Abstract
The extent to which humans facilitate zoonotic transmission of infectious diseases is unclear. Human encroachment into wildlife habitats as a consequence of expanding urbanization, cropland area and intensive animal farming is hypothesized to favour the emergence of zoonotic diseases. Here we analyse comprehensive, high-resolution datasets on forest cover, cropland distribution, livestock density, human population, human settlements, bat species’ distribution and land-use changes in regions populated by Asian horseshoe bats (>28.5 million km2)—the species that most commonly carry severe acute respiratory syndrome (SARS)-related coronaviruses. We identify areas at risk of SARS-related coronavirus outbreaks, showing that areas in China populated by horseshoe bats exhibit higher forest fragmentation and concentrations of livestock and humans than other countries. Our findings indicate that human–livestock–wildlife interactions in China may form hotspots with the potential to increase SARS-related coronavirus transmission from animals to humans.
Similar content being viewed by others
Main
Emerging infectious diseases frequently originate from pathogen spillovers from wildlife to humans1; contributing factors include forest fragmentation, habitat destruction, agricultural expansion, concentrated livestock production and human penetration into wildlife habitats2,3,4,5. However, a quantitative analysis of the nexus between ongoing land-use changes and the emergence of new zoonotic diseases is still missing5,6.
Genomic sequencing of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has shown that the virus is closely related (~96%) to a strain present in horseshoe bats7, but it is still unclear whether the spillover of SARS-CoV-2 occurred directly from bats to humans or through an intermediate species. For example, a strain of coronavirus very similar to SARS-CoV-2 was detected in Malayan pangolin (Manis javanica)8, a wild mammal that is frequently illegally smuggled from Southeast Asia into China and sold in markets8. Regardless of the specific pathway, the pathogen flow of emerging zoonotic diseases to humans is the result of human interactions with wildlife. We argue that the increasing incidence of emerging disease outbreaks is the result of a similar set of drivers able to change the distance and contact rates between wildlife and humans (as well as human–human interaction). Population growth, urbanization, increasing affluence in middle-income countries and the associated dietary shifts9,10,11, including increased demand for animal products, are driving agriculture expansion and changes in animal husbandry—often at the expense of natural ecosystems12,13. Intensive livestock production keeps a large number of animals—often immunosuppressed, with low genetic diversity and in poor conditions—in close proximity to one another, making them vulnerable to the emergence and spread of epidemics3,14,15.
Human encroachment into wildlife habitat favours the interaction between humans and wildlife species, either directly through activities such as hunting or indirectly through other species, particularly livestock that are in closer contact with humans16,17,18,19. The establishment of pastures, plantations or intensive livestock farms close to forest margins may increase pathogen flow from wildlife to humans17,19,20,21. Deforestation and forest fragmentation reshape the dynamics of wildlife communities, possibly leading to the extinction of habitat-specialist species while allowing generalists to thrive22. Wildlife species that are hosts of pathogens have been found to be relatively more abundant in managed landscapes (for example, agro-ecosystems and urban areas) than in adjacent undisturbed sites—particularly in the case of bats and other mammal species23. Among the four coronavirus genera, two (α and β coronaviruses) are found in bats, including the SARS-related CoVs (SARSr-CoV)24,25,26,27. For example, SARS and swine acute diarrhoea syndrome coronavirus (SADS-CoV) emerged in southeast China and were later detected in horseshoe bats, mainly Rhinolophus sinicus and Rhinolophus affinis7,24. SARSr-CoVs in China are most similar to the highly pathogenic human SARS-CoVs7,24.
Links between land-use change and the emergence of the COVID-19 pandemic have been hypothesized but have yet to be supported by a comprehensive high-resolution analysis of land-use patterns that combines forest fragmentation with agricultural expansion, livestock production and human encroachment into wildlife habitats21. Here we analyse environmental and land-use changes to identify locations at risk for SARS-related coronavirus outbreaks1,21 and relate such outbreak risks to ongoing agricultural production trends and dietary shifts. We analyse a set of factors that make a location suitable for the spillover to humans to occur. To that end, while we do not specifically link environmental change or bats as the immediate hosts of the SARS-CoV-2 ancestor7,8, we use horseshoe bats in the genus Rhinolophus (family Rhinolophidae) as a model system to understand the risk of future coronavirus outbreaks because China and Southeast Asia are reported to be regions with both highly diverse horseshoe bats and bat SARS-like CoV24.
