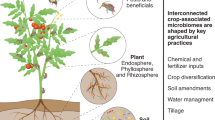
Abstract
The intimate interactions of indigenous crops with their associated microbiomes during long-term co-evolution strengthen the capacity and flexibility of crops to cope with biotic and abiotic stresses. This represents a promising untapped field for searching novel tools to sustainably increase crop productivity. However, the current capability of harnessing the power of indigenous crop microbiomes for sustainable crop production is limited due to low efficiency of separating the targeted functional microbes. Here, we highlight the potential benefits and existing challenges of utilizing indigenous crop microbiomes to reduce agrochemical inputs and increase crop resistance to biotic and abiotic stresses. We propose a framework using Raman-spectroscopy-based single-cell-sorting technology combined with a synthetic community approach to design and optimize a functionally reliable ‘beneficial biome’ under controlled conditions. This framework will offer opportunities for sustainable agriculture and provide a new direction for future studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Savci, S. An agricultural pollutant: chemical fertilizer. Int. J. Environ. Sci. Dev. 3, 77–80 (2012).
Guo, J. H. et al. Significant acidification in major Chinese croplands. Science 327, 1008–1010 (2010).
Raza, S. et al. Dramatic loss of inorganic carbon by nitrogen‐induced soil acidification in Chinese croplands. Glob. Change Biol. 26, 3738–3751 (2020).
Jez, J. M., Lee, S. G. & Sherp, A. M. The next green movement: plant biology for the environment and sustainability. Science 353, 1241–1244 (2016).
Cordovez, V., Dini-Andreote, F., Carrion, V. J. & Raaijmakers, J. M. Ecology and evolution of plant microbiomes. Annu. Rev. Microbiol. 73, 69–88 (2019).
Duran, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983.e914 (2018).
Dini-Andreote, F. & Raaijmakers, J. M. Embracing community ecology in plant microbiome research. Trends Plant Sci. 23, 467–469 (2018).
de Vries, F. T., Griffiths, R. I., Knight, C. G., Nicolitch, O. & Williams, A. Harnessing rhizosphere microbiomes for drought-resilient crop production. Science 368, 270–274 (2020).
Hubbard, C. J. et al. The effect of rhizosphere microbes outweighs host plant genetics in reducing insect herbivory. Mol. Ecol. 28, 1801–1811 (2019).
Oldroyd, G. E. D. & Leyser, O. A plant’s diet, surviving in a variable nutrient environment. Science 368, eaba0196 (2020).
Tedersoo, L., Bahram, M. & Zobel, M. How mycorrhizal associations drive plant population and community biology. Science 367, eaba1223 (2020).
Martín‐Robles, N. et al. Impacts of domestication on the arbuscular mycorrhizal symbiosis of 27 crop species. New Phytol. 218, 322–334 (2018).
Genre, A., Lanfranco, L., Perotto, S. & Bonfante, P. Unique and common traits in mycorrhizal symbioses. Nat. Rev. Microbiol. 18, 649–660 (2020).
Liu, X. et al. Partitioning of soil phosphorus among arbuscular and ectomycorrhizal trees in tropical and subtropical forests. Ecol. Lett. 21, 713–723 (2018).
Varoquaux, N. et al. Transcriptomic analysis of field-droughted sorghum from seedling to maturity reveals biotic and metabolic responses. Proc. Natl Acad. Sci. USA 116, 27124–27132 (2019).
Lazcano, C., Barrios-Masias, F. H. & Jackson, L. E. Arbuscular mycorrhizal effects on plant water relations and soil greenhouse gas emissions under changing moisture regimes. Soil Biol. Biochem. 74, 184–192 (2014).
Sprent, J. I. Evolving ideas of legume evolution and diversity: a taxonomic perspective on the occurrence of nodulation. New Phytol. 174, 11–25 (2007).
Soltis, D. E. et al. Chloroplast gene sequence data suggest a single origin of the predisposition for symbiotic nitrogen fixation in angiosperms. Proc. Natl Acad. Sci. USA 92, 2647–2651 (1995).
