Abstract
Macroautophagy (hereafter autophagy) degrades and recycles intracellular components to sustain metabolism and survival during starvation. Host autophagy promotes tumor growth by providing essential tumor nutrients. Autophagy also regulates immune cell homeostasis and function and suppresses inflammation. Although host autophagy does not promote a T-cell antitumor immune response in tumors with low tumor mutational burden (TMB), whether this was the case in tumors with high TMB was not known. Here we show that autophagy, especially in the liver, promotes tumor immune tolerance by enabling regulatory T-cell function and limiting stimulator of interferon genes, T-cell response and interferon-γ, which enables growth of high-TMB tumors. We have designated this as hepatic autophagy immune tolerance. Autophagy thereby promotes tumor growth through both metabolic and immune mechanisms depending on mutational load and autophagy inhibition is an effective means to promote an antitumor T-cell response in high-TMB tumors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
10X scRNA-seq data have been deposited in the Gene Expression Omnibus under accession ID GSE154654. Whole-exome sequencing data generated in this study can be found in the Gene Expression Omnibus under accession ID GSE155556. All other data supporting the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Change history
16 August 2021
A Correction to this paper has been published: https://doi.org/10.1038/s43018-021-00252-2
References
Karsli-Uzunbas, G. et al. Autophagy is required for glucose homeostasis and lung tumor maintenance. Cancer Discov. 4, 914–927 (2014).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004).
Guo, J. Y. et al. Autophagy provides metabolic substrates to maintain energy charge and nucleotide pools in Ras-driven lung cancer cells. Genes Dev. 30, 1704–1717 (2016).
Kamada, Y., Sekito, T. & Ohsumi, Y. Autophagy in yeast: a TOR-mediated response to nutrient starvation. Curr. Top. Microbiol. Immunol. 279, 73–84 (2004).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
Kimmelman, A. C. & White, E. Autophagy and tumor metabolism. Cell Metab. 25, 1037–1043 (2017).
Amaravadi, R., Kimmelman, A. C. & White, E. Recent insights into the function of autophagy in cancer. Genes Dev. 30, 1913–1930 (2016).
Poillet-Perez, L. & White, E. Role of tumor and host autophagy in cancer metabolism. Genes Dev. 33, 610–619 (2019).
Sousa, C. M. et al. Pancreatic stellate cells support tumour metabolism through autophagic alanine secretion. Nature 536, 479–483 (2016).
Yang, A. et al. Autophagy sustains pancreatic cancer growth through both cell-autonomous and nonautonomous mechanisms. Cancer Discov. 8, 276–287 (2018).
Poillet-Perez, L. et al. Autophagy maintains tumour growth through circulating arginine. Nature 563, 569–573 (2018).
Degenhardt, K. et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 10, 51–64 (2006).
Mathew, R. et al. Functional role of autophagy-mediated proteome remodeling in cell survival signaling and innate immunity. Mol. Cell 55, 916–930 (2014).
Merkley, S. D., Chock, C. J., Yang, X. O., Harris, J., Castillo, E. F. & Modulating, T. Cell responses via autophagy: the intrinsic influence controlling the function of both antigen-presenting cells and T cells. Front. Immunol. 9, 2914 (2018).
McFadden, D. G. et al. Mutational landscape of EGFR-, MYC-, and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc. Natl Acad. Sci. USA 113, E6409–E6417 (2016).
Bignell, G. R. et al. Signatures of mutation and selection in the cancer genome. Nature 463, 893–898 (2010).
Westcott, P. M. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015).
Wang, J. et al. UV-induced somatic mutations elicit a functional T cell response in the YUMMER1.7 mouse melanoma model. Pigment Cell Melanoma Res. 30, 428–435 (2017).
Wei, S. C., Duffy, C. R. & Allison, J. P. Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 8, 1069–1086 (2018).
Guo, J. Y. et al. Autophagy suppresses progression of K-Ras-induced lung tumors to oncocytomas and maintains lipid homeostasis. Genes Dev. 27, 1447–1461 (2013).
