Abstract
Understanding the intricacies of lethal prostate cancer poses specific challenges due to difficulties in accurate modeling of metastasis in vivo. Here we show that NPKEYFP mice (for Nkx3.1CreERT2/+; Ptenflox/flox; KrasLSL-G12D/+; R26R-CAG-LSL-EYFP/+) develop prostate cancer with a high penetrance of metastasis to bone, thereby enabling detection and tracking of bone metastasis in vivo and ex vivo. Transcriptomic and whole-exome analyses of bone metastasis from these mice revealed distinct molecular profiles conserved between human and mouse and specific patterns of subclonal branching from the primary tumor. Integrating bulk and single-cell transcriptomic data from mouse and human datasets with functional studies in vivo unravels a unique MYC/RAS co-activation signature associated with prostate cancer metastasis. Finally, we identify a gene signature with prognostic value for time to metastasis and predictive of treatment response in human patients undergoing androgen receptor therapy across clinical cohorts, thus uncovering conserved mechanisms of metastasis with potential translational significance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All relevant data supporting this study are provided in this paper, and reagents will be made available upon request. RNA-seq and WES data can be found in GEO (GSE143815). Previously published genomic data, reanalyzed here, were obtained from GEO (https://www.ncbi.nlm.nih.gov/geo/), dbGAP (https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs001141.v1.p1), TCGA data portal (https://portal.gdc.cancer.gov) and cBioPortal (https://github.com/cBioPortal/datahub/tree/master/public/prad_su2c_2019), as detailed in Supplementary Table 3. Databases used were HomoloGene (NCBI, https://www.ncbi.nlm.nih.gov/homologene) and hallmarks and C2 pathway gene sets from the molecular signatures database (MSigDB) 3.0 (https://www.gsea-msigdb.org/gsea/msigdb). Source data are provided with this paper.
Code availability
The code for UMAP visualization is available at https://github.com/simslab/cluster_diffex2018. The code for bootstrap analysis of evolution tree is available at https://github.com/RabadanLab.
References
Sartor, O. & de Bono, J. S. Metastatic prostate cancer. N. Engl. J. Med. 378, 645–657 (2018).
Scher, H. I. & Sawyers, C. L. Biology of progressive, castration-resistant prostate cancer: directed therapies targeting the androgen-receptor signaling axis. J. Clin. Oncol. 23, 8253–8261 (2005).
Watson, P. A., Arora, V. K. & Sawyers, C. L. Emerging mechanisms of resistance to androgen receptor inhibitors in prostate cancer. Nat. Rev. Cancer 15, 701–711 (2015).
Rickman, D. S., Beltran, H., Demichelis, F. & Rubin, M. A. Biology and evolution of poorly differentiated neuroendocrine tumors. Nat. Med. 23, 1–10 (2017).
Halabi, S. et al. Meta-analysis evaluating the impact of site of metastasis on overall survival in men with castration-resistant prostate cancer. J. Clin. Oncol. 34, 1652–1659 (2016).
Hernandez, R. K. et al. Incidence of bone metastases in patients with solid tumors: analysis of oncology electronic medical records in the United States. BMC Cancer 18, 44 (2018).
Logothetis, C., Morris, M. J., Den, R. & Coleman, R. E. Current perspectives on bone metastases in castrate-resistant prostate cancer. Cancer Metastasis Rev. 37, 189–196 (2018).
Nuhn, P. et al. Update on systemic prostate cancer therapies: management of metastatic castration-resistant prostate cancer in the era of precision oncology. Eur. Urol. 75, 88–99 (2019).
Crawford, E. D., Higano, C. S., Shore, N. D., Hussain, M. & Petrylak, D. P. Treating patients with metastatic castration resistant prostate cancer: a comprehensive review of available therapies. J. Urol. 194, 1537–1547 (2015).
