Abstract
Lack of tumor infiltration by immune cells is the main mechanism of primary resistance to programmed cell death protein 1 (PD-1) blockade therapies for cancer. It has been postulated that cancer cell-intrinsic mechanisms may actively exclude T cells from tumors, suggesting that the finding of actionable molecules that could be inhibited to increase T cell infiltration may synergize with checkpoint inhibitor immunotherapy. Here, we show that p21-activated kinase 4 (PAK4) is enriched in non-responding tumor biopsies with low T cell and dendritic cell infiltration. In mouse models, genetic deletion of PAK4 increased T cell infiltration and reversed resistance to PD-1 blockade in a CD8 T cell-dependent manner. Furthermore, combination of anti-PD-1 with the PAK4 inhibitor KPT-9274 improved anti-tumor response compared with anti-PD-1 alone. Therefore, high PAK4 expression is correlated with low T cell and dendritic cell infiltration and a lack of response to PD-1 blockade, which could be reversed with PAK4 inhibition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-Seq data supporting the findings of this study have been deposited in the National Center for Biotechnology Information database of Genotypes and Phenotypes (https://www.ncbi.nlm.nih.gov/gap/) with accession number phs001919. The data for the pan-cancer correlation analysis were derived from the TCGA Research Network (http://cancergenome.nih.gov/). Source data on unprocessed blots in Fig. 3 and Extended Data Figs. 3–5 and 7, as well as numerical raw data for Figs. 3, 5 and 7 and Extended Data Figs. 4–6 are provided with the paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Change history
22 January 2020
A Correction to this paper has been published: https://doi.org/10.1038/s43018-020-0025-7
References
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Chen, P. L. et al. Analysis of immune signatures in longitudinal tumor samples yields insight into biomarkers of response and mechanisms of resistance to immune checkpoint blockade. Cancer Discov. 6, 827–837 (2016).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e15 (2017).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997.e24 (2018).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Liu, C. et al. BRAF inhibition increases tumor infiltration by T cells and enhances the antitumor activity of adoptive immunotherapy in mice. Clin. Cancer Res. 19, 393–403 (2013).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Spranger, S. & Gajewski, T. F. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat. Rev. Cancer 18, 139–147 (2018).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Nsengimana, J. et al. β-Catenin-mediated immune evasion pathway frequently operates in primary cutaneous melanomas. J. Clin. Invest. 128, 2048–2063 (2018).
Grasso, C. S. et al. Genetic mechanisms of immune evasion in colorectal cancer. Cancer Discov. 8, 730–749 (2018).
Luke, J. J., Bao, R., Sweis, R. F., Spranger, S. & Gajewski, T. F. WNT/β-catenin pathway activation correlates with immune exclusion across human cancers. Clin. Cancer Res. 25, 3074–3083 (2019).
Li, Y. et al. Nucleo-cytoplasmic shuttling of PAK4 modulates β-catenin intracellular translocation and signaling. Biochim. Biophys. Acta 1823, 465–475 (2012).
Rane, C. K. & Minden, A. P21 activated kinase signaling in cancer. Semin. Cancer Biol. 54, 40–49 (2019).
Yun, C. Y. et al. p21-activated kinase 4 critically regulates melanogenesis via activation of the CREB/MITF and β-catenin/MITF pathways. J. Invest. Dermatol. 135, 1385–1394 (2015).
Vershinin, Z., Feldman, M., Chen, A. & Levy, D. PAK4 methylation by SETD6 promotes the activation of the Wnt/β-catenin pathway. J. Biol. Chem. 291, 6786–6795 (2016).
Moll, R., Divo, M. & Langbein, L. The human keratins: biology and pathology. Histochem. Cell Biol. 129, 705–733 (2008).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Radu, M., Semenova, G., Kosoff, R. & Chernoff, J. PAK signalling during the development and progression of cancer. Nat. Rev. Cancer 14, 13–25 (2014).
Chen, S. et al. Wnt-1 signaling inhibits apoptosis by activating β-catenin/T cell factor-mediated transcription. J. Cell Biol. 152, 87–96 (2001).
Li, J. et al. LATS2 suppresses oncogenic Wnt signaling by disrupting β-catenin/BCL9 interaction. Cell Rep. 5, 1650–1663 (2013).
Aboukameel, A. et al. Novel p21-activated kinase 4 (PAK4) allosteric modulators overcome drug resistance and stemness in pancreatic ductal adenocarcinoma. Mol. Cancer Ther. 16, 76–87 (2017).