Wei, H. et al. Suppression of autophagy by FIP200 deletion inhibits mammary tumorigenesis. Genes Dev. 25, 1510–1527 (2011).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Panda, A. et al. Identifying a clinically applicable mutational burden threshold as a potential biomarker of response to immune checkpoint therapy in solid tumors. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00146 (2017).
Yang, A. S., Monken, C. E. & Lattime, E. C. Intratumoral vaccination with vaccinia-expressed tumor antigen and granulocyte macrophage colony-stimulating factor overcomes immunological ignorance to tumor antigen. Cancer Res. 63, 6956–6961 (2003).
Poillet-Perez, L. et al. Autophagy maintains tumour growth through circulating arginine. Nature 563, 569–573 (2018).
Chu, H. et al. Gene-microbiota interactions contribute to the pathogenesis of inflammatory bowel disease. Science 352, 1116–1120 (2016).
Wei, J. et al. Autophagy enforces functional integrity of regulatory T cells by coupling environmental cues and metabolic homeostasis. Nat. Immunol. 17, 277–285 (2016).
Sawant, D. V. et al. Adaptive plasticity of IL-10(+) and IL-35(+) Treg cells cooperatively promotes tumor T cell exhaustion. Nat. Immunol. 20, 724–735 (2019).
Sliter, D. A. et al. Parkin and PINK1 mitigate STING-induced inflammation. Nature 561, 258–262 (2018).
Castro, F., Cardoso, A. P., Goncalves, R. M., Serre, K. & Oliveira, M. J. Interferon-γ at the crossroads of tumor immune surveillance or evasion. Front. Immunol. 9, 847 (2018).
Yamamoto, K. et al. Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I. Nature 581, 100–105 (2020).
Cunha, L. D. et al. LC3-associated phagocytosis in myeloid cells promotes tumor immune tolerance. Cell 175, 429–441 (2018).
Lee, H. K. et al. In vivo requirement for Atg5 in antigen presentation by dendritic cells. Immunity 32, 227–239 (2010).
Xu, X. et al. Autophagy is essential for effector CD8(+) T cell survival and memory formation. Nat. Immunol. 15, 1152–1161 (2014).
DeVorkin, L. et al. Autophagy regulation of metabolism is required for CD8(+) T cell anti-tumor immunity. Cell Rep. 27, 502–513 (2019).
Yu, Y. et al. STING-mediated inflammation in Kupffer cells contributes to progression of nonalcoholic steatohepatitis. J. Clin. Invest. 129, 546–555 (2019).
Appleman, L. J. et al. Targeting autophagy and immunotherapy with hydroxychloroquine and interleukin 2 in patients with metastatic renal cell carcinoma (mRCC): a Cytokine Working Group study. J. Clin. Oncol. 36, 106–106 (2018).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Gan, B. et al. Role of FIP200 in cardiac and liver development and its regulation of TNF-α and TSC-mTOR signaling pathways. J. Cell Biol. 175, 121–133 (2006).
Meeth, K., Wang, J. X., Micevic, G., Damsky, W. & Bosenberg, M. W. The YUMM lines: a series of congenic mouse melanoma cell lines with defined genetic alterations. Pigment Cell Melanoma Res. 29, 590–597 (2016).
Summerhayes, I. C. & Franks, L. M. Effects of donor age on neoplastic transformation of adult mouse bladder epithelium in vitro. J. Natl Cancer Inst. 62, 1017–1023 (1979).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Koboldt, D. C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
This work was supported by National Institutes of Health grants: R01CA193970 (to E.W.), R01CA163591 (to E.W. and J.D.R.), R01CA193970 (to J.M.M.), R01CA211066 (to J.L.G), R01CA243547 (to E.W., S.G. and E.C.L.) and P30CA072720 (Rutgers Cancer Institute of New Jersey) and by the Krauss Foundation (to E.W. and E.C.L.). C.S.C. is supported in part by Stand Up to Cancer and National Science Foundation grant no. 1546101. L.P.P. and S.V.L. received support from a postdoctoral fellowship from the New Jersey Commission for Cancer Research (DHFS16PPC034 and DCHS19PPC009, respectively). Services, results and/or products in support of the research project were generated by funding from the Rutgers Cancer Institute of New Jersey Comprehensive Genomics and Immune Monitoring Shared Resources supported, in part by NCI-CCSG P30CA072720-5920 and by the Lewis-Sigler Institute for Integrative Genomics (Genomics Core Facility).