Results
Unfortunately, the location of the first infection events of SARS-like CoV in humans is not known. Therefore, we performed our local analyses at horseshoe bat locations in China (Fig. 1a and Supplementary Table 1) and within horseshoe bat distributions in both the larger region including South, East and Southeast Asia (and additionally in Western Europe and North Africa) and then in China. Within these distributions we generated 10,000 random sampling points (Fig. 1a). Within 30 km from every random sampling point we calculate livestock density (number per km2), forest cover and fragmentation, cropland cover, population density, and the fractional cover of human settlements (Supplementary Figs. 2–7). Hotspots were calculated (Fig. 1a) using the Getis–Ord algorithm to show where the areas with high or low values of land-use attributes cluster (Fig. 2).
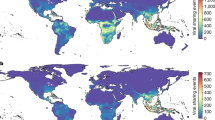
a, Sampling points randomly generated within and outside China and bat location points, weighted by the horseshoe bat species distributions present in East, South and Southeast Asia. b, Hotspots (red) and coldspots (blue) of livestock density. c, Hotspots of forest fragmentation. d, Hotspots of human settlement. Hotspots are based on the Getis–Ord \(G_i^*\) statistic and classified according to their two-tailed significance levels, corresponding to those of a standard normal distribution. Basemap adapted from IUCN Red List database (https://www.iucnredlist.org/search). Map projection EPSG:3395 WGS84/World Mercator.
a–c, Average distributions of livestock (a), land cover and use (b) and human population (c) in areas that are likely to be suitable for horseshoe bat occurrence in China and in the rest of their distribution. Error bars correspond to the 20% and 80% sample percentiles. Different capital letters indicate statistically different samples according to Mann–Whitney tests with significance α = 0.05.
Within the large region analysed, China exhibits a relatively high concentration of livestock production in horseshoe bat distributions (Fig. 1a and Supplementary Table 1). Indeed, China is a hotspot of livestock density within this region (Fig. 1b), with statistically significant higher concentrations of chickens, ducks, pigs, goats and cattle than exist outside China (Fig. 2a). Within a 30 km radius from observed bat locations the density of chicken, ducks, pigs, goats and cattle was again significantly greater than randomly selected locations outside China. Conversely the sheep density is lower in China, although sheep density was low overall, as it was for other ruminants. The density of chickens, pigs, goats and cattle surrounding (<30 km) the points where these bats were recorded and at the randomly selected locations in China within the suitability region were not significantly different, indicating that these random locations have livestock densities that are representative of the areas in which the actual presence of horseshoe bats has been documented.
Forest cover and fragmentation have been related to virus outbreaks from wildlife (including bats) for other zoonotic diseases such as Ebola virus disease2. China exhibits on average lower forest cover and cropland density and greater forest fragmentation than the other regions analysed (Fig. 1c). The average forest cover and forest fragmentation in the surroundings (within 30 km distance) of random points selected in China and the other regions (Figs. 1a and 2b) show that these differences are statistically significant. Likewise, statistically significant differences (that is, lower average cover and higher average fragmentation) are found between the points of actual observations of horseshoe bats and randomly selected locations in the regions outside China within the distributions of these bats (Fig. 2b).
China also exhibits higher levels of human presence in horseshoe bat distributions, as evidenced by population density and the fraction of the landscape covered by villages, towns and other human settlements (Fig. 2c). Indeed, the region of China suitable for horseshoe bats coincides with hotspots of human settlements (Fig. 1d). Collectively, these results demonstrate that China exhibits stronger signs of human encroachment, livestock density and forest disturbance of SARSr-CoV-hosting horseshoe bat distributions than other regions. In China, regions close to forest fragments are more densely used for livestock production and human settlements—and consequently exhibit lower forest and cropland cover (Fig. 1b)—thereby favouring the contact between wildlife and humans either directly or through intermediate animals such as livestock. The fact that China is a global hotspot in the concurrence of these three factors (fragmentation, livestock density and human settlement) is highlighted by the multivariate hotspot analysis (Fig. 3). These three attributes account for bat habitat (fragmentation), livestock and human presence, which are major factors contributing to the spillover of zoonotic infectious diseases3. Interestingly, we find that China is the global hotspot of simultaneously high forest fragmentation, livestock density and human settlement. The other major global hotspots outside China are found in Java, Bhutan, east Nepal, northern Bangladesh, the state of Kerala (India) and northeast India, of which the Bangladeshi and northeast India regions are known for past outbreaks of Nipah virus, another bat-related zoonotic disease28.
The hotspot analysis is based on the average \(G_i^*\) z score values for fragmentation, livestock (cattle, goats, pigs, sheep) density, and human settlements. Hotspots are classified based on their two-tailed significance levels, corresponding to those of a standard normal distribution. Basemap adapted from the IUCN Red List database (https://www.iucnredlist.org/search).