Arriaga, J. M. & Abate-Shen, C. Genetically engineered mouse models of prostate cancer in the postgenomic era. Cold Spring Harb. Perspect. Med. 9, a030528 (2019).
Aytes, A. et al. ETV4 promotes metastasis in response to activation of PI3-kinase and Ras signaling in a mouse model of advanced prostate cancer. Proc. Natl Acad. Sci. USA 110, E3506–E3515 (2013).
Aytes, A. et al. NSD2 is a conserved driver of metastatic prostate cancer progression. Nat. Commun. 9, 5201 (2018).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Wang, X. et al. A luminal epithelial stem cell that is a cell of origin for prostate cancer. Nature 461, 495–500 (2009).
Wang, Z. A., Toivanen, R., Bergren, S. K., Chambon, P. & Shen, M. M. Luminal cells are favored as the cell of origin for prostate cancer. Cell Rep. 8, 1339–1346 (2014).
Zou, M. et al. Transdifferentiation as a mechanism of treatment resistance in a mouse model of castration-resistant prostate cancer. Cancer Discov. 7, 736–749 (2017).
Wang, C., Shen, Y. & Zhu, S. Distribution features of skeletal metastases: a comparative study between pulmonary and prostate cancers. PLoS ONE 10, e0143437 (2015).
Carver, B. S. et al. Reciprocal feedback regulation of PI3K and androgen receptor signaling in PTEN-deficient prostate cancer. Cancer Cell 19, 575–586 (2011).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Stanbrough, M. et al. Increased expression of genes converting adrenal androgens to testosterone in androgen-independent prostate cancer. Cancer Res. 66, 2815–2825 (2006).
McFadden, D. G. et al. Mutational landscape of EGFR-, MYC- and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc. Natl Acad. Sci. USA 113, E6409–E6417 (2016).
Westcott, P. M. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015).
Mueller, S. et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature 554, 62–68 (2018).
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Abida, W. et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proc. Natl Acad. Sci. USA 116, 11428–11436 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Kumar, A. et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat. Med. 22, 369–378 (2016).
Zeller, K. I., Jegga, A. G., Aronow, B. J., O’Donnell, K. A. & Dang, C. V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
Sabo, A. et al. Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis. Nature 511, 488–492 (2014).
Koh, C. M. et al. MYC and prostate cancer. Genes Cancer 1, 617–628 (2010).
Gurel, B. et al. Nuclear MYC protein overexpression is an early alteration in human prostate carcinogenesis. Mod. Pathol. 21, 1156–1167 (2008).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Kaighn, M. E., Narayan, K. S., Ohnuki, Y., Lechner, J. F. & Jones, L. W. Establishment and characterization of a human prostatic carcinoma cell line (PC-3). Invest. Urol. 17, 16–23 (1979).
Wu, T. T. et al. Establishing human prostate cancer cell xenografts in bone: induction of osteoblastic reaction by prostate-specific antigen-producing tumors in athymic and SCID/bg mice using LNCaP and lineage-derived metastatic sublines. Int. J. Cancer 77, 887–894 (1998).
Corey, E. et al. Establishment and characterization of osseous prostate cancer models: intra-tibial injection of human prostate cancer cells. Prostate 52, 20–33 (2002).
Hubbard, G. K. et al. Combined MYC activation and Pten loss are sufficient to create genomic instability and lethal metastatic prostate cancer. Cancer Res. 76, 283–292 (2016).
Magnon, C. et al. Autonomic nerve development contributes to prostate cancer progression. Science 341, 1236361 (2013).
Leibold, J. et al. Somatic tissue engineering in mouse models reveals an actionable role for WNT pathway alterations in prostate cancer metastasis. Cancer Discov. 10, 1038–1057 (2020).
Nowak, D. G. et al. MYC drives Pten/Trp53-deficient proliferation and metastasis due to IL-6 secretion and AKT suppression via PHLPP2. Cancer Discov. 5, 636–651 (2015).