Young, N. D. et al. The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480, 520–524 (2011).
van Velzen, R. et al. Comparative genomics of the nonlegume Parasponia reveals insights into evolution of nitrogen-fixing Rhizobium symbioses. Proc. Natl Acad. Sci. USA 115, E4700–E4709 (2018).
Smil, V. Nitrogen in crop production: an account of global flows. Glob. Biogeochem. Cycles 13, 647–662 (1999).
O’Hara, G. W. The role of nitrogen fixation in crop production. J. Crop Prod. 1, 115–138 (1998).
Remigi, P., Zhu, J., Young, J. P. W. & Masson-Boivin, C. Symbiosis within symbiosis: evolving nitrogen-fixing legume symbionts. Trends Microbiol. 24, 63–75 (2016).
Garcia, K., Delaux, P. M., Cope, K. R. & Ané, J. M. Molecular signals required for the establishment and maintenance of ectomycorrhizal symbioses. New Phytol. 208, 79–87 (2015).
Fisher, R. F. & Long, S. R. Rhizobium–plant signal exchange. Nature 357, 655–660 (1992).
Cao, Y., Halane, M. K., Gassmann, W. & Stacey, G. The role of plant innate immunity in the legume–Rhizobium symbiosis. Annu. Rev. Plant Biol. 68, 535–561 (2017).
Ferguson, B. J. et al. Legume nodulation: the host controls the party. Plant Cell Environ. 42, 41–51 (2019).
Remans, R. et al. Effect of Rhizobium–Azospirillum coinoculation on nitrogen fixation and yield of two contrasting Phaseolus vulgaris L. genotypes cultivated across different environments in Cuba. Plant Soil 312, 25–37 (2008).
Cassán, F. & Diaz-Zorita, M. Azospirillum sp. in current agriculture: from the laboratory to the field. Soil Biol. Biochem. 103, 117–130 (2016).
Han, Q. et al. Variation in rhizosphere microbial communities and its association with the symbiotic efficiency of rhizobia in soybean. ISME J. 14, 1915–1928 (2020).
Saharan, B. S. & Nehra, V. Plant growth promoting rhizobacteria: a critical review. Life Sci. Med. Res. 21, 30 (2011).
Cheng, Y. T., Zhang, L. & He, S. Y. Plant–microbe interactions facing environmental challenge. Cell Host Microbe 26, 183–192 (2019).
Dini-Andreote, F. Endophytes: the second layer of plant defense. Trends Plant Sci. 25, 319–322 (2020).
Carrión, V. J. et al. Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366, 606–612 (2019).
Sieber, M. et al. Neutrality in the metaorganism. PLoS Biol. 17, e3000298 (2019).
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T. & Singh, B. K. Plant–microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 18, 607–621 (2020).
Burns, A. R. et al. Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. 10, 655–664 (2016).
Sloan, W. T. et al. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ. Microbiol. 8, 732–740 (2006).
Ning, D., Deng, Y., Tiedje, J. M. & Zhou, J. A general framework for quantitatively assessing ecological stochasticity. Proc. Natl Acad. Sci. USA 116, 16892–16898 (2019).
Carlström, C. I. et al. Synthetic microbiota reveal priority effects and keystone strains in the Arabidopsis phyllosphere. Nat. Ecol. Evol. 3, 1445–1454 (2019).
Purugganan, M. D. & Fuller, D. Q. The nature of selection during plant domestication. Nature 457, 843–848 (2009).
Chen, Y. H., Gols, R. & Benrey, B. Crop domestication and its impact on naturally selected trophic interactions. Annu. Rev. Entomol. 60, 35–58 (2015).
Szoboszlay, M. et al. Comparison of root system architecture and rhizosphere microbial communities of Balsas teosinte and domesticated corn cultivars. Soil Biol. Biochem. 80, 34–44 (2015).
Perez-Jaramillo, J. E., Mendes, R. & Raaijmakers, J. M. Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol. Biol. 90, 635–644 (2016).
Perez-Jaramillo, J. E., Carrion, V. J., de Hollander, M. & Raaijmakers, J. M. The wild side of plant microbiomes. Microbiome 6, 143 (2018).
