance. Int. J. Oncol. 45, 1184–1192 (2014).
Sprynski, A. C. et al. Insulin is a potent myeloma cell growth factor through insulin/IGF-1 hybrid receptor activation. Leukemia 24, 1940–1950 (2010).
Riz, I. & Hawley, R. G. Increased expression of the tight junction protein TJP1/ZO-1 is associated with upregulation of TAZ–TEAD activity and an adult tissue stem cell signature in carfilzomib-resistant multiple myeloma cells and high-risk multiple myeloma patients. Oncoscience 4, 79–94 (2017).
Herath, N. I., Rocques, N., Garancher, A., Eychène, A. & Pouponnot, C. GSK3-mediated MAF phosphorylation in multiple myeloma as a potential therapeutic target. Blood Cancer J. 4, e175 (2014).
Navarro, A. et al. Improved classification of leukemic B-cell lymphoproliferative disorders using a transcriptional and genetic classifier. Haematologica 102, 360–363 (2017).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Chapuy, B. et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 24, 679–690 (2018).
Schmitz, R. et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N. Engl. J. Med. 378, 1396–1407 (2018).
Aran, D., Toperoff, G., Rosenberg, M. & Hellman, A. Replication timing-related and gene body-specific methylation of active human genes. Hum. Mol. Genet. 20, 670–680 (2011).
Beerman, I. et al. Proliferation-dependent alterations of the DNA methylation landscape underlie hematopoietic stem cell aging. Cell Stem Cell 12, 413–425 (2013).
Landan, G. et al. Epigenetic polymorphism and the stochastic formation of differentially methylated regions in normal and cancerous tissues. Nat. Genet. 44, 1207–1214 (2012).
Siegmund, K. D., Marjoram, P., Woo, Y.-J., Tavaré, S. & Shibata, D. Inferring clonal expansion and cancer stem cell dynamics from DNA methylation patterns in colorectal cancers. Proc. Natl Acad. Sci. USA 106, 4828–4833 (2009).
Spencer, D. H. et al. CpG island hypermethylation mediated by DNMT3A is a consequence of AML progression. Cell 168, 801–816 (2017).
Yang, Z. et al. Correlation of an epigenetic mitotic clock with cancer risk. Genome Biol. 17, 205 (2016).
Zhou, W. et al. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat. Genet. 50, 591–602 (2018).
Youn, A. & Wang, S. The MiAge Calculator: a DNA methylation-based mitotic age calculator of human tissue types. Epigenetics 13, 192–206 (2018).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat. Genet. 44, 40–46 (2011).
Vandiver, A. R., Idrizi, A., Rizzardi, L., Feinberg, A. P. & Hansen, K. D. DNA methylation is stable during replication and cell cycle arrest. Sci. Rep. 5, 1–8 (2015).
Caron, G. et al. Cell-cycle-dependent reconfiguration of the DNA methylome during terminal differentiation of human B cells into plasma cells. Cell Rep. 13, 1059–1071 (2015).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Issa, J. CpG island methylator phenotype in cancer. Nat. Rev. Cancer 4, 988–993 (2004).
Rakyan, V. K. et al. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res. 20, 434–439 (2010).
Teschendorff, A. E. et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 20, 440–446 (2010).
Bell, C. G. et al. DNA methylation aging clocks: challenges and recommendations. Genome Biol. 20, 249 (2019).
Field, A. E. et al. DNA methylation clocks in aging: categories, causes, and consequences. Mol. Cell 71, 882–895 (2018).
Horvath, S. & Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 19, 371–384 (2018).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Queirós, A.C. et al. A B-cell epigenetic signature defines three biological subgroups of chronic lymphocytic leukemia with clinical impact. Leukemia 29, 598–605 (2015).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Shuai, S. et al. The U1 spliceosomal RNA is recurrently mutated in multiple cancers. Nature 574, 712–716 (2019).
Rodríguez-Paredes, M. et al. Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin. Nat. Commun. 9, 577 (2018).
Gaiti, F. et al. Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia. Nature 569, 576–580 (2019).
Meir, Z., Mukamel, Z., Chomsky, E., Lifshitz, A. & Tanay, A. Single-cell analysis of clonal maintenance of transcriptional and epigenetic states in cancer cells. Nat. Genet. 52, 709–718 (2020).