Ellwood-Yen, K. et al. Myc-driven murine prostate cancer shares molecular features with human prostate tumors. Cancer Cell 4, 223–238 (2003).
Wang, L. et al. A prospective genome-wide study of prostate cancer metastases reveals association of wnt pathway activation and increased cell cycle proliferation with primary resistance to abiraterone acetate-prednisone. Ann. Oncol. 29, 352–360 (2018).
Karnes, R. J. et al. Validation of a genomic classifier that predicts metastasis following radical prostatectomy in an at risk patient population. J. Urol. 190, 2047–2053 (2013).
Ross, A. E. et al. Tissue-based genomics augments post-prostatectomy risk stratification in a natural history cohort of intermediate- and high-risk men. Eur. Urol. 69, 157–165 (2016).
Beaver, J. A., Kluetz, P. G. & Pazdur, R. Metastasis-free survival: a new end point in prostate cancer trials. N. Engl. J. Med. 378, 2458–2460 (2018).
Armenia, J. et al. The long tail of oncogenic drivers in prostate cancer. Nat. Genet. 50, 645–651 (2018).
Chung, J. H. et al. Prospective comprehensive genomic profiling of primary and metastatic prostate tumors. JCO Precis. Oncol. https://doi.org/10.1200/PO.18.00283 (2019).
Kornberg, Z., Cooperberg, M. R., Spratt, D. E. & Feng, F. Y. Genomic biomarkers in prostate cancer. Transl. Androl. Urol. 7, 459–471 (2018).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Easterly, M. E., Foltz, C. J. & Paulus, M. J. Body condition scoring: comparing newly trained scorers and micro-computed tomography imaging. Lab. Anim. (NY) 30, 46–49 (2001).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://arxiv.org/abs/1303.3997?upload=1://arxiv.org/abs/1303.3997?upload=1 (2013).
Benjamin, D. et al. Calling somatic SNVs and indels with Mutect2. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/861054v1 (2019).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Oshima, K. et al. Mutational landscape, clonal evolution patterns, and role of RAS mutations in relapsed acute lymphoblastic leukemia. Proc. Natl Acad. Sci. USA 113, 11306–11311 (2016).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Levitin, H. M. et al. De novo gene signature identification from single-cell RNA-seq with hierarchical Poisson factorization. Mol. Syst. Biol. 15, e8557 (2019).
Trabzonlu, L. et al. Molecular pathology of high-grade prostatic intraepithelial neoplasia: challenges and opportunities. Cold Spring Harb. Perspect. Med. 9, a030403 (2019).
Thorek, D. L. et al. Internalization of secreted antigen-targeted antibodies by the neonatal Fc receptor for precision imaging of the androgen receptor axis. Sci. Transl. Med. 8, 367ra167 (2016).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Acknowledgements
We are grateful to D. Adams for assistance with the initial WES and E. Davicioni and T. Lotan for sharing clinical cohorts. We thank A. Aytes, R. Dalla-Favera, G. Karsenty and M. Shen for comments on the manuscript. This research was funded in part through the National Institutes of Health (NIH)/NCI Cancer Center Support Grant P30CA013696, which supported the Molecular Pathology, Flow Cytometry, Genomics and High Throughput Screening and Oncology Precision Therapeutics and Imaging Cores. The research was supported by NIH grants R01 CA193442, R01 CA173481 and R01 CA183929 (to C.A.S.), P50 CA211024 (to M.A.R.), U54CA209997 (to P.A.S. and C.A.S.), P50 CA097186 (to P.S.N.) and DOD grant W81XWH-17-1-0415 (to P.S.N.). P.S.N. was supported by Stand Up To Cancer-Prostate Cancer Foundation Prostate Cancer Dream Team Award (SU2C-AACR-DT0712). A.M. was supported by start-up funds from Rutgers School of Health Professions. J.M.A. was supported by a postdoctoral training grant from the Department of Defense Prostate Cancer Research Program (W81XWH-15-1-0185), an Irving Institute/Clinical Trials Office Pilot Award funded by the National Center for Advancing Translational Sciences, NIH (UL1TR001873) and the Dean’s Precision Medicine Research Fellowship from the Irving Institute for Clinical and Translational Research at CUIMC (UL1TR001873). S.P. was supported by New Jersey Commission on Cancer Research Pre-Doctoral Fellowship (DCHS20PPC028). M.Z. was supported by the National Center for Advancing Translational Sciences, NIH, grant number UL1TR001873. A.G. is a recipient of a Prostate Cancer Foundation Young Investigator Award. C.J.M. was supported by NIH fellowship F31CA210607. C.A.S. is supported by the TJ Martell Foundation for Leukemia, Cancer and AIDS Research and the Prostate Cancer Foundation and is an American Cancer Society Research Professor.