We then use the multivariate hotspot framework to identify regions at high potential risk of SARSr-CoVs spillovers to humans as a result of land-use change. To that end the results of the multivariate hotspot analysis were clustered into 30 groups, based on geographic contiguity and similarity in the above three attributes (Fig. 4 and Supplementary Table 3). We then perturbed one attribute at a time in each group to evaluate that group’s susceptibility to transitioning from non-significant conditions (Fig. 3) to a hotspot state (Fig. 5). This sensitivity analysis (Fig. 5) shows areas at risk of transitioning to hotspots as a result of a future increase in at least one of the analysed attributes (that is, forest fragmentation, livestock density or human settlement). Interestingly the Chinese region south of Shanghai is at high risk of potentially turning into a hotspot as a result of fragmentation increase. Other regions susceptible to hotspot transition as a result of forest fragmentation include Japan and north Philippines. Likewise, the transition region between China’s hotspot and Indochina’s coldspot and the region surrounding the hotspot of Thailand could turn into hotspots for SARSr-CoV spillover as a result of increasing presence of livestock or humans, respectively (Fig. 5). These results point both to regions of the world currently suitable for SARSr-CoV spillover from wildlife to humans as well as those at risk of becoming prone to spillover as a result of trajectories of land-use change and human penetration (Fig. 6 and Supplementary Fig. 8)
Group naming is indicative and the colour code has purely graphic purposes. To see the complete characterization of all 30 groups, please refer to Supplementary Table 3 and Supplementary Fig. 8. Basemap adapted from the IUCN Red List database (https://www.iucnredlist.org/search).
Light blue and red dots represent, respectively, multivariate coldspots and hotspots, whereas grey dots are neutral and not sensitive to perturbation in the attributes. Hotspots/coldspots are classified according to the two-tailed 95% significance levels of the \(G_i^*\) z score, corresponding to those of a standard normal distribution. Basemap adapted from the IUCN Red List database (https://www.iucnredlist.org/search).
Discussion
Food systems are often related to human health via the impact of unhealthy diets on the emergence of chronic diseases. However, they can also affect human health more indirectly, through land-use changes induced by the increasing demand for food commodities such as meat or other animal products, a phenomenon known as the ‘livestock revolution’13. In many regions of the world livestock production growth has often led to agricultural expansion, forest destruction and the encroachment of cropland and intensive livestock farms into disturbed wildlife habitat21. This study connected the dots between the risk of SARSr-CoV epidemics and land-use changes resulting from the increase in human population and intensive farming and from agricultural expansion. Our approach uses horseshoe bats as a model family because of their key role as hosts of Sarbecovirus coronaviruses, which have caused SARS and COVID-19, and SADS7,20,22,24. Other strains of related viruses have been found in other bat genera, but these relationships are less clear24. The widespread sampling of other bats may find species-specific relationships, although horseshoe bats appear to be the reservoirs where most SARSr-CoVs have their evolutionary ancestors and so we assume they are the most appropriate models. The risk to humans from other coronaviruses, therefore, will be different, because their host distributions are different, and two CoV genera (γ and δ coronaviruses) are mostly bird viruses. Similarly, the potential intermediate or amplifying host, such as other wildlife (for example, pangolin) or livestock species might differ for different coronaviruses. Here we present the results for all livestock (except poultry), because, for example, while pigs are not reportedly susceptible to SARS-CoV29 and SARS-CoV-230, SARS-CoV and SARS-CoV-2 have sporadically naturally infected numerous different animals and been shown experimentally to be able to infect others (for example, ref. 31 and references therein) and SADS-CoV has infected pigs multiple times20,32. SADS-CoV is a coronavirus related to Rhinolophus bat coronavirus HKU2, so we include pigs in our analyses. We do not include birds (poultry) in our main analyses (Figs. 1–6 and Supplementary Figs. 8), because there is no evidence of these coronaviruses in birds, but include poultry for comparison in the Supplementary Information (Supplementary Figs. 9–13). Moreover, poultry and pigs have been associated with the spread of, for example, influenza viruses, and therefore it is also interesting to consider all livestock types.
The bat location data and species distribution data also suffer from different, but related, issues to the virus data. The bat location data are presence-only data. True absence data are difficult to obtain, and therefore we randomly sampled within different locations to generate pseudo-absence data. Choosing from where to sample from also presents difficulties, and therefore we chose horseshoe bat distribution data for species that existed within China and East, South and Southeast Asia. This presents further issues because the distribution of one species, the greater horseshoe bat (Rhinolophus ferrumequinum), encompasses Western Europe, North Africa, and Central and East Asia. We therefore weighted our sampling based on the number of overlapping species distributions to account for this. However, these species distributions are large polygons and the realized niches used within them by the species probably differ; therefore, better niche models using presence and, ideally, presence–absence data are required to develop better predictions of species presence33. However, our results for random locations in China and outside China and reported bat observations were comparable, suggesting the results were insensitive to these changes. Land-use change and climate change may change their distributions in the future. Here we limit our analyses to data reported since 2000, but future analyses may be needed to better capture location and distribution changes.