Borssén, M. et al. DNA methylation holds prognostic information in relapsed precursor B-cell acute lymphoblastic leukemia. Clin. Epigenetics 10, 31 (2018).
Sandoval, J. et al. Genome-wide DNA methylation profiling predicts relapse in childhood B-cell acute lymphoblastic leukaemia. Br. J. Haematol. 160, 406–409 (2013).
Rhein, P. et al. Gene expression shift towards normal B cells, decreased proliferative capacity and distinct surface receptors characterize leukemic blasts persisting during induction therapy in childhood acute lymphoblastic leukemia. Leukemia 21, 897–905 (2007).
Oakes, C. C. et al. Evolution of DNA methylation is linked to genetic aberrations in chronic lymphocytic leukemia. Cancer Discov. 4, 348–361 (2014).
Reinius, L. E. et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE 7, e41361 (2012).
Vento-Tormo, R. et al. IL-4 orchestrates STAT6-mediated DNA demethylation leading to dendritic cell differentiation. Genome Biol. 17, 4 (2016).
Brönneke, S. et al. DNA methylation regulates lineage-specifying genes in primary lymphatic and blood endothelial cells. Angiogenesis 15, 317–329 (2012).
Aryee, M. J. et al. Minfi: a flexible and comprehensive bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Maksimovic, J., Gordon, L. & Oshlack, A. SWAN: subset–quantile within array normalization for Illumina Infinium HumanMethylation450 BeadChips. Genome Biol. 13, R44 (2012).
Bergmann, A. K. et al. DNA methylation profiling of pediatric B-cell lymphoblastic leukemia with KMT2A rearrangement identifies hypomethylation at enhancer sites. Pediatr. Blood Cancer 64, 1–5 (2017).
Gabriel, A. S. et al. Epigenetic landscape correlates with genetic subtype but does not predict outcome in childhood acute lymphoblastic leukemia. Epigenetics 10, 717–726 (2015).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 13, 86 (2012).
Scott, D. W. & Gascoyne, R. D. The tumour microenvironment in B cell lymphomas. Nat. Rev. Cancer 14, 517–534 (2014).
Teschendorff, A. E. & Relton, C. L. Statistical and integrative system-level analysis of DNA methylation data. Nat. Rev. Genet. 19, 129–147 (2018).
Navarro, A. et al. Molecular subsets of mantle cell lymphoma defined by the IGHV mutational status and SOX11 expression have distinct biologic and clinical features. Cancer Res. 72, 5307–5316 (2012).
Broyl, A. et al. Gene expression profiling for molecular classification of multiple myeloma in newly diagnosed patients. Blood 116, 2543–2553 (2010).
Stunnenberg, H. G., International Human Epigenome Consortium & Hirst, M. The International Human Epigenome Consortium: a blueprint for scientific collaboration and discovery. Cell 167, 1145–1149 (2016).
Debaize, L. et al. Interplay between transcription regulators RUNX1 and FUBP1 activates an enhancer of the oncogene c-KIT and amplifies cell proliferation. Nucleic Acids Res. 46, 11214–11228 (2018).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Beekman, R. et al. The reference epigenome and regulatory chromatin landscape of chronic lymphocytic leukemia. Nat. Med. 24, 868–880 (2018).
Guyon, I., Weston, J., Barnhill, S. & Vapnik, V. Gene selection for cancer classification using Support Vector Machines. Machine Learning 46, 389–422 (2002).
Dietrich, S. et al. Drug-perturbation-based stratification of blood cancer. J. Clin. Invest. 128, 427–445 (2017).
Sánchez-Vega, F., Gotea, V., Margolin, G. & Elnitski, L. Pan-cancer stratification of solid human epithelial tumors and cancer cell lines reveals commonalities and tissue-specific features of the CpG island methylator phenotype. Epigenetics Chromatin 8, 1–24 (2015).
Maura, F. et al. A practical guide for mutational signature analysis in hematological malignancies. Nat. Commun. 10, 2969 (2019).
