Abstract
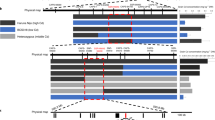
Global contamination of soils with toxic cadmium (Cd) is a serious health threat. Here we found that a tandem duplication of a gene encoding a manganese/Cd transporter, OsNramp5, was responsible for low-Cd accumulation in Pokkali, an old rice cultivar. This duplication doubled the expression of OsNramp5 gene but did not alter its spatial expression pattern and cellular localization. Higher expression of OsNramp5 increased uptake of Cd and Mn into the root cells but decreased Cd release to the xylem. Introgression of this allele into Koshihikari, an elite rice cultivar, through backcrossing significantly reduced Cd accumulation in the grain when cultivated in soil heavily contaminated with Cd but did not affect both grain yield and eating quality. This study not only reveals the molecular mechanism underlying low-Cd accumulation but also provides a useful target for breeding rice cultivars with low-Cd accumulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
Code availability
Sequence data of this article have been registered in the DDBJ/GenBank/EMBL databases under the accession numbers LC717507 for duplicated OsNramp5 in Pokkali.
References
Ma, J. F., Shen, R. F. & Shao, J. F. Transport of cadmium from soil to grain in cereal crops: a review. Pedosphere 31, 3–10 (2021).
Zhao, F. J. & Wang, P. Arsenic and cadmium accumulation in rice and mitigation strategies. Plant Soil 446, 1–21 (2020).
Horiguchi, H. et al. Hypoproduction of erythropoietin contributes to anemia in chronic cadmium intoxication: clinical study on Itai-itai disease in Japan. Arch. Toxicol. 68, 632–636 (1994).
Song, Y. et al. Dietary cadmium exposure assessment among the Chinese population. PLoS ONE 12, e0177978 (2017).
Sui, F. et al. Nramp5 expression and functionality likely explain higher cadmium uptake in rice than in wheat and maize. Plant Soil 433, 377–389 (2018).
Sasaki, A., Yamaji, N., Yokosho, K. & Ma, J. F. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. Plant Cell 24, 2155–2167 (2012).
Ishikawa, S. et al. Ion-beam irradiation, gene identification, and marker-assisted breeding in the development of low-cadmium rice. Proc. Natl Acad. Sci. USA 109, 19166–19171 (2012).
Yang, M. et al. OsNRAMP5 contributes to manganese translocation and distribution in rice shoots. J. Exp. Bot. 65, 4849–4861 (2014).
Yan, H. et al. Variation of a major facilitator superfamily gene contributes to differential cadmium accumulation between rice subspecies. Nat. Commun. 10, 2562 (2019).
Chang, J. D. et al. OsNRAMP1 transporter contributes to cadmium and manganese uptake in rice. Plant Cell Environ. 43, 2476–2491 (2020).
Tan, L. et al. ZINC TRANSPORTER5 and ZINC TRANSPORTER9 function synergistically in zinc/cadmium uptake. Plant Physiol. 183, 1235–1249 (2020).
Ueno, D. et al. Gene limiting cadmium accumulation in rice. Proc. Natl Acad. Sci. USA 107, 16500–16505 (2010).
Miyadate, H. et al. OsHMA3, a P1B-type of ATPase affects root-to-shoot cadmium translocation in rice by mediating efflux into vacuoles. New Phytol. 189, 190–199 (2011).
Takahashi, R. et al. The OsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice. Plant Cell Environ. 35, 1948–1957 (2012).
Yamaji, N., Xia, J., Mitani-Ueno, N., Yokosho, K. & Ma, J. F. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2. Plant Physiol. 162, 927–939 (2013).
Uraguchi, S. et al. Low-affinity cation transporter (OsLCT1) regulates cadmium transport into rice grains. Proc. Natl Acad. Sci. USA 108, 20959–20964 (2011).
Hao, X. et al. A node-expressed transporter OsCCX2 is involved in grain cadmium accumulation of rice. Front. Plant Sci. 9, 476 (2018).
Lu, C. et al. Producing cadmium-free Indica rice by overexpressing OsHMA3. Environ. Int. 126, 619–626 (2019).
Arao, T. & Ae, N. Genotypic variations in cadmium levels of rice grain. Soil Sci. Plant Nutr. 49, 473–479 (2003).
Ueno, D. et al. A major quantitative trait locus controlling cadmium translocation in rice (Oryza sativa). New Phytol. 182, 644–653 (2009).
Liu, C. L. et al. Natural variation in the promoter of OsHMA3 contributes to differential grain cadmium accumulation between Indica and Japonica rice. J. Integr. Plant Biol. 62, 314–329 (2020).
Ebana, K., Kojima, Y., Fukuoka, S., Nagamine, T. & Kawase, M. Development of mini core collection of Japanese rice landrace. Breed. Sci. 58, 281–291 (2008).
Yu, E., Yamaji, N., Mao, C., Wang, H. & Ma, J. F. Lateral roots but not root hairs contribute to high uptake of manganese and cadmium in rice. J. Exp. Bot. 72, 7219–7228 (2021).
Yang, C., Zhang, Y. & Huang, C. Reduction in cadmium accumulation in japonica rice grains by CRISPR/Cas9-mediated editing of OsNRAMP5. J. Integr. Agr. 18, 688–697 (2019).
Chang, J. D. et al. Overexpression of the manganese/cadmium transporter OsNRAMP5 reduces cadmium accumulation in rice grain. J. Exp. Bot. 71, 5705–5715 (2020).
Hoshikawa, K. The Growing Rice Plant (Nosan Gyoson Bunka Kyokai, 1989).
