Abstract
B-cell receptors (BCRs) are a crucial player in the development and activation of B cells, and their mature forms are secreted as antibodies, which execute functions such as the neutralization of invading pathogens. All current analytical approaches for BCRs solely investigate the BCR sequences and ignore their correlations with the transcriptomics of the B cells, yielding conclusions of unknown functional relevance regarding the roles of BCRs and B cells, and could generate biased interpretation. Many single-cell RNA-sequencing (scRNA-seq) techniques can now capture both the gene expression and BCR of each B cell, which could potentially address this issue. Here, we investigated 43,938 B cells from 13 scRNA-seq datasets with matched scBCR sequencing, and we observed an association between the BCRs and the B cells’ transcriptomics. Motivated by this, we developed the Benisse model (BCR embedding graphical network informed by scRNA-seq) to provide refined analyses of BCRs guided by single-cell gene expression. Benisse revealed a gradient of B-cell activation along BCR trajectories. We discovered a stronger coupling between BCRs and B-cell gene expression during COVID-19 infections. We found that BCRs form a directed pattern of continuous and linear evolution to achieve the highest antigen targeting efficiency, compared with the convergent evolution pattern of T-cell receptors. Overall, a simultaneous digestion of the BCR and gene expression of B cells, viewed through the lens of Benisse, will lead to a more insightful interpretation of the functional relevance of the BCR repertoire in different biological contexts.
Similar content being viewed by others
Main
B-cell receptors (BCRs) recognize antigenic epitopes and control the activation and maturation of B cells1,2,3,4,5. B cells with mature BCRs differentiate into plasma cells that secrete antibodies, which are the secreted forms of BCRs and carry out a variety of functions, such as neutralization of invading pathogens6,7. In addition to their key roles in infectious diseases and autoimmune diseases, recent studies have also discovered curious parts that tumour-infiltrating B lymphocytes play in all stages of cancers, potentially in a BCR/antibody-dependent manner8,9,10,11,12,13,14.
Due to the importance and the complexity of BCRs, profiling of the BCR repertoire has been the core interest of many studies. BCR sequencing enables the direct sequencing of BCRs of thousands of cells at one time. Following the sequencing of BCRs, BCRs with the same variable (V) and joining (J) gene segments and the same complementarity-determining region 3 (CDR3) lengths are sometimes grouped as being clonally related15. Moreover, the phylogenetic relationships of the BCRs are usually derived using metrics such as Levenshtein distance16,17,18. Many studies employ certain diversity metrics to characterize the clonal richness of BCRs19,20,21. However, a fundamental challenge exists for all the aforementioned approaches—all conclusions are drawn solely on the basis of interrogating the BCR sequences, without knowing the functional relevance of the BCRs/antibodies.
To address this challenge, we need to investigate the coupling between the BCR repertoire and the transcriptomic status of the B cells, which could reveal the true functional implication of the BCR repertoire under various biomedical contexts. Several recently developed single-cell RNA-sequencing (scRNA-seq) technologies provide the necessary data to answer this question. Among them, the 10x Genomics Chromium platform, which directly amplifies the BCRs while capturing the expression information of the other genes at the same time, is currently the dominant technique. However, until now, most studies generating such data have simply analysed these two modalities of data using separate and ad hoc methods. Critical opportunities from an integrative analysis of scBCR-seq and scRNA-seq data to discover interesting biological insights have been missed.
In this work, we develop a mathematical model, named Benisse (Fig. 1a), to integrate the high-dimensional BCR and single-B-cell expression data. Benisse is based on a correlation effect that we observed between BCRs and B-cell gene expression. By validating and applying Benisse on 43,938 B cells from 13 scRNA-seq + scBCR-seq datasets, we showed that Benisse is capable of mapping the functional relevance of the BCR repertoire in various biological contexts, at single-cell resolution and supported by empirical evidence from single-B-cell expression.
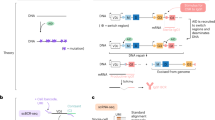
a, Schematic diagram showing how Benisse digests the BCR sequence data and the single-B-cell expression data to detect an embedding space of the BCR clonotypes that are supervised by the transcriptomic information, and also to detect a graph of BCR networks that connect BCR clonotypes (from a crude BCR graph that connects BCRs with the same V and J genes). Uniform manifold approximation and projection (UMAP) was performed for dimension reduction of the single-cell expression data. The BCR encoder outputs a 20-dimensional embedding of the BCRs. Thus they are labelled differently (‘UMAP 1’ versus ‘Dim 1’). B cell expression and BCRs of the same B cells, marked by the same colour (e.g., red, blue, yellow), are merged into one single latent space embedding point. b,c, Spearman correlation between BCR distances and distances between LIBRA-seq scores of all antigens (b) and the HA_indo antigen (c) for all pairs of BCR clonotypes. d, Benchmark analyses showing the correlations between BCRs and LIBRA-seq scores, for BCR embeddings derived from our contrastive learning model, from BcRep and from Lindenbaum et al.25 e, The distances in latent space between the BCR clonotypes that are connected in the BCR networks built by Benisse (left), that are not connected but share V/J genes (middle) and that do not share V/J genes (right). Results for all datasets are averaged. f, The correlations between expression and BCR of the B cells of the same networks, with BCR distances calculated from the Benisse latent space (‘cor(a, b)’, left-hand box of the boxplot) or the original BCR embedding space (‘cor(a, c)’, right-hand box).