More generally, although we used relatively specific bat and virus relationships, we took a high-level approach to understand the more distal or ultimate (rather than proximal) causes of infectious disease emergence, linking environmental change and human drivers such as agricultural intensification. Different infectious diseases have different transmission mechanisms and life cycles, and not all will respond to such changes in the same way. For example, directly transmitted, acute infections with short incubation and infectious periods, such as SARSr-CoVs, will probably be dependent on hosts having greater densities, as in China, for them to emerge. The epidemic potential is also increased through local and global movement and trade, either of people, wildlife or livestock16,34,35. Along with the biological properties of the virus and hosts, the true risk of both the initial cross-species transmission and epidemic potential is either increased or limited by more proximal mechanisms, such as biosecurity, health and safety measures that can reduce risk, even if the ultimate factors are present and increasing through the processes of habitat fragmentation and human encroachment2,16.
Spillover of infectious disease such as SARS, COVID-19 and SADS from wildlife to humans probably requires the coexistence of horseshoe bats and humans in the same environment and is favoured by the presence of intermediate animal species, particularly livestock because this is in closer contact with humans. The fragmentation and disturbance of forest ecosystems probably favours habitat-generalist bat species. This study demonstrates that in China these important factors responsible for reducing the distance between wildlife and humans co-occur both in horseshoe bat distributions and in the surroundings of actual documented bat occurrence. These results are consistent with the notion that population growth and increasing meat consumption associated with urbanization and economic growth have expanded the footprint of agriculture, leading to human encroachment in wildlife habitats and increased livestock density in areas adjacent to fragmented forest patches. China has dramatically increased animal consumption36, probably as the result of increasing affluence. In China, meat supply is largely reliant on domestic production using imported feed (for example, soy from the Americas)36, which explains the high livestock density in many rural areas, including those at the forest margins. Indeed, about 94% of meat consumption is contributed by domestic production (96% for pig meat, 92% for poultry, 94% for mutton and goat, 80% for bovine meat). Likewise, economic growth and the shift to diets richer in animal products explains the increasing demand for wild animal meat delicacies, increasing human–wildlife interactions through multiple pathways and the disturbance of forest habitat in more remote locations—frequently abroad—through trade-related connections37. China is also undergoing major urbanization trends, with a ~24.6 × 103 km2 increase in built-up land between 2010 and 2015 at the expense of cropland, woodland and grassland38. At the same time, China accounts for about 25% of recent global greening trends, largely from tree plantations and forests39. Despite these greening trends, between 2000 and 2018, the increase in forest cover was only a fraction of the forest loss (Supplementary Table 5). Moreover, tree cover increase does not necessarily go hand in hand with a reversal of fragmentation trends. For reforestation to reduce forest fragmentation it would need to take place within forest fragments and lead to a more continuous tree cover. Planting trees in discontinuous land patches has the effect of increasing forest fragmentation. In fact, when the forest gain that occurred between 2000 and 2018 was accounted for, we detected an even higher degree of forest fragmentation than without considering the effect of forest gain (Supplementary Table 6 and Fig. 15). Moreover, it has been reported that in China tree planting often occurs with monocultures growing within forest fragments, thereby contributing to the persistence of fragmented habitats40.
The multivariate hotspot analysis highlights how China is the largest hotspot for the concurrence of high forest fragmentation, livestock density and human presence in our analysis (Fig. 3). The analysis does not prove any causal relationship between these land-use attributes and virus transmission to humans but highlights the existence of a remarkable co-dependence pattern among different risk factors in areas where horseshoe bats occur. While the distribution of horseshoe bats can be reshaped by climate change, the patterns identified by this study can be used to investigate the nexus between coronavirus emergence and land-use change. The sensitivity analyses identifying the possible transition to new hotspots in response to an increase in one of these attributes (Fig. 5) highlights areas that could become suitable for spillover and the type of land-use change that could induce hotspot activation. Therefore, this analysis highlights region-specific targeted interventions that are urgently needed to increase resilience to SARSr-CoV spillovers. For instance, the green dots in Fig. 4 could be turned into hotspots as a result of forest fragmentation. In these regions resilience can be built through restoration efforts. Indeed, land-use change evaluations should consider the risk of activating new hotspots suitable for wildlife-to-human spillover of pathogens such as SARSr-CoV, an aspect that has seldom been included in the impact analysis of land-use change. Likewise, other regions such as the China–Indochina transition zone or central Thailand are prone to hotspot transitioning as a result of increased livestock density or urbanization, respectively. Here, mitigation of SARSr-CoV emergence can be enhanced by reducing livestock or human density, respectively, thereby reversing ongoing dietary and urbanization trends. Thus, environmental health is tightly connected to both animal and human health, as recently stressed by planetary and ‘one health’ discourses, which advocate for more holistic views of global health, encompassing environment, animals and people, and the interactions among these factors41.