Takao, S. et al. Targeting the vulnerability to NAD+ depletion in B-cell acute lymphoblastic leukemia. Leukemia 32, 616–625 (2018).
Abu Aboud, O. et al. Dual and specific inhibition of NAMPT and PAK4 by KPT-9274 decreases kidney cancer growth. Mol. Cancer Ther. 15, 2119–2129 (2016).
Rane, C. et al. A novel orally bioavailable compound KPT-9274 inhibits PAK4, and blocks triple negative breast cancer tumor growth. Sci. Rep. 7, 42555 (2017).
Wang, K. et al. Inhibition of p21 activated kinase enhances tumour immune response and sensitizes pancreatic cancer to gemcitabine. Int. J. Oncol. 52, 261–269 (2018).
Mosely, S. I. et al. Rational selection of syngeneic preclinical tumor models for immunotherapeutic drug discovery. Cancer Immunol. Res. 5, 29–41 (2017).
Ueha, S. et al. Robust antitumor effects of combined anti-CD4-depleting antibody and anti-PD-1/PD-L1 immune checkpoint antibody treatment in mice. Cancer Immunol. Res. 3, 631–640 (2015).
Homet Moreno, B. et al. Response to programmed cell death-1 blockade in a murine melanoma syngeneic model requires costimulation, CD4, and CD8 T cells. Cancer Immunol. Res. 4, 845–857 (2016).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Li, X. & Minden, A. PAK4 functions in tumor necrosis factor (TNF) α-induced survival pathways by facilitating TRADD binding to the TNF receptor. J. Biol. Chem. 280, 41192–41200 (2005).
Li, Q. et al. p21-activated kinase 4 as a switch between caspase-8 apoptosis and NF-κB survival signals in response to TNF-α in hepatocarcinoma cells. Biochem. Biophys. Res. Commun. 503, 3003–3010 (2018).
Dunn, G. P., Old, L. J. & Schreiber, R. D. The three Es of cancer immunoediting. Annu. Rev. Immunol. 22, 329–360 (2004).
Nicholas, N. S. et al. PAK4 suppresses PDZ-RhoGEF activity to drive invadopodia maturation in melanoma cells. Oncotarget 7, 70881–70897 (2016).
Chen, S. et al. Copy number alterations in pancreatic cancer identify recurrent PAK4 amplification. Cancer Biol. Ther. 7, 1793–1802 (2008).
Wells, C. M., Whale, A. D., Parsons, M., Masters, J. R. & Jones, G. E. PAK4: a pluripotent kinase that regulates prostate cancer cell adhesion. J. Cell Sci. 123, 1663–1673 (2010).
Bi, Y. et al. Study on the expression of PAK4 and P54 protein in breast cancer. World J. Surg. Oncol. 14, 160 (2016).
Rane, C. K. et al. Decrypting the PAK4 transcriptome profile in mammary tumor forming cells using next generation sequencing. Genomics 110, 248–256 (2018).
Wong, L. E., Chen, N., Karantza, V. & Minden, A. The Pak4 protein kinase is required for oncogenic transformation of MDA-MB-231 breast cancer cells. Oncogenesis 2, e50 (2013).
Kim, H., Woo, D. J., Kim, S. Y. & Yang, E. G. p21-activated kinase 4 regulates HIF-1α translation in cancer cells. Biochem. Biophys. Res. Commun. 486, 270–276 (2017).
King, H. et al. PAK4 interacts with p85 alpha: implications for pancreatic cancer cell migration. Sci. Rep. 7, 42575 (2017).
Fu, X. et al. PAK4 confers cisplatin resistance in gastric cancer cells via PI3K/Akt- and MEK/ERK-dependent pathways. Biosci. Rep. 34, e00094 (2014).
He, L. F. et al. Activated-PAK4 predicts worse prognosis in breast cancer and promotes tumorigenesis through activation of PI3K/AKT signaling. Oncotarget 8, 17573–17585 (2017).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Escuin-Ordinas, H. et al. COX-2 inhibition prevents the appearance of cutaneous squamous cell carcinomas accelerated by BRAF inhibitors. Mol. Oncol. 8, 250–260 (2014).
Wei, S. C. et al. Distinct cellular mechanisms underlie anti-CTLA-4 and anti-PD-1 checkpoint blockade. Cell 170, 1120–1133 e1117 (2017).
Chen, H. et al. Cytofkit: a bioconductor package for an integrated mass cytometry data analysis pipeline. PLoS Comput. Biol. 12, e1005112 (2016).