Author information
Authors and Affiliations
Contributions
J.M.A, A.M. and C.A.S. conceived and designed the study. J.M.A., S.P., M.A., J.Z., M.Z., A.G., C.J.M., J.Y.K., A.R., I.C., R.K.V., H.H., O.E., B.O., J.F., J.L., E.S.A., P.S.N., F.Y.F., M.A.R., A.M.D., R.R., P.A.S. and A.M. collected and assembled data. J.M.A., M.Z. and A.G. collected mouse data. M.A., A.R., I.C., R.K.V., H.H., O.E., B.O., J.F., R.J.K., J.L., E.S.A, P.S.N., F.Y.F., A.M.D. and M.A.R. provided study materials or patient samples and J.M.A., S.P., M.A., J.Z., M.Z., A.G., C.J.M., J.Y.K., A.R., I.C., O.E., B.O., J.L., E.S.A., P.S.N., F.Y.F., M.A.R., A.M.D., R.R., P.A.S., A.M. and C.A.S. analyzed and interpreted the data. S.P., A.M. and P.A.S. processed and analyzed single-cell data, J.Z., C.J.M. and R.R. processed and analyzed whole-exome data and J.M.A., S.P. and A.M. processed and analyzed bulk RNA-seq data. S.P., M.A., I.C., P.S.N. and A.M. analyzed human tumor sequencing data. J.M.A., J.Y.K. and A.D.M performed immunohistochemistry. All authors participated in writing and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.M.D. has sponsored research funding from Janssen R&D and consults for Cepheid. R.J.K. receives Royalties from GenomeDx. E.S.A. has served as a paid consultant/advisor to Janssen, Pfizer, Sanofi, Dendreon, Bayer, Bristol Myers Squibb, Amgen, Merck, AstraZeneca and Clovis; has received research grants to his institution from Janssen, Johnson&Johnson, Sanofi, Bristol Myers Squibb, Pfizer, AstraZeneca, Celgene, Merck, Bayer and Clovis; and is an inventor of a biomarker technology that has been licensed to QIAGEN. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional phenotypic analyses of NPKEYFP prostate tumors.
a, Representative bright field and ex vivo fluorescence images of prostate, lung, and bone (spine) from NPEYFP mice (n = 35), NPKEYFP mice without bone metastases (n = 59), and NPKEYFP mice with bone metastases (n = 47). b–j, Comparison of NPKEYFP mice with (n = 47) or without (n = 59) bone metastasis. See also Supplementary Table 1. b, Overall survival; p-value calculated using a two-tailed log-rank test. c, Bladder obstruction; p-value calculated using a two-sided Fisher’s exact test. d–g, Dot-plots showing tumor weight (d) and metastatic load (number of metastases/mouse) to lungs (e), liver (f) and brain (g). p-values were calculated by two-tailed Mann-Whitney test, center-lines show the mean and error bars depict SD. h, Distribution of metastases to specific bone types in NPKEYFP mice at the time of euthanasia. Shown is the mean with standard deviation; n = 106 mice (n = 59 without bone metastases and n = 47 with bone metastases). i, j, Longitudinal analysis of micro-metastasis in non-terminal mice dissected at the ages shown (n = 26). i, Bar graphs showing the percentage of mice with micro-metastasis at 3 months (n = 7), 4 months (n = 17) and 5 months (n = 2). j, Bar graphs showing the percentage of mice with bone or lung micro-metastases (n = 26).