Emmett, B. D., Buckley, D. H., Smith, M. E. & Drinkwater, L. E. Eighty years of maize breeding alters plant nitrogen acquisition but not rhizosphere bacterial community composition. Plant Soil 431, 53–69 (2018).
Mutch, L. A. & Young, J. P. W. Diversity and specificity of Rhizobium leguminosarum biovar viciae on wild and cultivated legumes. Mol. Ecol. 13, 2435–2444 (2004).
Kiers, E. T., Hutton, M. G. & Denison, R. F. Human selection and the relaxation of legume defences against ineffective rhizobia. Proc. R. Soc. B 274, 3119–3126 (2007).
Pérez-Jaramillo, J. E. et al. Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J. 11, 2244–2257 (2017).
Zachow, C., Müller, H., Tilcher, R. & Berg, G. Differences between the rhizosphere microbiome of Beta vulgaris ssp. maritima—ancestor of all beet crops—and modern sugar beets. Front. Microbiol. 5, 415 (2014).
Coleman‐Derr, D. et al. Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol. 209, 798–811 (2016).
Warschefsky, E., Penmetsa, R. V., Cook, D. R. & von Wettberg, E. J. Back to the wilds: tapping evolutionary adaptations for resilient crops through systematic hybridization with crop wild relatives. Am. J. Bot. 101, 1791–1800 (2014).
Brozynska, M., Furtado, A. & Henry, R. J. Genomics of crop wild relatives: expanding the gene pool for crop improvement. Plant Biotechnol. J. 14, 1070–1085 (2016).
Zhang, H., Mittal, N., Leamy, L. J., Barazani, O. & Song, B. H. Back into the wild—apply untapped genetic diversity of wild relatives for crop improvement. Evol. Appl. 10, 5–24 (2017).
Maxted, N. & Kell, S. P. Establishment of a Global Network for the In Situ Conservation of Crop Wild Relatives: Status and Needs (FAO Commission on Genetic Resources for Food and Agriculture, 2009).
Stenberg, J. A., Heil, M., Åhman, I. & Björkman, C. Optimizing crops for biocontrol of pests and disease. Trends Plant Sci. 20, 698–712 (2015).
Heil, M. & Baldwin, I. T. Fitness costs of induced resistance: emerging experimental support for a slippery concept. Trends Plant Sci. 7, 61–67 (2002).
Liu, H. & Brettell, L. E. Plant defense by VOC-induced microbial priming. Trends Plant Sci. 24, 187–189 (2019).
Schulz-Bohm, K. et al. Calling from distance: attraction of soil bacteria by plant root volatiles. ISME J. 12, 1252–1262 (2018).
Ehlers, B. K. et al. Plant secondary compounds in soil and their role in belowground species interactions. Trends Ecol. Evol. 35, 716–730 (2020).
Preece, C. & Penuelas, J. A return to the wild: root exudates and food security. Trends Plant Sci. 25, 14–21 (2020).
Rasmann, S. et al. Recruitment of entomopathogenic nematodes by insect-damaged maize roots. Nature 434, 732–737 (2005).
Köllner, T. G. et al. A maize (E)-β-caryophyllene synthase implicated in indirect defense responses against herbivores is not expressed in most American maize varieties. Plant Cell 20, 482–494 (2008).
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015).
Vorholt, J. A., Vogel, C., Carlstrom, C. I. & Muller, D. B. Establishing causality: opportunities of synthetic communities for plant microbiome research. Cell Host Microbe 22, 142–155 (2017).
Zhang, J. et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 37, 676–684 (2019).
Hatzenpichler, R., Krukenberg, V., Spietz, R. L. & Jay, Z. J. Next-generation physiology approaches to study microbiome function at single cell level. Nat. Rev. Microbiol. 18, 241–256 (2020).
Cui, L., Zhang, D., Yang, K., Zhang, X. & Zhu, Y. G. Perspective on surface-enhanced Raman spectroscopic investigation of microbial world. Anal. Chem. 91, 15345–15354 (2019).
Wang, Y., Huang, W. E., Cui, L. & Wagner, M. Single cell stable isotope probing in microbiology using Raman microspectroscopy. Curr. Opin. Biotechnol. 41, 34–42 (2016).