Acknowledgements
This study was funded in part by the Parker Institute for Cancer Immunotherapy, NIH grants R35 CA197633 and P01 CA168585, the Ressler Family Foundation, and support from K. Schultz and D. Schultz (to A.R.). G.A.-R. was supported by the Isabel and Harvey Kibel Fellowship award and Alan Ghitis Fellowship Award for Melanoma Research. D.Y.T. was supported by a Young Investigator Award from the American Society of Clinical Oncology, a grant from the Spanish Society of Medical Oncology for Translational Research in Reference Centers and the V Foundation–Gil Nickel Family Endowed Fellowship in Melanoma Research. J.M.Z. was part of the UCLA Medical Scientist Training Program supported by NIH training grant GM08042. T.S.N. was supported by NIH/NICHD grant K12-HD000850 (Pediatric Scientist Development Program). S.H.-L. was supported by a Young Investigator Award and a Career Development Award from the American Society of Clinical Oncology, a Tower Cancer Research Foundation Grant and a Dr. Charles A. Coltman Fellowship Award from the Hope Foundation. C.-Y.W. was supported by NIH/NIDCR grant R01DE15964. We acknowledge X. Li, L. Dong, J. Yoshizawa and J. Zhou from the UCLA Clinical Microarray Core for sequencing expertise, and J. Min Chen and J. Trent from the Parker Institute for Cancer Immunotherapy Center at UCLA for administrative support. Flow and mass cytometry were performed in the UCLA Jonsson Comprehensive Cancer Center and the Center for AIDS Research Flow Cytometry Core Facility (supported by NIH awards P30 CA016042 and 5P30 AI028697), as well as by the Jonsson Comprehensive Cancer Center, UCLA AIDS Institute and David Geffen School of Medicine at UCLA. The authors thank A. Minden from Rutgers, the State University of New Jersey, for helpful comments.
Author information
Authors and Affiliations
Contributions
G.A.-R., C.S.G. and A.R. conceived and designed the study. G.A.-R., D.Y.T., W.L., J.M.Z., C.P.-S., T.S.N., A.K., A.J.G., G.C.-L., B.C.-A., S.H.-L., C.-Y.W., C.S.G. and A.R. developed the methodology. B.B.-M., I.B.C., S.H.-L., C-Y.W. and A.R. acquired the data (provided animals, acquired and managed patients, provided facilities, and so on). G.A.-R., D.Y.T., W.L., J.T., E.M., M.J.Q., W.S., E.B., B.C.-A., C-Y.W., C.S.G. and A.R. analyzed and interpreted the data (including statistical analysis, biostatistics and computational analysis). G.A.-R. and A.R. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
G.A.-R. has received honoraria for consulting with Arcus Biosciences. W.S. and E.B. were employees of Karyopharm Therapeutics when this study was conducted. A.R. has received honoraria for consulting with Amgen, Bristol-Myers Squibb, Chugai, Genentech, Merck, Novartis, Roche and Sanofi, is or has been a member of the scientific advisory board, and holds stock in Advaxis, Arcus Biosciences, Bioncotech Therapeutics, Compugen, CytomX, Five Prime, FLX Bio, ImaginAb, IsoPlexis, Gilead Kite, Lutris Pharma, Merus, PACT Pharma, Rgenix and Tango Therapeutics. G.A.-R., D.Y.T., C.S.G. and A.R. are inventors in a patent application covering the use of PAK4 inhibitors for cancer immunotherapy.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
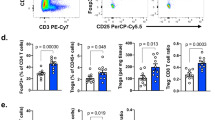
Extended Data Fig. 1 Differential change in immune populations between non-responding and responding biopsies during anti-PD-1 therapy.
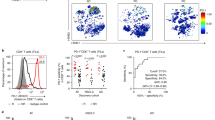
Comparison (two-sided, paired T-test) of each of the immune populations and immune markers between baseline and on-treatment tumour samples for responding (n= 5) and non-responding (n= 6) biopsies. From left to right: T cell score (R P= 0.007, NR P= 0.44), Dendritic cell score (R P= 0.009, NR P= 0.08), CD8 T cell score (R P= 0.006, NR P= 0.48), CTL score (R P= 0.01, NR P= 0.43), NK cell score (R P= 0.006, NR P= 0.32), Monocyte lineage score (R P= 0.004, NR P= 0.48), IFNg (R P= 0.01, NR P= 0.47), TNF (R P= 0.01, NR P= 0.9), GZMA (R P= 0.01, NR P= 0.73), PRF1 (R P= 0.004, NR P= 0.29) and CD8A (R P= 0.004, NR P= 0.52) expression. Increase in all immune populations and markers was significant (P < 0.05) only in responding biopsies. *P <0.05, **P < 0.01; ns, not significant
Extended Data Fig. 2 PAK4 expression analysis with immune infiltration and overlap with S100 and β-catenin staining.