Extended Data Fig. 2 Analyses of androgen-intact and castrated of NPKEYFP mice.
Comparison of intact (n = 106) and castrated (n = 22) NPKEYFP mice. See also Supplementary Table 1. a, Kaplan-Meier curves showing overall survival; p-value was calculated using a two-tailed log-rank test. b, c, Dot-plots showing tumor weight (b) and number of bone metastasis (c). p-values were calculated by two-tailed Mann-Whitney test, center-lines show the mean and error bars depict SD.d, Bar graphs showing the percentage of mice with metastasis to the indicated organs. e, f, Dot-plots showing relative AR activity levels (that is, NES defined based on enrichment of AR signature (based on18) in each sample) (e) and relative neuroendocrine (NE) activity (that is, NES defined based on enrichment of NEPC signature (based on19) in each sample (f) comparing intact (n = 13) or castrated (n = 6) primary tumors, intact (n = 9) or castrated (n = 2) lung metastasis and intact (n = 10) or castrated (n = 2) bone metastasis from NPKEYFP mice (Supplementary Table 2). g, Gene Set Enrichment Analyses (GSEA) comparing a bone metastasis signatures from castrated mice used as a query and bone metastasis signature from non-castrated (intact) NPKEYFP mice used as a reference (Supplementary Table 2i,j). NES (normalized enrichment score) and p-values were estimated using 1,000 gene permutations. NS, non-significant (P < 0.05).
Extended Data Fig. 3 Comparison of MYC and RAS in mouse and human prostate cancer.
a–c, Copy number variant (CNV) analyses of KRAS and MYC in mouse and human prostate cancer. a, Kras, Cdkn2a/b and Myc loci, inferred from whole-exome sequencing of NPKEYFP prostate tumors. Color coding reflects amplifications or deletions in five individual mice (Supplementary Table 4b). b, c, Summary of gains in MYC (b) and KRAS (c) in human prostate cancer comparing primary tumors from TCGA (n = 489) and metastases from SU2C (n = 429) using cBioportal49. P values were calculated using a Fisher’s exact test comparing samples with all gains versus no gains. d, e, Box plots depicting Myc pathway and Ras pathway activation in primary tumors and metastases from intact or castrated NPKEYFP mice (primary tumors: n = 13 intact and n = 6 castrated; lung metastases: n = 9 intact and n = 2 castrated; bone metastases: n = 10 intact and n = 2 castrated). The distribution of the activity scores (y-axis) for Myc activity is based on single-sample GSEA in panel d, and Ras activity levels is based on the absolute-valued average of RAS-related genes as in11,33 in panel e. P-values were estimated using two-sample one-tailed Welch t-test, boxes show the 25th–75th percentile with the median, and whiskers show the minimum–maximum values. f, g, Violin plots depicting the distribution of MYC and RAS pathway activation in primary tumors and metastases comparing human primary tumors (TCGA, n = 497) versus metastases (SU2C, n = 270). In panel f, the distribution of the NESs (y-axis) represent MYC activity levels based on single-sample GSEA (see Extended data Fig. 4d) In panel g, the activity scores (y-axis) represent RAS pathway activity levels based on the absolute-valued average of RAS-related genes (as in11,33). P-values were estimated using two-sample one-tailed Welch t-test. In the violin plots with embedded box plots, boxes show the 25th–75th percentile, center-lines show the median, and whiskers show the minimum–maximum values. h, i, Heatmap representation showing the correlation of MYC and RAS pathway activity in mouse (h) and human (i) prostate cancer. Panel h shows Myc and Ras pathway activity in mouse NPKEYFP primary tumors and bone metastases. Panel i shows MYC and RAS pathway activity in human primary tumors (TCGA, n = 497) and metastases (SU2C, n = 270). Gleason scores are shown for the primary tumors; metastases include all metastases in the SU2C cohort. In panels h, i, Spearman correlation rho- and p-values are shown. j, Mouse NPKEYFP primary tumors and bone metastases classified as MYC- or RAS-activated are depicted in a heatmap in red, whereas those without MYC- or RAS-activation are represented in blue. Samples were considered Myc-activated if Myc activity scores were greater than the average across the cohorts. Samples were considered Ras-activated if Ras activity scores were greater than the average across the cohorts. The percentage of cases in which Myc and Ras are co-activated are shown; two-tailed p-value was calculated using Fisher’s exact test.