Methods
Bat location data
Most SARS-related CoVs are detected in horseshoe bats, although some strains have also been detected in other genera24,25,26,27. SARSr-CoVs in China are most similar to the highly pathogenic human SARS-CoVs24,42.
We restricted our local analyses of disturbance at bat locations to rhinolophid bats in China. We performed a Web of Science search on 10 April 2020 using the following Boolean operators: Rhinoloph* AND China AND Monitor* OR Survey OR Niche OR Distribution. We found 129 unique references. We removed all those published before 2000, reporting data outside China, review articles and non-English-language publications (specifically 23 Chinese-language publications), those with no rhinolophid data and those reporting only fossil records. We retained infection studies. This left 48 publications. We then further manually reviewed the publications for those reporting location data but more specifically those with latitude and longitude, leaving 17 publications and 264 observations (Fig. 1a and Supplementary Table 1).
Bat distribution data
We restricted our analyses of disturbance in bat distributions to rhinolophid bats in both the larger South, East and Southeast Asia region (but see the main text and ‘Results’ section) and then China. We searched the IUCN Red List database (https://www.iucnredlist.org/search) using Taxonomy: Rhinolophidae and Region: East Asia and South & South East Asia (hereinafter ‘regional’) followed by Taxonomy: Rhinolophidae and Region: East Asia: China, Hong Kong & Taiwan (hereinafter ‘Chinese’) classifications and downloaded the shapefiles for the 55 regional and 22 Chinese Rhinolophus species present in the region. We consider these areas as regions of suitable habitat for Rhinolophidae. The extent of this study area exceeds 28.5 million km2.
Within these putative species distributions, we generated 10,000 random sampling points with a local sampling density that is proportional to the number of species whose distributions were reported at the point. Horseshoe bats are largely sedentary, foraging within a few kilometres (typically 1–5 km and nearly always <10 km) of their roosts; their roosts are ~13–90 km apart and they only travel 20–60 km between winter and summer roosts (maximum recorded, 320 km)43,44. Therefore, for every random sampling point we consider a circular area of 30 km radius within which we calculate livestock density, forest cover and fragmentation, cropland cover, population density and the fractional cover of human settlements as explained below. The average values of these statistics are then calculated for China and the other regions of the world with habitat suitable for Rhinolophidae and compared and the difference is tested for significance using the Mann–Whitney non-parametric test in Mathematica.
Livestock, forest cover and population data
We took livestock data from the GeoWiki database that provides georeferenced livestock counts (in number of animals per km2) at 1 km resolution for chickens, ducks, pigs, goats, sheep and cattle (https://livestock.geo-wiki.org/home-2/)45. We quantified human presence both in terms of population density at 1 km resolution and as a fraction of the landscape taken by villages, towns or other settlements from the WorldPop database at 1 km resolution. We used cropland data (at 30 × 30 m2 resolution) from ref. 46. Forest cover data are available at 30 m resolution annually between 2000 and 201847. Forest cover is associated with the presence of trees taller than 5 m. Forest loss or gain was determined as the difference in forest cover between these two years.
Data uncertainties and consistency
The Global Livestock of the World (GLW) maps at 1 km resolution were validated by Robinson et al.45 with overall satisfactory results for our study area, with observed–simulated correlations ranging between a minimum of 0.54 for ducks in Australia to a maximum of 0.81 for ducks and pigs in Asia.
Gilbert at al.48 presented a new version of the GLW at a lower resolution (~10 km at the equator instead of 1 km) to avoid misinterpretations in local analyses. We checked the robustness of our analysis with respect to the different spatial resolution of GLW maps by computing the linear correlation between hotspot and coldspot results obtained with the two inputs (Supplementary Fig. 14). We obtained values higher than 0.95 for all species except poultry and higher than 0.80 for poultry, and chose to employ the high-resolution maps because they are better suited for the type of analysis performed in this study.