Cui, L. et al. Functional single-cell approach to probing nitrogen-fixing bacteria in soil communities by resonance Raman spectroscopy with 15N2 labeling. Anal. Chem. 90, 5082–5089 (2018).
Yang, K. et al. Rapid antibiotic susceptibility testing of pathogenic bacteria using heavy-water-labeled single-cell Raman spectroscopy in clinical samples. Anal. Chem. 91, 6296–6303 (2019).
Li, H. Z. et al. D2O-isotope-labeling approach to probing phosphate-solubilizing bacteria in complex soil communities by single-cell Raman spectroscopy. Anal. Chem. 91, 2239–2246 (2019).
Moutia, J.-F. Y., Saumtally, S., Spaepen, S. & Vanderleyden, J. Plant growth promotion by Azospirillum sp. in sugarcane is influenced by genotype and drought stress. Plant Soil 337, 233–242 (2010).
Bashan, Y. & De-Bashan, L. E. How the plant growth-promoting bacterium Azospirillum promotes plant growth—a critical assessment. Adv. Agron. 108, 77–136 (2010).
Figueiredo, M. V. B., Burity, H. A., Martínez, C. R. & Chanway, C. P. Alleviation of drought stress in the common bean (Phaseolus vulgaris L.) by co-inoculation with Paenibacillus polymyxa and Rhizobium tropici. Appl. Soil Ecol. 40, 182–188 (2008).
Uma, C., Sivagurunathan, P. & Sangeetha, D. Performance of bradyrhizobial isolates under drought conditions. Int. J. Curr. Microbiol. App. Sci. 2, 228–232 (2013).
Tank, N. & Saraf, M. Salinity-resistant plant growth promoting rhizobacteria ameliorates sodium chloride stress on tomato plants. J. Plant Interact. 5, 51–58 (2010).
Tahir, H. A. et al. Plant growth promotion by volatile organic compounds produced by Bacillus subtilis SYST2. Front. Microbiol. 8, 171 (2017).
Vardharajula, S., Zulfikar Ali, S., Grover, M., Reddy, G. & Bandi, V. Drought-tolerant plant growth promoting Bacillus spp.: effect on growth, osmolytes, and antioxidant status of maize under drought stress. J. Plant Interact. 6, 1–14 (2011).
Santoyo, G., Orozco-Mosqueda, M. D. C. & Govindappa, M. Mechanisms of biocontrol and plant growth-promoting activity in soil bacterial species of Bacillus and Pseudomonas: a review. Biocontrol Sci. Technol. 22, 855–872 (2012).
Leclere, V. et al. Mycosubtilin overproduction by Bacillus subtilis BBG100 enhances the organism’s antagonistic and biocontrol activities. Appl. Environ. Microbiol. 71, 4577–4584 (2005).
Hu, J. et al. Probiotic Pseudomonas communities enhance plant growth and nutrient assimilation via diversity-mediated ecosystem functioning. Soil Biol. Biochem. 113, 122–129 (2017).
Kohler, J., Hernández, J. A., Caravaca, F. & Roldán, A. Plant-growth-promoting rhizobacteria and arbuscular mycorrhizal fungi modify alleviation biochemical mechanisms in water-stressed plants. Funct. Plant Biol. 35, 141–151 (2008).
Nassar, A. H., El-Tarabily, K. A. & Sivasithamparam, K. Growth promotion of bean (Phaseolus vulgaris L.) by a polyamine-producing isolate of Streptomyces griseoluteus. Plant Growth Reg. 40, 97–106 (2003).
Gopalakrishnan, S. et al. Evaluation of Streptomyces strains isolated from herbal vermicompost for their plant growth-promotion traits in rice. Microbiol. Res. 169, 40–48 (2014).
Kwak, M.-J. et al. Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 36, 1100–1109 (2018).
Sang, M. K. & Kim, K. D. The volatile‐producing Flavobacterium johnsoniae strain GSE09 shows biocontrol activity against Phytophthora capsici in pepper. J. Appl. Microbiol. 113, 383–398 (2012).