Acknowledgements
This research was funded by the European Union’s Seventh Framework Programme through the Blueprint Consortium (grant agreement 282510); the European Research Council under the European Union’s Horizon 2020 research and innovation program (Project BCLLATLAS, grant agreement 810287); Generalitat de Catalunya Suport Grups de Recerca AGAUR 2017-SGR-1142 (to E.C.) and 2017-SGR-736 (to J.I.M.-S.); Ministerio de Ciencia, Innovación y Universidades of the Spanish Government, grants RTI2018-094274-B-I00 (to E.C.) and SAF2017-86126-R (to J.I.M.-S.); Proyecto Medicina Personalizada PERMED (grant PMP15/00007), which is part of Plan Nacional de I+D+I and is cofinanced by the ISCIII-Sub-Directorate General for Evaluation and the European Regional Development Fund (FEDER; “Una manera de Hacer Europa”); CIBERONC (CB16/12/00225, CB16/12/00334, CB16/12/00236 and CB16/12/00489); the Accelerator award CRUK/AIRC/AECC joint funder-partnership; research funding from Fondo de Investigaciones Sanitarias, Instituto de Salud Carlos III PI17/01061 (S.B.); Ministerio de Ciencia, Innovación y Universidades, RTI2018-094274-B-I00, SAF2015-64885-R (E.C.); NIH grant number 1P01CA229100 (E.C.) and the European Regional Development Fund “Una manera de fer Europa”, CERCA Programme/Generalitat de Catalunya. F.N. is supported by a predoctoral fellowship of the Ministerio de Economía y Competitividad (MINECO, BES-2016-076372). E.C. is an Academia Researcher of the Institució Catalana de Recerca i Estudis Avançats (ICREA) of the Generalitat de Catalunya. This work was partially developed at the Centro Esther Koplowitz (CEK, Barcelona, Spain). We thank F. Maura for his help with the analysis of mutational signatures.
Author information
Authors and Affiliations
Contributions
M.D.-F., G. Clot, F.N. and R.B. performed DNA methylome, ChIP–seq, transcriptome, genetic, clinical and/or statistical analyses. T.B., J.N., Y.M.-Z., G.L., A.R.-D., S.M., R.O., G. Castellano, M.K., A.C.Q., S.-T.L., J.W., J.L., E.G., S.B., P.J., X.A., F.P., C.L.-O., X.S.P., C.C.O., T.Z., J.D., A.L.-G. and E.C. contributed to sample biological and/or clinical annotation. R.R. and M.P. provided computational support. M.D.-F. and J.I.M.-S. participated in study design. M.D.-F., F.N., R.B., T.B., J.D., A.L.-G., E.C. and J.I.M.-S. participated in data interpretation. J.I.M.-S. directed the research and wrote the manuscript together with M.D.-F.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analyses related to sample selection and annotation of stably-methylated CpGs.
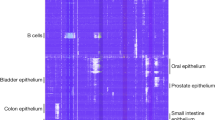
a, Principal component analysis and hierarchical clustering of synchronic unpurified/purified DNA methylation profiles obtained with EPIC array from MCL and CLL patients. Colors represent the same sample, with FCM-based purities highlighted in each sample. MCL, mantle cell lymphoma. CLL, chronic lymphocytic leukemia. b, Correlations and Passing Bablock regression fits of gold-standard methods for tumor purity prediction (FCM and genetic-based) against DNA methylation-based tumor purity prediction for MCL and CLL patients in initial and validation series. Samples sizes are: MCL initial series, n=32; MCL validation series, n=56; CLL cohort 1, n=109 and CLL cohort 2, n=178 patients. Shaded area represents 95% confidence intervals. Pearson correlation and derived p-values are also shown. c, Pearson correlations and Passing Bablock regression fits for gold-standard methods for tumor purity predictions (FCM and genetic-based) against DNA methylation-based tumor purity predictions in MM and DLBCL patients. Sample sizes are: MM, n=100 and DLBCL, n=55 patients and are the same as in panel d. Shaded area represents 95% confidence intervals. Pearson correlation and derived p-values are also shown. d, Pan-B cell DNA methylation signature used to deconvolute DNA methylation data and obtain B-cell tumor purities in B-cell tumors. The DNA methylation levels for the Pan-B-cell DNA methylation signature is shown for microenvironmental cells as well as MM and DLBCL. Bar plots representing DNA-methylation based predictions as well as gold standard-based predictions for MM and DLBCL are represented on the top of the heatmaps. e, Chromatin state genome segmentation with the CHMM software using the 6 histone marks used in the whole study for normal B cells, MCL, CLL and MM primary cases as well as for KARPAS-422 and SUDHL-5 DLBCL cells lines. f, Genomic distribution of stably methylated and unmethylated CpGs in normal and neoplastic B cell. Barplots represent single data values. g, Example gene showing stably unmethylated CpGs at promoters and stably methylated CpGs at gene body in normal and neoplastic B cells. A total of 98 CpGs are shown. h, Gene ontology analysis of genes showing both stably methylated and stably unmethylated CpGs in normal and neoplastic B cells.