The authors of the other datasets used for our analysis reported an accuracy greater than 95% for forest data47 and 91.7% for cropland data49, ranging, for the tiles including the study area, from 88.6% in Southeast Asia to 94% in China. The framework used by WorldPop to compute the built settlement maps is explained in Nieves et al.50. Validation results show a proportion of correctly predicted transition pixels ranging from 0.79 in Vietnam to 0.997 in Switzerland.
We checked for the consistency of cropland and settlement data against the tree cover data. Forest covers 25.2% of the study area; settlements and croplands cover 7.4% and 34.2% of the study area, respectively. Overall, 95.8% of the area classified as settlements and 96.2% of the area classified as cropland fall within areas not classified as forest. The overall consistency, calculated as the unambiguously classified share of the study area, is 98.7% between the maps of tree cover and croplands and 99.7% between tree cover and settlements. More detailed results of the consistency analyses are found in Supplementary Table 4.
Forest fragmentation analyses
We performed a forest fragmentation analyses based on Vogt et al.51 using the 30 m forest cover data. This method distinguishes forest cores from forest margins and patches. Every 30 m pixel is classified as wooded or non-wooded, based on whether its tree cover was greater or smaller than 50% in the year 2018. Forest cores are wooded pixels that are not adjacent to non-wooded pixels. Conversely, forest patches are made of wooded pixels that are not adjacent to forest core pixels. Wooded pixels that are neither core nor patch pixels occur at the margins of forest cores. Forest fragmentation was then quantified in terms of a composite fragmentation index2, defined as the ratio between the sum of number of pixels classified as ‘margins’, ‘patches’ or smaller core areas (that is, <200 ha), and the total number of pixels (wooded + non-wooded) in the 30 km circles used to characterize land cover and land use in the surroundings of the points of actual bat observations or the randomly generated points. This index ranges between 0 and 1.
Hotspot analyses and multivariate clustering
We then used two different methods to generate a multivariate distribution for the three indicators (livestock density, forest fragmentation and human settlements). First, we averaged their \(G_i^*\). Since the \(G_i^*\) is a z score, that is, it has a standard normal distribution, a linear combination of the three \(G_i^*\) indicators, such as their average, is a standard normal distribution and can still be represented with the same significance levels (Fig. 2). Second, we performed a spatially constrained multivariate clustering analysis. A minimum spanning tree from the connectivity graph of the features was built, and then the SKATER (Spatial ‘K’luster Analysis by Tree Edge Removal) clustering method was used52. SKATER iteratively cuts branches in the minimum spanning tree, based on data variability among and within groups and on a spatial constraint, until it reaches the user-defined number of groups. The spatial constraint defined here is a ‘k nearest neighbours’ type with eight neighbours, meaning each feature in a group must have at least one of its eight nearest neighbours in the same group. We chose 30 as the number of groups, calculated a set of summary statistics and boxplots for the groups and compared them to their global values (Supplementary Table 2). For each indicator, we calculated the R2 value as the reduction in variance of the indicator obtained by grouping, divided by the original variance of the indicator (Supplementary Table 2). While the modularity analysis based on pseudo F-statistics shows that the optimal number of groups (the maximum differences between groups while maximizing within-group similarity) is 12, here we studied 30 groups to analyse distinct regional patterns. Having a greater number of groups allows us to identify groups that are susceptible to transitioning to a hotspot because they are not ‘too different’ from hotspots.
References
Jones, K. E. et al. Global trends in emerging infectious diseases. Nature 451, 990–993 (2008).
Rulli, M. C., Santini, M., Hayman, D. T. & D’Odorico, P. The nexus between forest fragmentation in Africa and Ebola virus disease outbreaks. Sci. Rep. 7, 41613 (2017).
Espinosa, R., Tago, D. & Treich, N. Infectious diseases and meat production. Environ. Resource Econ. 76, 1019–1044 (2020).
Young, H., Griffin, R. H., Wood, C. L. & Nunn, C. L. Does habitat disturbance increase infectious disease risk for primates? Ecol. Lett. 16, 656–663 (2013).
Gottdenker, N. L., Streicker, D. G., Faust, C. L. & Carroll, C. R. Anthropogenic land use change and infectious diseases: a review of the evidence. EcoHealth 11, 619–632 (2014).
Rohr et al. Emerging human infectious diseases and the links to global food production. Nat. Sustain. 2, 445–456 (2019).
Zhou, P. et al. A pnemonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Lam, T. T. et al. Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature 583, 282–285 (2020).
Tilman, D. & Clark, M. Global diets link environmental sustainability and human health. Nature 515, 518–522 (2014).