a, Comparison of exclusion up Jerby-Arnon score expression (P = 3.28e-05) between tumour biopsies within the upper (n = 15) and lower (n = 15) quartile of PAK4 expression. b, PAK4 correlation with Jerby-Arnon score expression (n = 60) (R = 0.65, P = 1.78e-08). Exclusion up Jerby-Arnon was obtained based on the geometric mean of the 302 gene from Jerby-Arnon et al. c, CD8A (R = -0.39, P = 6.07e-05), TNF (R = -0.49, P = 1.89e-07), GZMA (R = -0.45, P = 2.47e-06), PRF1 (R = -0.28, P = 4e-03) and the different immune populations assessed using MCP-Counter: T cells (R = -0.39, P = 4.41e-05), CD8 T cells (R = -0.36, P = 1.71e-04), cytotoxic lymphocytes (R = -0.28, P = 4.9e-03) and dendritic cells (R = -0.57, P = 3.95e-10). n= 99 for all plots. d, Quantification of PAK4 positive cells out of S100 total positive cells. PT0158_tx2 and PT0112_tx are two biopsies with low T cell infiltration and high PAK4 expression while PT0294_tx2 and PT0349_tx have low PAK4 and high T cell infiltrate as determined by RNAseq. e, Quantification of PAK4 positive cells out of β-catenin total positive cells. From top to bottom box-plots define the maximum, 3rd quartile, median, 1st quartile and minimum values a. P values obtained using two-sided Welch’s t-test a. Correlations were calculated applying Pearson’s correlation coefficient test b, c
Extended Data Fig. 3 Validation of the generation of a PAK4 KO B16 cell line.
a, b, c TIDE analysis of the B16 PAK4 KO clones: 6.2, 8.1 and 8.2 respectively. d, e, Analysis of PAK4 protein expression in the three B16 PAK4 KO clones, B16 WT CRISPR control and rescue cell lines by Western blot. Results are representative from three independent experiments. Unprocessed blot images are provided as a Source Data file d, e.
Extended Data Fig. 4 PAK4 depletion impact on nuclear protein β-catenin and WNT signalling activity.
a, Negative control for the Topflash experiment using the Fopflash luciferase vector which contains a mutated version of the TCF/LEF binding motifs. There are no changes in Fopflash activity upon stimulation with Wnt-3a ligand for 8 hours in any of the tested cell lines (n= 3 per group) (P > 0.05 for all comparisons). b, Baseline WNT activity levels assessed by Topflash assay (n= 3 per group). Values were normalized to B16 WT CC cell lines and no significant WNT activity changes were observed between PAK4 WT and KO cell. c, Immunoblots for nuclear β-catenin protein levels show no differences between B16 WT CRISPR control, PAK4 KO and PAK4 rescue cells. Results are representative from three independent experiments. Means +/- SEM two-tailed unpaired t-test a, b. Unprocessed blot images and raw data are provided as a Source Data file a-c.
Extended Data Fig. 5 PAK4 inhibition disrupts WNT signalling and melanogenesis.
a, Cells were cultured with 2µM KPT-9274 for 72 hours before nuclear protein isolation. Showing immunoblots for nuclear β-catenin, nuclear phosphor-β-catenin (S675) and nuclear PAK4 protein levels. Results are representative from two independent experiments. b, Cells were cultured with 2µM KPT-9274 for 72 hours and Wnt-3a for 8 hours prior to Topflash assay (n= 3 per group). Pharmacological inhibition of PAK4 significantly decreases sensitivity to Wnt-3a stimulation (P= 0.005 for WT Wnt3a vs WT KPT-9274 + Wnt3a comparison). c, Baseline WNT activity levels assessed by Topflash assay of cell treated with 2µM KPT-9274 for 72 hours (n= 3 per group) (P > 0.05). Values were normalized to untreated B16 WT CC cells. d, RT-PCR for tyrosinase expression show that PAK4 depletion reduces the expression levels of this gene. Showing means +/- SEM. Results are normalized to B16 WT CRISPR control levels and then log2 transformed (n= 3). e, For image, cells were cultured and harvest upon reaching 80% confluency. B16 WT CRISPR Control cell line maintains melanin production over time while PAK4 KO clones lose their pigmentation. Results are representative from three independent experiments. Means +/- SEM two-tailed unpaired t-test b, c. Unprocessed blots and raw data are provided as a Source Data file a-c.