Huang, S., Wang, P., Yamaji, N. & Ma, J. F. Plant nutrition for human nutrition: hints from rice research and future perspectives. Mol. Plant 13, 825–835 (2020).
Ma, J. F. et al. A silicon transporter in rice. Nature 440, 688–691 (2006).
Ma, J. F. et al. An efflux transporter of silicon in rice. Nature 448, 209–212 (2007).
Huang, S. et al. Boron uptake in rice is regulated post-translationally via a clathrin-independent pathway. Plant Physiol. 188, 1649–1664 (2022).
Ueno, D. et al. A polarly localized transporter for efficient manganese uptake in rice. Nat. Plants 1, 15170 (2015).
Zou, C., Lehti-Shiu, M. D., Thomashow, M. & Shiu, S. H. Evolution of stress-regulated gene expression in duplicate genes of Arabidopsis thaliana. PLoS Genet. 5, e1000581 (2009).
Kondrashov, F. A. Gene duplication as a mechanism of genomic adaptation to a changing environment. Proc. R. Soc. B 279, 5048–5057 (2012).
Chow, E. W. L., Morrow, C. A., Djordjevic, J. T., Wood, I. A. & Fraser, J. A. Microevolution of Cryptococcus neoformans driven by massive tandem gene amplification. Mol. Biol. Evol. 29, 1987–2000 (2012).
Sutton, T. et al. Boron-toxicity tolerance in barley arising from efflux transporter amplification. Science 318, 1446–1449 (2007).
Nadeem, F. et al. Comparative response of two rice (Oryza sativa L.) cultivars to applied zinc and manganese for mitigation of salt stress. J. Soil Sci. Plant Nutr. 20, 2059–2072 (2020).
Ma, J. F., Tamai, K., Ichii, M. & Wu, G. F. A rice mutant defective in Si uptake. Plant Physiol. 130, 2111–2117 (2002).
Yonemaru, J., Ebana, K. & Yano, M. HapRice, an SNP haplotype database and a web tool for rice. Plant Cell Physiol. 55, e9 (2014).
Wang, S., Basten, C. J. & Zeng, Z. B. Windows QTL Cartographer 2.5 (North Carolina State Univ., 2012).
Hiei, Y. & Komari, T. Agrobacterium-mediated transformation of rice using immature embryos or calli induced from mature seed. Nat. Protoc. 3, 824 (2008).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Yamaji, N. & Ma, J. F. Spatial distribution and temporal variation of the rice silicon transporter Lsi1. Plant Physiol. 143, 1306–1313 (2007).
Yamaji, N., Sasaki, A., Xia, J. X., Yokosho, K. & Ma, J. F. A node-based switch for preferential distribution of manganese in rice. Nat. Commun. 4, 2442 (2013).
Tamai, K. & Ma, J. F. Reexamination of silicon effects on rice growth and production under field conditions using a low silicon mutant. Plant Soil 307, 21–27 (2008).
Iijima, K. et al. Endosperm enzyme activity is responsible for texture and eating quality of cooked rice grains in Japanese cultivars. Biosci. Biotech. Bioch. 83, 502–510 (2019).
Yamaji, N. & Ma, J. F. Bioimaging of multiple elements by high-resolution LAICP-MS reveals altered distribution of mineral elements in the nodes of rice mutants. Plant J. 99, 1254–1263 (2019).
Suzuki, T. et al. iQuant2: software for rapid and quantitative imaging using laser ablation-ICP mass spectrometry. Mass Spectrom. 7, A0065 (2018).
Acknowledgements
This work was supported by Japan Society for the Promotion of Science (JSPS) (KAKENHI grant numbers 16H06296 and 21H05034 to J.F.M.) and the Key Project of the National Natural Science Foundation of China (number 42020104004 to R.F.S.). We thank A. Morita, Y. Takahashi, M. Hikasa, Y. Murasawa and S. Rikiishi for their experimental assistance.
Author information
Authors and Affiliations
Contributions
J.F.M. designed the research. E.Y., W.W., N.Y., S.F., J.C., D.U., T.A., F.D., K.H., M.Y., R.F.S. and J.F.M. performed the experiment. E.Y., W.W., N.Y. and J.F.M. analysed the data. E.Y. and J.F.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Food thanks Steve P. McGrath, Sébastien Thomine and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1–5.
Source data
Source Data Fig. 1
Original data for Fig. 1.
Source Data Fig. 2
Original data for Fig. 2.
Source Data Fig. 4
Original data for Fig. 4.
Source Data Fig. 5
Original data for Fig. 5.
Source Data Fig. 6
Original data for Fig. 6.
Source Data Fig. 7
Original data for Fig. 7.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, E., Wang, W., Yamaji, N. et al. Duplication of a manganese/cadmium transporter gene reduces cadmium accumulation in rice grain. Nat Food 3, 597–607 (2022). https://doi.org/10.1038/s43016-022-00569-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43016-022-00569-w
This article is cited by
-
Knockout of OsHMA3 in an indica rice increases cadmium sensitivity and inhibits plant growth
Plant Growth Regulation (2024)
-
Does cadmium cause cascading effects on the development and reproduction of the striped stem borer, Chilo suppressalis (Walker)?
CABI Agriculture and Bioscience (2023)
-
Responses of Rice and Related Cadmium Transporter Genes to the Passivating Microbial Agent MBLHAP
Current Microbiology (2023)
-
Populus euphratica plant cadmium resistance 2 mediates Cd tolerance by root efflux of Cd ions in poplar
Plant Cell Reports (2023)
-
Keeping toxic cadmium out of the food chain
Nature Food (2022)