Hassell, J. M., Begon, M., Ward, M. J. & Fèvre, E. M. Urbanization and disease emergence: dynamics at the wildlife–livestock–human interface. Trends Ecol. Evol. 32, 55–67 (2017).
Shah, H. A., Huxley, P., Elmes, J. & Murray, K. A. Agricultural land-uses consistently exacerbate infectious disease risks in Southeast Asia. Nat. Commun. 10, 4299 (2019).
Godfray, H. C. J. et al. Meat consumption, health, and the environment. Science 361, eaam5324 (2018).
Delgado, C., Rosegrant, M., Steinfeld, H., Ehui, S. & Courbois, C. Livestock to 2020: The Next Food Revolution. Food, Agriculture, and the Environment Discussion Paper 28 (International Food Policy Research Institute, 1999).
Coker, R. et al. Towards a conceptual framework to support one-health research for policy on emerging zoonoses. Lancet Infect. Dis. 11, P326–P331 (2011).
Wu et al. Economic growth, urbanization, globalization, and the risks of emerging infectious diseases in China: a review. Ambio 46, 18–29 (2017).
Wilkinson, D. A., Marshall, J. C., French, N. P. & Hayman, D. T. Habitat fragmentation, biodiversity loss and the risk of novel infectious disease emergence. J. R. Soc. Interface 15, 20180403 (2018).
Johnson, C. K. et al. Global shifts in mammalian population trends reveal key predictors of virus spillover risk. Proc. R. Soc. B 287, 20192736 (2020).
Bloomfield, L. S. P., McIntosh, T. L. & Lambin, E. F. Habitat fragmentation, livelihood behaviors, and contact between people and nonhuman primates in Africa. Landsc. Ecol. 35, 985–1000 (2020).
Pulliam, J. R. et al. Agricultural intensification, priming for persistence and the emergence of Nipah virus: a lethal bat-borne zoonosis. J. R. Soc. Interface 9, 89–101 (2012).
Zhou, P. et al. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 5556, 255–258 (2018).
Allen, T. et al. Global hotspots and correlates of emerging zoonotic diseases. Nat. Commun. 8, 1124 (2017).
Meyer, C. F., Struebig, M. J. & Willig, M. R. in Bats in the Anthropocene: Conservation of Bats in a Changing World (eds Voigt, C.C. & Kingston, T.) 63–103 (Springer, 2016).
Gibb, R. et al. Zoonotic host diversity increases in human-dominated ecosystems. Nature 584, 398–402 (2020).
Cui, J., Li, F. & Shi, Z. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Hul V. et al. A novel SARS-CoV-2 related coronavirus in bats from Cambodia. Preprint at https://doi.org/10.1101/2021.01.26.428212 (2021).
Murakami, S. et al. Detection and characterization of bat Sarbecovirus phylogenetically related to SARS-CoV-2, Japan. Emerg. Infect. Dis. 26, 3025 (2020).
Wacharapluesadee, S. et al. Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia. Nat. Commun. 12, 972 (2021).
Soman Pillai, V., Krishna, G. & Valiya Veettil, M. Nipah virus: past outbreaks and future containment. Viruses. 12, 465 (2020).
Weingartl, H. M. et al. Susceptibility of pigs and chickens to SARS coronavirus. Emerg. Infect. Dis. 10, 179–184 (2004).
Schlottau, K. et al. SARS-CoV-2 in fruit bats, ferrets, pigs, and chickens: an experimental transmission study. Lancet Microbe 1, e218–e225 (2020).
Munnink, B. B. O. et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science 371, 172–177 (2021).
Zhou, L. et al. The re‐emerging of SADS‐CoV infection in pig herds in southern China. Transbound. Emerg. Dis. 66, 2180–2183 (2019).
Thuiller, W., Lafourcade, B., Engler, R. & Araújo, M. B. BIOMOD—a platform for ensemble forecasting of species distributions. Ecography 32, 369–373 (2009).
Chinazzi, M. et al. The effect of travel restrictions on the spread of the 2019 novel coronavirus (COVID-19) outbreak. Science 368, 395–400 (2020).
Yang, Q. et al. Assessing the role of live poultry trade in community-structured transmission of avian influenza in China. Proc. Natl Acad. Sci. USA 117, 5949–5954 (2020).
D’Odorico, P. et al. The global food–energy–water nexus. Rev. Geophys. 56, 456–531 (2018).
Meyfroidt, P., Lambin, E. F., Erb, K. H. & Hertel, T. W. Globalization of land use: distant drivers of land change and geographic displacement of land use. Curr. Opin. Environ. Sustain. 5, 438–444 (2013).
