Abstract
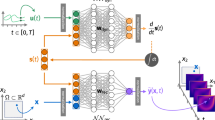
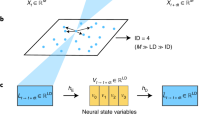
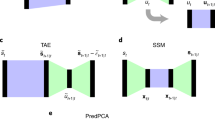
Dynamical models underpin our ability to understand and predict the behaviour of natural systems. Whether dynamical models are developed from first-principles derivations or from observational data, they are predicated on our choice of state variables. The choice of state variables is driven by convenience and intuition, and, in data-driven cases, the observed variables are often chosen to be the state variables. The dimensionality of these variables (and consequently the dynamical models) can be arbitrarily large, obscuring the underlying behaviour of the system. In truth these variables are often highly redundant and the system is driven by a much smaller set of latent intrinsic variables. In this study we combine the mathematical theory of manifolds with the representational capacity of neural networks to develop a method that learns a system’s intrinsic state variables directly from time-series data, as well as predictive models for their dynamics. What distinguishes our method is its ability to reduce data to the intrinsic dimensionality of the nonlinear manifold they live on. This ability is enabled by the concepts of charts and atlases from the theory of manifolds, whereby a manifold is represented by a collection of patches that are sewn together—a necessary representation to attain intrinsic dimensionality. We demonstrate this approach on several high-dimensional systems with low-dimensional behaviour. The resulting framework provides the ability to develop dynamical models of the lowest possible dimension, capturing the essence of a system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data are available at https://doi.org/10.5281/zenodo.7219159 (ref. 63). For data that are unavailable due to size restrictions, code that exactly reproduces the data has been deposited in the same repository.
Code availability
Code is available at https://doi.org/10.5281/zenodo.7219159 (ref. 63).
References
N. Watters et al. Visual interaction networks: Learning a physics simulator from video. In Advances in Neural Information Processing Systems (eds Garnett, R. et al.) Vol. 30 (Curran Associates, 2017); https://proceedings.neurips.cc/paper/2017/file/8cbd005a556ccd4211ce43f309bc0eac-Paper.pdf
Gonzalez, F. J. & Balajewicz, M. Deep convolutional recurrent autoencoders for learning low-dimensional feature dynamics of fluid systems. Preprint at https://arxiv.org/abs/1808.01346 (2018).
Vlachas, P. R., Byeon, W., Wan, Z. Y., Sapsis, T. P. & Koumoutsakos, P. Data-driven forecasting of high-dimensional chaotic systems with long short-term memory networks. Proc. R. Soc. A 474, 20170844 (2018).
Champion, K., Lusch, B., Kutz, J. N. & Brunton, S. L. Data-driven discovery of coordinates and governing equations. Proc. Natl Acad. Sci. USA 116, 22445–22451 (2019).
Carlberg, K. T. et al. Recovering missing CFD data for high-order discretizations using deep neural networks and dynamics learning. J. Comput. Phys. 395, 105–124 (2019).
Linot, A. J. & Graham, M. D. Deep learning to discover and predict dynamics on an inertial manifold. Phys. Rev. E 101, 062209 (2020).
Maulik, R. et al. Time-series learning of latent-space dynamics for reduced-order model closure. Physica D 405, 132368 (2020).
Hasegawa, K., Fukami, K., Murata, T. & Fukagata, K. Machine-learning-based reduced-order modeling for unsteady flows around bluff bodies of various shapes. Theor. Comput. Fluid Dyn. 34, 367–383 (2020).
Linot, A. J. & Graham, M. D. Data-driven reduced-order modeling of spatiotemporal chaos with neural ordinary differential equations. Chaos 32, 073110 (2022).
Maulik, R., Lusch, B. & Balaprakash, P. Reduced-order modeling of advection-dominated systems with recurrent neural networks and convolutional autoencoders. Phys. Fluids 33, 037106 (2021).
Rojas, C. J. G., Dengel, A. & Ribeiro, M. D. Reduced-order model for fluid flows via neural ordinary differential equations. Preprint at https://arxiv.org/abs/2102.02248 (2021)
Vlachas, P. R., Arampatzis, G., Uhler, C. & Koumoutsakos, P. Multiscale simulations of complex systems by learning their effective dynamics. Nat. Mach. Intell. 4, 359–366 (2022).
Takens, F. Detecting strange attractors in turbulence. In Dynamical Systems and Turbulence, Warwick 1980 (eds Rand, D. & Young, L.-S.) 366–381 (Springer, 1981).
Fefferman, C., Mitter, S. & Narayanan, H. Testing the manifold hypothesis. J. Am. Mathematical Soc. 29, 983–1049 (2016).
Hopf, E. A mathematical example displaying features of turbulence. Commun. Pure Appl. Math. 1, 303–322 (1948).