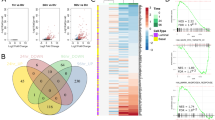
Extended Data Fig. 2 Characterization of tumor-specific DNA methylation signatures.
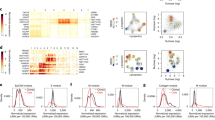
a, First 9 components of a Principal Component Analysis for normal and neoplastic B cells. Samples sizes are the same as in Fig. 1a. The same sample size applies also for panel b, c and d. b, Percentages of de novo DNA methylation signatures over the total DNA methylome. All de novo hyper- and hypomethylation from the five B-cell tumors analyzed are considered together to derive each respective percentage. c, Heatmap showing B-cell tumor-specific hypermethylation and the number of CpGs located at active regulatory regions (marked by H3K27ac). To calculate CpG enrichments in regulatory regions, the number of CpGs falling in regulatory regions were compared with the same number of de novo CpGs 10,000 times randomly chosen from the DNA methylome fraction with potential tumor-specific signatures falling in regulatory regions. d, Distribution of mean methylation levels of CpGs from de novo B-cell tumor-specific DNA methylation signatures across all normal and neoplastic B cell samples subtypes. The number of samples used to calculate the means is shown in Fig. 1a and the number of CpGs analyzed are those from Fig. 2b. e, Genomic distribution for de novo DNA methylation changes in B-cell tumors. Barplots represent single data values. f, Gene expression percentile of TFs showing the most significant p-values and frequencies for TFs binding site predictions (Methods) in de novo hypomethylation signatures in each B-cell tumor from Fig. 2d. Sample sizes for gene expression analyses in tumor samples are the same than in Fig. 4e. Center line, box limits, whiskers and points represent the median, 25th and 75th percentiles, 1.5x interquartile range and individual samples, respectively.
Extended Data Fig. 3 DNA methylation levels and analysis of the sensitivity of the epigenetic classifier of B cell neoplasms.
a, DNA methylation levels of all CpGs from the pan-B-cell diagnostic algorithm in normal and neoplastic B cells. Sample sizes are the training samples shown in Fig. 3b. b, Estimated sensitivity according to the number of CpGs used in the pan-B-cell diagnostic algorithm for the classification of an unknown B-cell tumor into ALL, MCL, CLL, DLBCL or MM (first step of Fig. 3a, predictor 1). The number of CpGs selected for the predictor was chosen by maximizing the highest balanced accuracy and is indicated with a red circle. This strategy was applied also in the remaining 4 predictors to classify B-cell tumor subtypes in panels c–f, (second step of Fig. 3a). Each B-cell tumor is represented with different shapes and colors. c, Estimated sensitivity according to the number of CpGs used in the pan-B-cell diagnostic algorithm (predictor 2 of Fig. 3a) for the classification of ALL into the subtypes HeH, 11q23/MLL, t(12;21), t(1;19), t(9;22) and dic(9;20) while incrementing the number of CpGs (predictor 2 in Fig. 3a). d, Estimated sensitivity according to the number of CpGs used in the pan-B-cell diagnostic algorithm (predictor 3 of Fig. 3a) for the classification of MCL into the subtypes C1 or C2 while incrementing the number of CpGs (predictor 3 in Fig. 3a). e, Estimated sensitivity according to the number of CpGs used in the pan-B-cell diagnostic algorithm for the classification of CLL into the subtypes n-CLL, i-CLL or m-CLL while incrementing the number of CpGs (predictor 4 in Fig. 3a). f, Estimated sensitivity according to the number of CpGs used in the pan-B-cell diagnostic algorithm for the classification of DLBCL into the subtypes ABC and GCB while incrementing the number of CpGs (predictor 5 in Fig. 3a).