Author information
Authors and Affiliations
Contributions
L.P.P. performed the majority of the experimental work and wrote the manuscript. D.W.S. performed/assisted with the MB49 allografts and CD4/CD8 or CD25 depletion and NanoString analysis. Y.Y. developed the methods and provided the mice required for generating Atg5∆/∆ hosts. S.V.L. and C.S.C. analyzed the whole-exome sequencing data and scRNA-seq data. M.I. assisted with the in vivo experiments. P.K.B. assisted with tumor processing and analyzed the data for the NanoString. Z.S.H. assisted with animal husbandry. J.V. performed the flow cytometry, analyzed T-cell exhaustion markers and performed cell isolation and library preparation for single-cell sequencing. M.W.B. provided the YUMM1.1 melanoma cells. J.D.R and S.G provided advice throughout the project and on the manuscript. J.C. provided expertise on CRISPR Cas9. M.H and J.L.G provided the Fip200flox/flox mice. J.M.M. provided expertise in melanoma. E.W. and E.C.L. are the leading principal investigators who conceived the project, supervised research and edited the paper.
Corresponding author
Ethics declarations
Competing interests
E.W. is co-founder of Vescor Therapeutics and is a consultant for Novartis. S.G. has consulted for Roche, Merck, Novartis, Foundation Medicine and Foghorn Therapeutics. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Loss of host autophagy does not decrease the growth of low TMB tumors in a T cell-dependent manner.
a, CD3+, CD4+, CD8+ T cells (percentage) in high TMB tumors (MB49) grown on female Atg7+/+ and Atg7Δ/Δ (n = 5 mice each) hosts following αCD4/8, analyzed by flow cytometry. b, Representative IHC images and quantification of CD3+, CD4+ and CD8+ T cells in YUMM1.3 tumors from male Atg7+/+ and Atg7Δ/Δ hosts (n = 5 mice each). c, d, e, f, Comparison of YUMM1.3 tumor growth on male (c, d) Atg7+/+, Atg7+/+ + αCD4/8, Atg7Δ/Δ and Atg7Δ/Δ + αCD4/8 (n = 10 tumors each) and female (e, f) Atg7+/+(n = 6 tumors), Atg7+/+ + αCD4/8 (n = 12 tumors), Atg7Δ/Δ (n = 12 tumors) and Atg7Δ/Δ + αCD4/8 (n = 12 tumors). Tumor pictures (c, e), and tumor weight and ratio (antibody treated/control) (d, f) from the experimental endpoint. g, h, Comparison of MB49 tumor growth on female Atg7+/+(n = 14 tumors), Atg7+/+ + αCD4 (n = 12 tumors), Atg7+/+ + αCD8 (n = 12 tumors), Atg7+/+ + αCD4/8 (n = 8 tumors), Atg7Δ/Δ (n = 12 tumors), Atg7Δ/Δ + αCD4 (n = 14 tumors), Atg7Δ/Δ + αCD8 (n = 14 tumors) and Atg7Δ/Δ + αCD4/8 (n = 12 tumors). Tumor pictures (g), and tumor weight and ratio (antibody treated/control) (h) from the experimental endpoint. All data are mean +/- s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.001 using two-sided Student’s t-test. Source data available for a, b, d, f, h.