Naznin, H. A. et al. Systemic resistance induced by volatile organic compounds emitted by plant growth-promoting fungi in Arabidopsis thaliana. PLoS ONE 9, e86882 (2014).
Kiss, L., Russell, J. C., Szentiványi, O., Xu, X. & Jeffries, P. Biology and biocontrol potential of Ampelomyces mycoparasites, natural antagonists of powdery mildew fungi. Biocontrol Sci. Technol. 14, 635–651 (2004).
Lee, S., Yap, M., Behringer, G., Hung, R. & Bennett, J. W. Volatile organic compounds emitted by Trichoderma species mediate plant growth. Fungal Biol. Biotechnol. 3, 1–14 (2016).
Zhang, S., Gan, Y. & Xu, B. Application of plant-growth-promoting fungi Trichoderma longibrachiatum t6 enhances tolerance of wheat to salt stress through improvement of antioxidative defense system and gene expression. Front. Plant Sci. 7, 1405 (2016).
van der Meij, A., Worsley, S. F., Hutchings, M. I. & van Wezel, G. P. Chemical ecology of antibiotic production by Actinomycetes. FEMS Microbiol. Rev. 41, 392–416 (2017).
Bhatti, A. A., Haq, S. & Bhat, R. A. Actinomycetes benefaction role in soil and plant health. Microb. Pathog. 111, 458–467 (2017).
Chaurasia, A. et al. Actinomycetes: an unexplored microorganisms for plant growth promotion and biocontrol in vegetable crops. World J. Microbiol. Biotechnol. 34, 1–16 (2018).
Ercoli, L., Schüßler, A., Arduini, I. & Pellegrino, E. Strong increase of durum wheat iron and zinc content by field-inoculation with arbuscular mycorrhizal fungi at different soil nitrogen availabilities. Plant Soil 419, 153–167 (2017).
Xu, L. et al. Arbuscular mycorrhiza enhances drought tolerance of tomato plants by regulating the 14-3-3 genes in the ABA signaling pathway. Appl. Soil Ecol. 125, 213–221 (2018).
Ghorchiani, M., Etesami, H. & Alikhani, H. A. Improvement of growth and yield of maize under water stress by co-inoculating an arbuscular mycorrhizal fungus and a plant growth promoting rhizobacterium together with phosphate fertilizers. Agric. Ecosyst. Environ. 258, 59–70 (2018).
Meeds, J. A. et al. Phosphorus deficiencies invoke optimal allocation of exoenzymes by ectomycorrhizas. ISME J. https://doi.org/10.1038/s41396-020-00864-z (2021).
Acknowledgements
This work was supported by the Australian Research Council (FT190100383 and DE210100271).
Author information
Authors and Affiliations
Contributions
All authors contributed intellectual input and assistance to this work. The original concepts were conceived by J.-Z.H., Q.-L.C., H.-W.H. and Y.-G.Z. Q.-L.C. and J.-Z.H. wrote the first draft, with subsequent input during revision from H.-W.H., Z.-Y.H., L.C. and Y.-G.Z. All the figures are original from Q.-L.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Food thanks Josep Penuelas, Zhong Wei, Andrea Genre and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, QL., Hu, HW., He, ZY. et al. Potential of indigenous crop microbiomes for sustainable agriculture. Nat Food 2, 233–240 (2021). https://doi.org/10.1038/s43016-021-00253-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43016-021-00253-5
This article is cited by
-
Serial cultures in invert emulsion and monophase systems for microbial community shaping and propagation
Microbial Cell Factories (2024)
-
Challenges and opportunities for increasing the use of low-risk plant protection products in sustainable production. A review
Agronomy for Sustainable Development (2024)
-
Frankia-actinorhizal symbiosis: a non-chemical biological assemblage for enhanced plant growth, nodulation and reclamation of degraded soils
Symbiosis (2024)
-
Plant domestication shapes rhizosphere microbiome assembly and metabolic functions
Microbiome (2023)
-
Root microbiota confers rice resistance to aluminium toxicity and phosphorus deficiency in acidic soils
Nature Food (2023)