Extended Data Fig. 4 Further characterization of patient-specific DNA methylation changes.
a, Variability of DNA methylation changes measured by the interquartile range (IQR) in normal and neoplastic B cells against the median number of DNA methylation changes per each subtype. R and p-values were derived from linear modelling. Shaded area represents 95% confidence interval. b, Correlations in all B cell tumors between B-cell independent DNA methylation changes and B-cell related changes for hypermethylation (top) and hypomethylation (bottom) changes. R and p-values were derived from linear models. c, Number of B-cell related or B-cell independent hyper- or hypomethylation in B-cell tumors showing consistent patterns (Methods). d, B-cell independent CpGs losing DNA methylation in B-cell tumors and the percentages of each chromatin state in normal and neoplastic B-cells. The mean of percentages per sample type is shown. The sample sizes are the same as in Fig. 4c and also apply for panel g. e, The mean of 2,000 representative CpGs per each sample subtype from panel d is represented. f, Gene density distributed along the expression percentiles of genes associated with B-cell independent CpGs losing DNA methylation at low signal heterochromatin in B-cell tumors. Expressed genes (H3K36me3) are displayed at right as control. Means within each B-cell subpopulation as well as B-cell tumors are represented. g, B-cell independent CpGs gaining DNA methylation in B-cell tumors and the percentages in each chromatin state in normal and neoplastic B-cells. h, The mean of 2,000 representative CpGs per each sample subtype from panel g is represented. i, Gene density distributed along the expression percentiles of genes associated with B-cell independent CpGs gaining DNA methylation at H3K27me3 regions in B-cell tumors. Expressed genes (H3K36me3) are displayed at right as control. Means within each B-cell subpopulation as well as B-cell tumors are represented. Sample size for DNA methylation analyzes in panels a, b, c, e and h are the same as in Fig. 4a. Samples sizes for gene expression analyses in panels f and i are the same as in Fig. 4e.
Extended Data Fig. 5 Additional analyses performed to validate the epiCMIT.
a, Illustrative scheme showing DNA methylation changes upon cell division and how they relate to epiCMIT scores. b, In vitro B-cell differentiation model used to experimentally validate the epiCMIT score. Primary naïve B cells are differentiated into plasma cells in 6 days. At day 0, primary human B cells are incubated with carboxyfluorescein succinimidyl ester (CFSE) and harvested with activation and proliferation cocktails necessary for plasma cell differentiation. The epiCMIT was calculated at day 0, day 4 and day 6 in B cells with different proliferative histories based on CFSE dilution. c, The epiCMIT is correlated with total number of mutations detected by WGS in each CLL epigenetic subtype. R and p-values are derived from linear modelling. 138 CLL patient samples with WGS and DNA methylation data are shown (66 n-CLL, 18 i-CLL and 54 m-CLL). The same sample size applies for panel e, f and g. d, The epiCMIT is correlated with CLL genomic complexity measured by the total number of driver alterations and thus with mutations with positive selection. Fitted linear regression models and derived R and p-values are shown for each group. The sample size for each number of driver alterations are: 0 drivers: n-CLL, n=2, i-CLL, n=5, m-CLL, n=44; 1 driver: n-CLL, n=14, i-CLL, n=19, m-CLL, n=119; 2 drivers: n-CLL, n=37, i-CLL, n= 25, m-CLL, n= 55; 3 drivers: n-CLL, n=38, i-CLL, n= 12, m-CLL, n=28; 4 drivers: n-CLL, n=27, i-CLL, n=4, m-CLL, n=12; 5 drivers: n-CLL, n=23, i-CLL, n=2, m-CLL, n=2; 6 drivers: n-CLL, n=10, i-CLL, n=0, m-CLL, n=0; 7 drivers: n-CLL, n=7, i-CLL, n=2, m-CLL, n=0; 8 drivers: n-CLL, n=1; 