Extended Data Fig. 2 Loss of host autophagy through Atg5 deletion decreases tumor growth through a T cell-dependent mechanism.
a, Experimental design to induce conditional whole-body Atg5 deletion (Atg5Δ/Δ), and wild-type (Atg5+/+) controls without and with T cell depletion. Ubc-CreERT2/+;Atg5+/+ and Ubc-CreERT2/+;Atg5flox/flox mice were injected with TAM to delete Atg5 and were then injected subcutaneously with tumor cells and intraperitoneally with αCD4/8. Tumor growth was monitored over three weeks. b, c, d, e, Comparison of MB49 tumor growth on male (b, c) Atg5+/+(n = 8 tumors), Atg5+/+ + αCD4/8 (n = 6 tumors), Atg5Δ/Δ (n = 8 tumors) and Atg5Δ/Δ + αCD4/8 (n = 8 tumors) and female (d, e) Atg5+/+(n = 12 tumors), Atg5+/+ + αCD4/8 (n = 10 tumors), Atg5Δ/Δ (n = 12 tumors) and Atg5Δ/Δ + αCD4/8 (n = 10 tumors). Tumor pictures (b, d), and tumor weight and ratio (antibody treated/control) (c, e) from the experimental endpoint. f, Representative IHC images and quantification of CD3+, CD4+ and CD8+ T cells in MB49 tumors from male Atg5+/+ and Atg5Δ/Δ hosts (n = 5 mice each). g, h, Comparison of UV YUMM1.1-9 tumor growth on male (g) Atg7+/+ (n = 10 tumors) and Atg7Δ/Δ (n = 8 tumors) and female (h) Atg7+/+ (n = 16 tumors) and Atg7Δ/Δ (n = 12 tumors) hosts. Tumor pictures and tumor weight from the experimental endpoint. All data are mean +/- s.e.m. *P < 0.05, ***P < 0.001, ****P < 0.0001 using two-sided Student’s t-test. Source data available for c, e, f, g, h.
Extended Data Fig. 3 Loss of autophagy reduces Tregs and increases immune cell fraction.
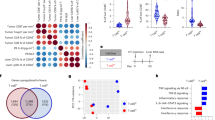
a, Overall NanoString gene expression analysis (750 genes) from tumors on female Atg7+/+ and Atg7Δ/Δ hosts with or without αCD4/8 (n = 2 tumors each). Red: high, blue: low expression level. b, Representative IHC images and quantification of FOXP3+ cells in MB49 tumors from female Atg7+/+ and Atg7Δ/Δ hosts (n = 5 tumors each). c, FOXP3+ Treg cells, CD4+ and CD8+ T cells (percentage) in high TMB tumors (MB49) grown on female (n = 3 tumors) Atg7+/+ and Atg7Δ/Δ hosts following αCD25, analyzed by flow cytometry. d, Representative IHC images and quantification of FOXP3+ cells in MB49 tumors from female Atg7+/+ and Atg7Δ/Δ hosts following αCD25 (n = 3 tumors each). e, Survival analysis of Atg7+/+ (n = 19 mice), Atg7Δ/Δ (n = 19 mice), Stinggt/gt;Atg7+/+ (n = 15 mice) and Stinggt/gt;Atg7Δ/Δ (n = 13 mice) mice. f, scRNA-seq UMAP projection of cell cluster from tumors from female Atg7+/+ and Atg7Δ/Δ hosts (n = 3 tumors each). All data are mean +/- s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 using two-sided Student’s t-test. Source data available for b, c, d.
Extended Data Fig. 4 Loss of autophagy increases high TMB tumor growth in an IFNγ-dependent manner.