Extended Data Fig. 4 Additional analyses of MYC activity in prostate tumors and metastases.
a, Cross-species pathway analysis. Pathway-based GSEA comparing pathways enriched in the FHCRC human bone metastasis signature (Supplementary Table 6d) with those enriched in the mouse bulk RNA bone metastasis signature (Supplementary Table 6a). NES and p-values were estimated using 1,000 pathway permutations. b, Stouffer integration of the leading-edge pathways from the GSEA comparing the mouse (Supplementary Table 6a) and the two human bone metastases signatures (Supplementary Tables 6c,d) from panel a and Fig. 4b. The x-axis shows the Stouffer integrated NES. c, Bar graphs summarizing NES scores from GSEA of bone metastasis signatures from NPKEYFP mice (Supplementary Table 2c), and the Balk and FHCRC human cohorts (Supplementary Tables 3c,d) showing enrichment of three independent MYC signatures: “Hallmarks” (human), “Dang” (human)29 and “Sabo” (mouse)30. NES and p-values were estimated using 1,000 gene permutations. d, Heatmap representation of single-sample GSEA enrichment of MYC activity based on enrichment of the Hallmarks MYC pathway in primary tumors from TCGA (n = 497) and metastases from SU2C (n = 270) (Supplementary Table 3). Gleason scores are shown for the primary tumors; metastases include all metastases in the SU2C cohort. Colors correspond to NES.
Extended Data Fig. 5 Additional analysis of Myc function in an allograft model of bone metastasis.
a, Bar graphs and crystal violet staining of colony formation assays of NPK bone cells two weeks after treatment with shControl or shRNAs targeting Myc (shMyc#1 and shMyc#2). b, Comparison of lung and bone from Nude mouse hosts implanted via intracardiac injection with green fluorescent protein (GFP)-tagged cells derived from primary tumors of non-metastatic NP mice (NPGFP cells, reported in;16 n = 2) or the NPKEYFP bone cells (n = 10). Shown are representative ex vivo fluorescence or H&E images. Scale bars represent 0.1 cm for the ex vivo fluorescence images and 50 μm for all other images.
Extended Data Fig. 6 Analyses of MYC silencing in a human tumor growth in bone.