9 drivers: n-CLL, n=1; 10 drivers: n-CLL, n=1. e, Mutational signatures found in CLL with available WGS. CLL subtypes are shown separately. f, The epiCMIT is correlated with the mitotic-like mutational signature SBS1. CLL samples are divided in CLL epigenetic subgroups. R and p-values are derived from linear models. g, The epiCMIT is correlated with the mitotic-like mutational signatures SBS9. CLL samples are separated with the classical IGHV mutational status (98%). R and p-values are shown for each respective linear model. h, epiCMIT-hyper CpGs and epiCMIT-hypo mitotic clocks are compared with other hyper- or hypomethylation based mitotic clocks as well as the total number of hyper- (rightmost top) or hypomethylation (rightmost bottom) changes per sample since HPC stage. R from linear models are shown. Samples sizes are the same as in Fig. 4a. i, Overlap among the CpG used to build each mitotic clock. Barplots represent single data values. j, Performance of all mitotic clocks in the in vitro B-cell differentiation model from panel c. The fraction of epiCMIT which gain methylation (epiCMIT-hyper) and the fraction that lose DNA methylation (epiCMIT-hypo) were analyzed together with hyper- and hypomethylation-based mitotic clocks, respectively. Biological independent sample sizes are the same as in Fig. 5e. P-values are derived from two-sided t-tests and from biological independent experiments. On the right, expression of genes containing any CpG of each respective mitotic clock as well as genes containing CpGs in H3K36me3 regions are depicted (n=14,598). The number of genes analyzed per each mitotic clock are: epiCMIT-hyper, n=155; epiTOC, n=412; MiAge, n=298; CIMP, n=102; epiCMIT-hypo, n1,123; PMDsoloWCGW, n=4053. For the box plot, center line, box limits, whiskers and points represent the median, 25th and 75th percentiles, 1.5x interquartile range and individual samples, respectively.
Extended Data Fig. 6 Comparison between the epiCMIT mitotic clock and the Horvath aging clock.
a, Correlations among epiCMIT, age and Horvath-predicted age in normal and neoplastic B cells. Samples sizes are: NBC, n=10 and MBC, n=9 donors; C1 MCL, n=40; C2 MCL, n=17; n-CLL, n=159; i-CLL, n=69; m-CLL, n=260; GCB DLBCL, n=20 and ABC DLBCL, n=28 patients. R and p-value are derived from linear models. Shaded areas represent 95% confidence intervals. b, epiCMIT and Horvath clocks do not have any CpG in common. CpGs of the Horvath model are divided into positively associated with age (gain of methylation) and negatively associated with age (loss of methylation). In addition, they are further classified into B-cell related or B-cell independent if they are extensively modulated or not during normal B-cell differentiation. Barplots represent single data values. c, The CpGs used to build the epiCMIT and Horvath clock show distinct genomic locations. Barplots represent single data values. d, DNA methylation levels of the CpGs from the epiCMIT and Horvath clocks in normal and neoplastic B cells. Sample sizes are the same as in Fig. 4a. e, The CpGs associated with the epiCMIT and Horvath clocks are located in markedly different chromatin states. Sample sizes are the same as in Fig. 4c. f, Genes associated with epiCMIT and Horvath CpGs show distinct transcriptional states in normal and neoplastic B cells. Gene probes shared across all normalized matrices from normal and neoplastic B cells were retained and were the following: epiCMIT-hyper, n=60; epiCMIT-hypo, n=327; Age positive B-cell related, n=44; Age positive B-cell independent, n=118; Age negative B-cell related, n=49; Age negative B-cell independent, n=101. For the box plot, center line, box limits, whiskers and points represent the median, 25th and 75th percentiles, 1.5x interquartile range and individual samples, respectively. Sample size are the same as in Fig. 4e.