a, Survival analysis of Atg7+/+ (n = 16 mice), Atg7Δ/Δ (n = 14 mice), IFNγ-/-;Atg7+/+ (n = 6 mice) and IFNγ-/-;Atg7Δ/Δ (n = 5 mice) hosts. b, Representative H&E tissue staining from IFNγ-/-;Atg7+/+ and IFNγ-/-;Atg7Δ/Δ hosts. Images are representative of two independent experiments. c, d, e, f. Comparison of UV YUMM1.1-9 tumor growth on male (c, d) Atg7+/+ (n = 18 tumors), IFNγ-/-;Atg7+/+ (n = 18 tumors), Atg7Δ/Δ (n = 10 tumors), IFNγ-/-;Atg7Δ/Δ (n = 18 tumors) and female (e, f) Atg7+/+ (n = 16 tumors), IFNγ-/-;Atg7+/+ (n = 18 tumors), Atg7Δ/Δ (n = 16 tumors), IFNγ-/-;Atg7Δ/Δ (n = 18 tumors) hosts. Tumor pictures (c, e), and tumor weight and ratio (antibody treated/control) (d, f) from the experimental endpoint. g, Cropped Western blotting showing expression of B2m in MB49 shC and shB2m tumors from Atg7+/+ and Atg7Δ/Δ hosts (n = 3 tumors each) from one experiment. Actin was used a loading control. All data are mean +/- s.e.m. **P < 0.01, ***P < 0.001 using two-sided Student’s t-test. Source data available for d, f.
Extended Data Fig. 5 Loss of autophagy, not LC3-associated phagocytosis (LAP), decreases tumor growth through a T cell-dependent mechanism.
a, Serum and b, liver cytokine and chemokine profiling of female and male hosts bearing MB49 tumors with or without liver-specific deletion of Atg7 (n = 6 mice each), showing those with significant differences among 26. c, Experimental design to induce liver-specific Fip200 deletion (liver Fip200Δ/Δ) and wild-type controls (liver Fip200+/+). Fip200flox/flox or C57BL/6J mice were injected with AAV-TBG-iCre and MB49 cells were then injected subcutaneously and tumor growth was monitored over three weeks. d, Cropped Western blotting showing expression of Fip200 in livers, kidneys and brains (n = 3 each) from one experiment. e, Comparison of MB49 tumor growth on male liver Fip200+/+(n = 8 tumors) and liver Fip200Δ/Δ (n = 6 tumors) hosts. f, Experimental design to induce conditional Fip200 deletion (Fip200Δ/Δ) and wild-type (Fip200+/+) controls. C57BL/6J mice were injected with lentiCRISPR scramble or Fip200 to delete Fip200 and were then injected subcutaneously with tumor cells. Tumor growth was monitored over three weeks. g, Representative surveyor assay for Fip200 gRNA screening using liver gDNA from Fip200+/+ and Fip200Δ/Δ hosts (n = 10 mice each) from one experiment. h, Representative IHC images of FIP200+ and p62+ cells in liver from Fip200+/+ and Fip200Δ/Δ hosts (n = 10 mice each). i, j, Comparison of MB49 tumor growth on female Fip200+/+ and Fip200Δ/Δ hosts with or without CD4/8 depletion (n = 10 tumors each). Tumor pictures (i), and tumor weight and ratio (antibody treated/control) (j) from the experimental endpoint. Data are mean +/- s.e.m. *P < 0.05, **P < 0.01, ***P<0.001, ****P < 0.0001 using two-sided Student’s t-test. Source data available for a, b, e, j.
Supplementary information
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Extended Data Fig. 4
Unprocessed western blots and gels
Source Extended Data Fig. 5
Unprocessed western blots and gels
Rights and permissions
About this article
Cite this article
Poillet-Perez, L., Sharp, D.W., Yang, Y. et al. Autophagy promotes growth of tumors with high mutational burden by inhibiting a T-cell immune response. Nat Cancer 1, 923–934 (2020). https://doi.org/10.1038/s43018-020-00110-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00110-7
This article is cited by
-
Cancer cell metabolism and antitumour immunity
Nature Reviews Immunology (2024)
-
Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology
Nature Reviews Molecular Cell Biology (2024)
-
Autophagy and nuclear morphometry are associated with histopathologic features in esophageal squamous cell carcinoma
Journal of Molecular Medicine (2024)
-
Exploiting autophagy balance in T and NK cells as a new strategy to implement adoptive cell therapies
Molecular Cancer (2023)
-
Autophagy and autophagy-related pathways in cancer
Nature Reviews Molecular Cell Biology (2023)