Ning, J. et al. Spatiotemporal patterns and characteristics of land-use change in China during 2010–2015. In J. Geogr. Sci. 28, 547–562 (2018).
Chen, C. et al. China and India lead in greening of the world through land-use management. Nat. Sustain. 2, 122–129 (2019).
Liu, J. et al. Forest fragmentation in China and its effect on biodiversity. Biol. Rev. 94, 1636–1657 (2019).
Whitmee, S. et al. Safeguarding human health in the Anthropocene epoch: report of the Rockefeller Foundation–Lancet Commission on planetary health. Lancet 386, 1973–2028 (2015).
Andersen, K. G., Rambaut, A., Lipkin, W. I., Holmes, E. C. & Garry, R. F. The proximal origin of SARS-CoV-2. Nat. Med. 26, 450–452 (2020).
Dietz, C., Dietz, I., Ivanova, T. & Siemers, B. M. Seasonal and regional scale movements of horseshoe bats (Rhinolophus, Chiroptera: Rhinolophidae) in northern Bulgaria. Nyctalus NF 14, 52–64 (2009).
Wang, J. et al. Seasonal habitat use by greater horseshoe bat Rhinolophus ferrumequinum (Chiroptera: Rhinolophidae) in Changbai Mountain temperate forest, northeast China. Mammalia 74, 257–266 (2010).
Robinson, T. P. et al. Mapping the global distribution of livestock. PLoS ONE 9, e96084 (2014).
Teluguntla, P. et al. in Land Resources: Monitoring, Modelling, and Mapping, Remote Sensing Handbook Vol. II (eds Prasad, S. & Thenkabail, P. S.) Ch. 7 (CRC Press Inc, 2014).
Hansen, M. C. et al. High-resolution global maps of 21st-century forest cover change. Science 342, 850–853 (2013).
Gilbert, M. et al. Global distribution data for cattle, buffaloes, horses, sheep, goats, pigs, chickens and ducks in 2010. Sci. Data 5, 180227 (2018).
Congalton, R. G. et al. NASA Making Earth System Data Records for Use in Research Environments (MEaSUREs) Global Food Security-support Analysis Data (GFSAD) @ 30-m: Cropland Extent Validation (GFSAD30VAL) (NASA EOSDIS Land Processes DAAC, 2017); https://doi.org/10.5067/MEaSUREs/GFSAD/GFSAD30VAL.001
Nieves, J. J. et al. Annually modelling built-settlements between remotely-sensed observations using relative changes in subnational populations and lights at night. Comput. Environ. Urban Syst. 80, 101444 (2020).
Vogt, P., Riitters, K. H., Estreguil, C. J., Kozak, T. G. & Wade, J. D. Wickham mapping spatial patterns with morphological image processing. Landsc. Ecol. 22, 171–177 (2007).
Assuncao, R. M., Neves, M. C., Camara, G. & Da Costa Freitas, C. Efficient regionalisation techniques for socio-economic geographical units using minimum spanning trees. Int. J. Geogr. Inf. Sci. 20, 797–811 (2006).
Acknowledgements
M.C.R and N.G. are supported by the ENI Enrico Mattei Foundation (FEEM), the Cariplo Foundation (SusFeed project 0737 CUP D49H170000300007) and Regione Lombardia (RUD0CONV01/ASSO project D44I20002000002). D.T.S.H. is supported by Royal Society Te Apārangi Rutherford Discovery Fellowship RDF‐MAU1701 MAU1701 and Bryce Carmine and Anne Carmine (née Percival), through the Massey University Foundation.
Author information
Authors and Affiliations
Contributions
M.C.R. designed the research. M.C.R., N.G. and D.T.S.H performed the research. M.C.R., P.D. and N.G. analysed the data. P.D., M.C.R. and D.T.S.H wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Food thanks Ruediger Schaldach, Xia Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Figs. 1–15.
Rights and permissions
About this article
Cite this article
Rulli, M.C., D’Odorico, P., Galli, N. et al. Land-use change and the livestock revolution increase the risk of zoonotic coronavirus transmission from rhinolophid bats. Nat Food 2, 409–416 (2021). https://doi.org/10.1038/s43016-021-00285-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43016-021-00285-x
This article is cited by
-
Organoids in virology
npj Viruses (2024)
-
Pathogen spillover driven by rapid changes in bat ecology
Nature (2023)
-
Generation of three-dimensional meat-like tissue from stable pig epiblast stem cells
Nature Communications (2023)
-
Using drivers and transmission pathways to identify SARS-like coronavirus spillover risk hotspots
Nature Communications (2023)
-
An argument for pandemic risk management using a multidisciplinary One Health approach to governance: an Australian case study
Globalization and Health (2022)