Extended Data Fig. 7 Additional characterization of the clinical impact of the epiCMIT in B cell tumors.
a, Kaplan-Meier curves for relapse-free survival in ALL patients with low or high epiCMIT according to the maxstat rank statistics-based cutoff. Hazard ratio and p-value for the univariate Cox regression model are shown. A multivariate Cox regression model with epiCMIT as continuous variable and ALL cytogenetic groups is shown on the right. b, epiCMIT preserves its prognostic value in multivariate Cox regressions for time to first treatment in CLL patients whose samples were acquired at maximum 30 months after diagnosis both in initial and validation series. c, epiCMIT shows independent prognostic value from major prognostic variables in CLL including IGHV mutational status and TP53 alterations (deletions and mutations) in multivariate Cox regressions for time to first treatment (TTT). d, Multivariate cox regression models in initial and validation CLL series for overall survival with epiCMIT and important prognostic variables. e, Kaplan-Meier curves for overall survival in GCB and ABC DLBCL patients with low or high epiCMIT according to the maxstat rank statistics-based cutoff. A multivariate Cox regression model with epiCMIT as continuous variable, the DLBCL subtype and age is shown on the right. On the right, univariate cox regression model for all mitotic clocks. All hazard ratios for epiCMIT correspond to 0.1 increments.
Extended Data Fig. 8 Clinical impact of the epiCMIT as compared to other mitotic clocks.
a, On the left, epiCMIT and hypermethylation-based mitotic clocks are highly correlated in ALL, creating a collinearity phenomenon in multivariate cox regression models with multiple mitotic clocks. On the right, multivariate Cox regression models with epiCMIT and PMDsoloWCGW mitotic clocks and ALL cytogenetic subgroups for overall survival, relapse-free survival and overall survival after relapse. b, In CLL, epiCMIT shows superior prognostic value in multivariate cox models for time to first treatment than all the other mitotic clocks in both initial and validation series. c, In MCL, epiCMIT shows an overall superior prognostic value in multivariate Cox models for overall survival in both initial series (with C1 and C2 MCL subtypes) and in the validation series, which only contain C1 MCL subtypes. In the initial series, MCL subtypes with different cellular origin were not introduced in multivariate Cox regression models due to few events, and thus the epiCMIT of each MCL patient was centered according to its cellular origin (C1 or C2) to account for normal B-cell development epiCMIT (Fig. 6a). Hazard ratios for the mitotic clocks correspond to scaled values and are comparable among them in each corresponding Cox model.
Extended Data Fig. 9 Additional data regarding the link between the epiCMIT and genetic changes in CLL.
a, Oncoprint showing all genetic driver alterations considered in the whole CLL initial series composed by 490 CLL patient samples grouped by epigenetic subtypes and ordered according to increasing levels of epiCMIT (from left to right within each epigenetic subgroup). Other clinico-biological features including MBL or CLL, IGHV status, Age, Binet stage, epiCMIT subgroups based on maxstat rank statistic, need for treatment and patient status are shown. Distinct genetic driver alterations are depicted with different colors and shapes. The percentage of mutated patients and number of mutated patients for each alteration is shown at right. b, Driver genetic alterations without clear associations with epiCMIT. Analyses were done in the whole cohort as well as within each epigenetic subgroup. Point estimates with 95% confidence intervals were derived in the whole cohort using linear modelling between epiCMIT and alterations adjusted for CLL subtypes, and with two-sided t-tests within CLL subtypes. Point estimates then represent the coefficient of each respective alteration in each corresponding linear model (whole cohort analysis) or the difference between means (CLL subtypes analysis). Point estimates are color-coded according to FDR correction. Treated and untreated patients at the moment of sampling were considered for these analyses.
Supplementary information
Rights and permissions
About this article
Cite this article
Duran-Ferrer, M., Clot, G., Nadeu, F. et al. The proliferative history shapes the DNA methylome of B-cell tumors and predicts clinical outcome. Nat Cancer 1, 1066–1081 (2020). https://doi.org/10.1038/s43018-020-00131-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00131-2
This article is cited by
-
Genome-wide DNA methylation-analysis of blastic plasmacytoid dendritic cell neoplasm identifies distinct molecular features
Leukemia (2024)
-
Refining risk prediction in pediatric acute lymphoblastic leukemia through DNA methylation profiling
Clinical Epigenetics (2024)
-
Comprehensive analyses of partially methylated domains and differentially methylated regions in esophageal cancer reveal both cell-type- and cancer-specific epigenetic regulation
Genome Biology (2023)
-
MALAT1 expression is associated with aggressive behavior in indolent B-cell neoplasms
Scientific Reports (2023)
-
Multimodal classification of molecular subtypes in pediatric acute lymphoblastic leukemia
npj Precision Oncology (2023)