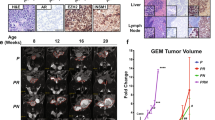
a, Strategy. PC3 cells engineered to express luciferase and GFP (PC3-Luciferase-GFP cells). Cells were infected with a control shRNA (shControl) or shRNA to silence MYC (shMYC#1 or shMYC#2) and implanted into the tibia of NOD-SCID mouse hosts. b, Western blot image of total protein extracts. Shown are the approximate molecular weight markers (kDa); Actin is a control for protein loading. Shown is a representative blot from two independent experiments. The uncropped Western image is shown in Source data Extended Fig. 6. c. Immunostaining for MYC in tumors from mice that had been injected with cells expressing the indicated shRNAs. Scale bars represent 50 μm. d, Bar graphs and crystal violet staining of colony formation assays of PC3-Luciferase-GFP cells two weeks after treatment with the shRNAs as indicated. e, Growth curves comparing PC3-Luciferase-GFP cells infected with shRNA (n = 10/group). P-value shown for day 52 was estimated by two-way ANOVA with Sidak’s multiple comparisons against shControl. f. Representative IVIS bioluminescence imaging used for panel e. g. Representative images from the time of sacrifice of tibiae implanted with the PC3-Luciferase-GFP cells infected with shRNA (n = 10/group). Shown are ex vivo imaging of GFP fluorescence, to visualize the tumor, and corresponding micro-computed tomography (CT) images, to show areas of osteolysis as is typical of PC3 tumors in bone36. Also shown are representative H&E and immunostaining for GFP. In a and f, bars show mean and error bars the SD, (n = 3) and p-value is shown for One-way ANOVA with Dunnett’s multiple comparison test, compared to shControl. Scale bars represent 0.1 cm for the ex vivo fluorescence images and 50 μm for all other images.
Extended Data Fig. 7 Additional analyses of discovery of a MYC-correlated signature in prostate cancer metastasis.
a,b, GSEA using the PROMOTE-559 gene signature (Supplementary Table 8) to query the bone metastasis gene signature from the NPKEYFP mice (Supplementary Table 2c) (in a) and the human bone metastasis gene signature (Supplementary Table 3c) (in b); NESs and p-values were estimated using 1,000 gene permutations. c, Association with adverse outcome for metastasis. Each of the META-55 genes was evaluated by univariable Cox proportional hazards analysis for time-to-metastasis outcome in the TCGA dataset (n = 336 with available time to follow-up, Supplementary Table 3) and ranked by the strength of the association (that is, Wald test p-value), with a cutoff at p-value<10–7 from Wald test used to identify the 16 top-genes constituting the META-16 gene signature (Supplementary Table 8). d, Random model. To evaluate the probability that not any random group of 16 genes would be upregulated in the SU2C (n = 270) versus the TCGA (n = 497) cohorts, we constructed a null model using 10,000 iterations, with the x-axis showing -log2 p-value (from the two-sample one-tailed Welch t-test) between TCGA and SU2C comparisons and y-axis showing its probability density. The p-value of this random model thus represents an estimate of the number of times two-sample one-tailed Welch t-test p-values for a random 16 genes reached or outperformed two-sample one-tailed Welch t-test p-values for the META-16 genes. The p-value for the analogous random model for META-55 was P = 0.036.
Extended Data Fig. 8 Additional analyses of META-55 and META-16 in prostate cancer metastasis.
a, Scaled expression (DESeq2 normalized values) of META-55 in single-cell UMAP projections of primary tumors and bone metastases (see Fig. 7c, d). Shown is the correlation between META-55 expression at the single-cell level with MYC pathway activity (Spearman’s rank correlation rho and p-values). b, c, Heatmap representation of single-sample GSEA enrichment of the META-16 (b) and META-55 (c) gene signatures in primary tumors from TCGA (n = 497) and metastases from SU2C (n = 270) (Supplementary Table 3). Colors correspond to NES. d, e, Violin plots depicting the distribution of the NESs (y-axis) which reflect activity levels of META-16 (d) and META-55 (e) in primary tumors from TCGA (n = 497) compared with metastases from SU2C (n = 270). The p-value was estimated using two-sample one-tailed Welch t-test. In inset box-plots, boxes show the 25th–75th percentile, center-lines show the median, and whiskers show the minimum–maximum values. f, g, Heatmap representation of expression levels of META genes (as indicated) in each of the individual samples from the TCGA (n = 497) and SU2C (n = 270) cohorts. Gleason scores are shown for the primary tumors; metastases include all metastases in the SU2C cohort. Shown are row-scaled expression values (color). Panel f shows the 10 genes from the META-16 signature that do not co-reside with MYC on chromosome 8q, indicated as META-10. Panel g shows the META-55 genes.