Extended Data Fig. 6 In vivo experiments with additional B16 PAK4 KO and rescue clones and CD8 depletion validation.
a, Tumour growth curves for B16 PAK4 KO 8.1 tumours treated with isotype (blue, n = 10) or anti-PD-1 (red, n = 12) (P= 0.00024, day 15). b, Tumour growth curves for B16 PAK4 KO 8.2 tumours treated with isotype (blue, n = 10) or anti-PD-1 (red, n = 10) (P= 0.02, day 15). In both PAK4 KO cell lines anti-PD-1 treated tumours showed decreased tumour growth compared to untreated tumours. c, Tumour growth curves for B16 8.1 PAK4 rescue tumours treated with isotype (blue, n = 5) or anti-PD-1 (red, n = 5). Anti-PD-1 treatment did not result in any significant anti-tumour efficacy (P= 0.80, day 15). d, Flow cytometry analysis of CD8 positive splenocytes after CD8 depletion. Left panel show splenocytes pattern without anti-CD8 treatment (CD8 population = 18.9%) while middle and right panel show splenocytes derived from two independent mice treated with anti-CD8 antibody (CD8 population = 0.77% and 0.50% respectively). Plotting the mean +/- s.e.m a-c. Statistical significance and correction for multiple comparisons was calculated using Holm-Sidak method a-c. Raw data is provided as a Source Data file a-c. *P <0.05, **P < 0.01, ***P <0.001, ****P < 0.0001. ns, not significant.
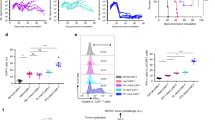
Extended Data Fig. 7 PAK4 KO validation and sensitivity to TNF in MC38 cells.
a, TIDE analysis of the MC38 PAK4 KO 6.9 clone. b, Analysis of PAK4 protein expression in MC38 PAK4 KO 6.9 clone and MC38 WT by Western blot. Results are representative from two independent experiments. c, Cells were plated by triplicate into 96 well plates and then treated with TNF at 100ng/mL. Cell proliferation was measured by cell confluence using the IncuCyte S3 Live Cell Analysis System. TNF treatment decreased proliferation of MC38 WT, MC38 PAK4 KO 6.9 and MC38 PAK4 KO 6.10 cells by 41%, 95% and 74% respectively compared to untreated cells (means +/- SEM). Results are representative from three biologically independent experiments. Unprocessed blots are provided as a Source Data file b.
Supplementary information
Source data
Source Data Fig. 3
Unprocessed Western Blots for Fig. 3f
Source Data Fig. 3
Raw data for d, e.
Source Data Fig. 5
Raw data for a-c, f, g.
Source Data Fig. 7
Raw data for a-c
Source Data Extended Data Fig. 3
Unprocessed Western Blots for Extended Data Fig. 3d, e
Source Data Extended Data Fig. 4
Unprocessed Western Blots for Extended Data Fig. 4c
Source Data Extended Data Fig. 4
Raw data for 4a, b
Source Data Extended Data Fig. 5
Unprocessed Western Blots for Extended Data Fig. 5a
Source Data Extended Data Fig. 5
Raw data for b, c
Source Data Extended Data Fig. 6
Raw data for a-c
Source Data Extended Data Fig. 7
Unprocessed Western Blots for Extended Data Fig. 7b
Rights and permissions
About this article
Cite this article
Abril-Rodriguez, G., Torrejon, D.Y., Liu, W. et al. PAK4 inhibition improves PD-1 blockade immunotherapy. Nat Cancer 1, 46–58 (2020). https://doi.org/10.1038/s43018-019-0003-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-019-0003-0
This article is cited by
-
Systematic investigation of chemo-immunotherapy synergism to shift anti-PD-1 resistance in cancer
Nature Communications (2024)
-
Mi-2β promotes immune evasion in melanoma by activating EZH2 methylation
Nature Communications (2024)
-
CAR-T cell therapy targeting surface expression of TYRP1 to treat cutaneous and rare melanoma subtypes
Nature Communications (2024)
-
Engineered exosomes from different sources for cancer-targeted therapy
Signal Transduction and Targeted Therapy (2023)
-
Genetic alterations shape innate immune cells to foster immunosuppression and cancer immunotherapy resistance
Clinical and Experimental Medicine (2023)