Foias, C., Sell, G. R. & Temam, R. Inertial manifolds for nonlinear evolutionary equations. J. Differ. Equ. 73, 309–353 (1988).
Temam, R. & Wang, X. M. Estimates on the lowest dimension of inertial manifolds for the Kuramoto–Sivashinsky equation in the general case. Differ. Integral Equ. 7, 1095–1108 (1994).
Doering, C. R. & Gibbon, J. D. Applied Analysis of the Navier-Stokes Equations Cambridge Texts in Applied Mathematics No. 12 (Cambridge Univ. Press, 1995)
Schölkopf, B., Smola, A. & Müller, K.-R. Nonlinear component analysis as a kernel eigenvalue problem. Neural Comput. 10, 1299–1319 (1998).
Tenenbaum, J. B., De Silva, V. & Langford, J. C. A global geometric framework for nonlinear dimensionality reduction. Science 290, 2319–2323 (2000).
Roweis, S. T. & Saul, L. K. Nonlinear dimensionality reduction by locally linear embedding. Science 290, 2323–2326 (2000).
Belkin, M. & Niyogi, P. Laplacian eigenmaps for dimensionality reduction and data representation. Neural Comput. 15, 1373–1396 (2003).
Donoho, D. L. & Grimes, C. Hessian eigenmaps: locally linear embedding techniques for high-dimensional data. Proc. Natl Acad. Sci. USA 100, 5591–5596 (2003).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, 2016).
Ma, Y. & Fu, Y. Manifold Learning Theory and Applications Vol. 434 (CRC, 2012)
Bregler, C. & Omohundro, S. Surface learning with applications to lipreading. In Advances in Neural Information Processing Systems (eds Alspector, J.) Vol. 6 (Morgan-Kaufmann, 1994); https://proceedings.neurips.cc/paper/1993/file/96b9bff013acedfb1d140579e2fbeb63-Paper.pdf
Hinton G. E., Revow, M. & Dayan, P. Recognizing handwritten digits using mixtures of linear models. In Advances in Neural Information Processing Systems (eds Leen, T. et al.) Vol. 7 (MIT Press, 1995); https://proceedings.neurips.cc/paper/1994/file/5c936263f3428a40227908d5a3847c0b-Paper.pdf
Kambhatla, N. & Leen, T. K. Dimension reduction by local principal component analysis. Neural Comput. 9, 1493–1516 (1997).
Roweis, S., Saul, L. & Hinton, G. E. Global coordination of local linear models. In Advances in Neural Information Processing Systems (eds Ghahramani, Z.) Vol. 14 (MIT Press, 2002); https://proceedings.neurips.cc/paper/2001/file/850af92f8d9903e7a4e0559a98ecc857-Paper.pdf
Brand, M. Charting a manifold. In Advances in Neural Information Processing Systems (eds. Obermayer, K.) Vol. 15, 985–992 (MIT Press, 2003); https://proceedings.neurips.cc/paper/2002/file/8929c70f8d710e412d38da624b21c3c8-Paper.pdf
Amsallem, D., Zahr, M. J. & Farhat, C. Nonlinear model order reduction based on local reduced-order bases. Int. J. Numer. Meth. Eng. 92, 891–916 (2012).
Pitelis, N., Russell, C. & Agapito, L. Learning a manifold as an atlas. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 1642–1649 (IEEE, 2013).
Schonsheck, S, Chen, J. & Lai, R. Chart auto-encoders for manifold structured data. Preprint at https://arxiv.org/abs/1912.10094 (2019)
Lee, J. M. Introduction to Smooth Manifolds (Springer, 2013)
MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proc. 5th Berkeley Symposium on Mathematical Statistics and Probability Vol. 5.1, (eds Neyman, J.) 281–297 (Statistical Laboratory of the University of California, 1967).
Steinhaus, H. Sur la division des corps matériels en parties. Bull. Acad. Polon. Sci 4, 801–804 (1957).
Lloyd, S. Least squares quantization in PCM. IEEE Trans. Inform. Theory 28, 129–137 (1982).
Forgy, E. W. Cluster analysis of multivariate data: efficiency versus interpretability of classifications. Biometrics 21, 768–769 (1965).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Cybenko, G. Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 2, 303–314 (1989).
Hornik, K. Approximation capabilities of multilayer feedforward networks. Neural Netw. 4, 251–257 (1991).
Pinkus, A. Approximation theory of the MLP model in neural networks. Acta Numerica 8, 143–195 (1999).
Bottou, L. & Bousquet, O. The tradeoffs of large scale learning. In Advances in Neural Information Processing Systems (edited Roweis, S.) Vol. 20 (Curran Associates, 2007); https://proceedings.neurips.cc/paper/2007/file/0d3180d672e08b4c5312dcdafdf6ef36-Paper.pdf
Jing, L., Zbontar, J. & LeCun, Y. Implicit Rank-Minimizing Autoencoder. In Advances in Neural Information Processing Systems (eds Lin, H. et al.) Vol. 33 (Curran Associates, 2020); https://proceedings.neurips.cc/paper/2020/file/a9078e8653368c9c291ae2f8b74012e7-Paper.pdf
Chen, B. et al. Automated discovery of fundamental variables hidden in experimental data. Nat. Comput. Sci. 2, 433–442 (2022).