Extended Data Fig. 9 Additional validation of the META-16 gene signature.
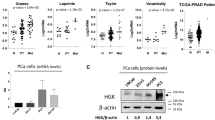
a, Quantitative reverse transcriptase PCR (qRT-PCR) of META-16 in the CUIMC cohort of bone metastases (n = 5) compared with high-Gleason grade primary prostate tumors (n = 10). Indicated p-values were estimated using a two-tailed Mann-Whitney test compared to the average of all primary tumors. In box plots, boxes show the 25th–75th percentile, center-lines show the median, and whiskers show the minimum–maximum values. b, c, Heatmaps showing expression levels of META-16 genes determined by qRT-PCR following MYC silencing in human and mouse prostate cancer cells. b, qRT-PCR using RNA obtained from subcutaneous PC3-Luc-GFP tumors expressing shRNA against MYC (shMYC#1) or control shRNA (shControl). c, qRT-PCR using RNA obtained from NPKEYFP bone cells grown in vitro and infected with the indicated shRNAs. Scaled values represent ratios of expression compared to shControl for each gene. In b, c, p-values were estimated using z-score sums of all genes using two-tailed, unpaired t-test (b) or one-way ANOVA with Dunnett’s multiple comparisons against shControl (c).
Extended Data Fig. 10 Additional validation of the META-55/META-16 gene signatures and survival analyses.
a, b, Heatmaps of hierarchical consensus clustering analysis used to define tumors with high (brown cluster) and low (green cluster) expression of META-16 in MAYO (n = 235) and JHMI (n = 260) cohorts, as indicated (Supplementary Table 3). Brown vertical bars on the second from top row represent patients that developed distant metastasis. Colors represent row-scaled expression values. c, d, Kaplan-Meier survival analyses comparing patients with low and high overall expression of META-55. The p-values were estimated using a log-rank test. e, Multivariable survival analysis of the META-55 gene signature in the JHMI and MAYO cohorts showing significant association with metastasis-free survival but not with prostate-cancer specific mortality (HR = hazard ratio, CI = confidence interval, p-values estimated from Coxproportional hazards model), adjusted for age, pathological Gleason score/grade at diagnosis, pre-PSA, seminal vesicle invasion (SVI), lymph node invasion (LNI), and extra-prostatic extension (EPE). f, g, Kaplan-Meier survival analyses comparing patients from SU2C cohort with the low and high MYC activity with respect to treatment-associated survival (that is, time from the start of treatment with androgen receptor signaling inhibitor (ARSi) therapy, to death or last follow-up, n = 75 patients) or treatment-associated disease progression (that is, time on treatment with ARSIs, n = 56) as defined in26. The p-values were estimated using a log-rank test.
Supplementary information
Supplementary Tables
Supplementary Tables 1–10
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Arriaga, J.M., Panja, S., Alshalalfa, M. et al. A MYC and RAS co-activation signature in localized prostate cancer drives bone metastasis and castration resistance. Nat Cancer 1, 1082–1096 (2020). https://doi.org/10.1038/s43018-020-00125-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00125-0
This article is cited by
-
CRISPR/Cas9 model of prostate cancer identifies Kmt2c deficiency as a metastatic driver by Odam/Cabs1 gene cluster expression
Nature Communications (2024)
-
In vivo genome-wide CRISPR screening identifies CITED2 as a driver of prostate cancer bone metastasis
Oncogene (2024)
-
Mechanism-centric regulatory network identifies NME2 and MYC programs as markers of Enzalutamide resistance in CRPC
Nature Communications (2024)
-
Loss of Pip4k2c confers liver-metastatic organotropism through insulin-dependent PI3K-AKT pathway activation
Nature Cancer (2024)
-
Engineering prostate cancer in vitro: what does it take?
Oncogene (2023)