Kirby, M. & Armbruster, D. Reconstructing phase space from PDE simulations. Zeit. Angew. Math. Phys. 43, 999–1022 (1992).
Kevrekidis, I. G., Nicolaenko, B. & Scovel, J. C. Back in the saddle again: a computer assisted study of the Kuramoto–Sivashinsky equation. SIAM J. Appl.Math. 50, 760–790 (1990).
Whitney, H. The self-intersections of a smooth n-manifold in 2n-space. Ann Math 45, 220–246 (1944).
Graham, M. D. & Kevrekidis, I. G. Alternative approaches to the Karhunen-Loeve decomposition for model reduction and data analysis. Comput. Chem. Eng. 20, 495–506 (1996).
Takeishi, N., Kawahara, Y. & Yairi, T. Learning Koopman invariant subspaces for dynamic mode decomposition. In Advances in Neural Information Processing Systems (eds Garnett, R. et al.) Vol. 30 (Curran Associates, 2017); https://proceedings.neurips.cc/paper/2017/file/3a835d3215755c435ef4fe9965a3f2a0-Paper.pd
Lusch, B., Kutz, J. N. & Brunton, S. L. Deep learning for universal linear embeddings of nonlinear dynamics. Nat. Commun. 9, 1–10 (2018).
Otto, S. E. & Rowley, C. W. Linearly recurrent autoencoder networks for learning dynamics. SIAM J. Appl. Dyn. Syst. 18, 558–593 (2019).
Pathak, J., Lu, Z., Hunt, B. R., Girvan, M. & Ott, E. Using machine learning to replicate chaotic attractors and calculate Lyapunov exponents from data. Chaos 27, 121102 (2017).
Pathak, J., Hunt, B., Girvan, M., Lu, Z. & Ott, E. Model-free prediction of large spatiotemporally chaotic systems from data: a reservoir computing approach. Phys. Rev. Lett. 120, 024102 (2018).
Vlachas, P. R. et al. Backpropagation algorithms and reservoir computing in recurrent neural networks for the forecasting of complex spatiotemporal dynamics. Neural Netw. 126, 191–217 (2020).
Cornea, O., Lupton, G., Oprea, J. & Tanré, D. Lusternik-Schnirelmann Category 103 (American Mathematical Society, 2003).
Camastra, F. & Staiano, A. Intrinsic dimension estimation: advances and open problems. Inform. Sci. 328, 26–41 (2016).
Nash, J. C1 isometric imbeddings. Ann. Math. 60, 383–396 (1954).
Kuiper, N. H. On C1-isometric imbeddings. I. Indag. Math. 58, 545–556 (1955).
Nash, J. The imbedding problem for Riemannian manifolds. Ann. Math. 63, 20–63 (1956).
Borrelli, V., Jabrane, S., Lazarus, F. & Thibert, B. Flat tori in three-dimensional space and convex integration. Proc. Natl Acad. Sci. USA 109, 7218–7223 (2012).
Floryan, D. & Graham, M. D. dfloryan/neural-manifold-dynamics: v1.0 (Zenodo, 2022); https://doi.org/10.5281/zenodo.7219159
Acknowledgements
We acknowledge the use of the Sabine cluster from the Research Computing Data Core at the University of Houston, and the assistance of D. A. Kaji and the Luke cluster. This work was supported by the Air Force Office of Scientific Research (grant no. FA9550-18-0174 to M.D.G.) and an Office of Naval Research grant (grant no. N00014-18-1-2865, Vannevar Bush Faculty Fellowship, to M.D.G.).
Author information
Authors and Affiliations
Contributions
D.F. and M.D.G. designed and performed the research, analysed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Stefan Schonscheck and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary discussion, tables and figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Floryan, D., Graham, M.D. Data-driven discovery of intrinsic dynamics. Nat Mach Intell 4, 1113–1120 (2022). https://doi.org/10.1038/s42256-022-00575-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-022-00575-4
This article is cited by
-
Predicting multiple observations in complex systems through low-dimensional embeddings
Nature Communications (2024)
-
Generative learning for nonlinear dynamics
Nature Reviews Physics (2024)
-
Learning the intrinsic dynamics of spatio-temporal processes through Latent Dynamics Networks
Nature Communications (2024)
-
Data-driven discovery of linear dynamical systems from noisy data
Science China Technological Sciences (2024)
-
Reconstructing computational system dynamics from neural data with recurrent neural networks
Nature Reviews Neuroscience (2023)