Results
A numeric embedding for BCRs of B cells based on deep contrastive learning
To build Benisse, we need to first mathematically describe the peptide sequences of the BCRs. To achieve this, we focused on the complementarity-determining region of the heavy chain (CDR3H) regions of BCRs and built a numeric embedding of BCR CDR3H sequences. We encoded the BCR sequences by the ‘Atchley factors’22, representing each amino acid with five numeric values. Next, we further reduced the dimension of this Atchley factor matrix into a short numeric vector through contrastive learning23. During this step, the contrastive learning model learns a 20-dimensional embedding space where similar CDR3H peptide sequences are closer to each other, whereas the dissimilar ones are far apart (Extended Data Fig. 1).
We tested whether the CDR3H embedding is reflective of antigen specificity using the LIBRA-seq data24, which allows high-throughput mapping of antigen specificity of BCR sequences. A total of nine antigens were profiled in one setting, and the antigen specificities of 2,321 BCRs against these nine antigens were obtained as a continuous variable for each antigen. We calculated the pairwise similarities between BCRs, in terms of their LIBRA-seq antigen specificity scores (Euclidean distances between LIBRA-seq scores of all antigens), and also of their numeric embeddings of BCRs (also Euclidean distances). We showed that the correlations between BCR sequence embedding similarities and BCR antigen specificity similarities reached 0.616 (Fig. 1b), suggesting that our embedding can indeed reasonably reflect the key features of BCR CDR3Hs. We also performed this analysis with each individual antigen, and observed an overall positive but reduced correlation for each (Fig. 1c and Extended Data Fig. 2). As a benchmark, we performed the same analyses with the work of Lindenbaum et al.25 and bcRep26, in both of which we can also calculate similarity scores between different BCRs. In Fig. 1d, we show that our BCR CDR3H embeddings achieved a higher association with the LIBRA-seq scores compared with Lindenbaum et al. and bcRep.
We also accessed the BCR-sequencing data from Liao et al.27, who performed BCR sequencing for a single lineage of antibodies derived against one human immunodeficiency virus (HIV) epitope, collected at a series of times (weeks after HIV infection). We generated a pairwise Euclidean distance matrix between the BCR embeddings. A phylogenetic tree of BCRs was constructed from this distance matrix28, rooted at the unmutated common ancestor (UCA) of these BCRs. We observed that BCRs from earlier times are closer to the UCA on the tree (Extended Data Fig. 3a). The phylogenetic tree then branched out, and the leaves in the only major branch contain many BCRs from later times. In particular, the group of the best antigen-targeting antibodies of week 144, CH103, CH104, CH105 and CH106 (sharing the same CDR3H), were found at almost the farthest leaf of the tree. The waterfall plot with BCRs ordered by the Euclidean distances between the UCA BCR and all the other BCRs (Extended Data Fig. 3b) also confirms that BCRs from earlier times are closer to the UCA BCR in general. Taken together, these analyses validate our BCR embedding approach, and reveal an interesting linear evolution pattern of BCRs/antibodies.
Benisse integrates BCR and expression of single B cells
With the BCR embeddings, we then sought to investigate whether the BCRs are indeed correlated with the expression of the B cells. We investigated a total of 13 datasets that were generated from scRNA-seq with paired scBCR-seq (Extended Data Table 1). In each dataset, we embedded the BCRs as described above, and then we calculated the pairwise distances between BCR clonotypes using their BCR distances and also their gene expression distances. Interestingly, we observed a positive correlation between these two distances, suggesting that the BCR clonotypes with similar BCR sequences have similar gene expression profiles (Supplementary Note 2). Across all 13 datasets, we observed a positive correlation with an average of 0.32. Importantly, in this analysis, we removed pairs of BCRs that are the same (BCR distance = 0). We also separately examined the expression distances between pairs of B cells with BCRs that are the same (in the same clonotype) or differ. We observed that B cells in the same clonotype have much more similar expressions than those from different clonotypes (Supplementary Note 2).
Motivated by this observation, we built the core Benisse model (Supplementary Note 1), to enable a more refined interrogation of the relationships between BCRs and B-cell expression. The model searches for a latent space of the BCRs, supervised by the empirical evidence of their functional relevance provided through the B-cell gene expression. Many BCRs in one sample are usually clonally related, representing BCRs that were generated from one parental BCR due to somatic hypermutations15,29. Therefore, we need to detect sparse graphs of BCRs under the new latent space so that closely related BCRs will be connected into what we term ‘BCR networks’ (Fig. 1a). Mathematically, Benisse employs a sparse graph learning model to handle these requirements, where the BCRs are to be embedded in a low-dimensional manifold that may be reasonably expressed by a graph. Each vertex is a B-cell clonotype marked by a unique BCR (same V/J gene and same CDR3H) and the weight of each edge represents the similarity between two BCRs. This latent space is learned via the supervision of gene expression, and we require that BCRs closer to each other in the latent space should have similar BCR sequences and represent B cells with similar transcriptomic features. We also incorporated a prior requirement that an edge exists only when two BCRs share the same V gene and the same J gene. The resulting Benisse graph is comprised of many small BCR networks, with each network containing BCRs with the same V/J genes and similar CDR3Hs in the latent space.
In the scRNA-seq data that we collected, we calculated the distances in the latent space between the BCR clonotypes that are connected in the BCR networks, that are not connected but share V/J genes or that do not share the same V/J genes. In Fig. 1e, we show that the BCR distances in the latent space are the smallest for the BCR clonotypes that are connected within the same networks by Benisse (the first group). This is consistent with our expectation, as Benisse is supposed to group B-cell clonotypes with the most similar BCRs into individual BCR networks. In Fig. 1f, we also calculate the correlation between the expression-wise distances and BCR-wise distances of the BCR clonotypes that are connected in the same networks. We found that their correlation indeed increased in the Benisse latent space compared with the original BCR numeric embedding space. These results suggest that Benisse successfully achieved the maximal sharing of information between B-cell expression and the BCRs of B cells.
Benisse reveals a gradient of B-cell activation along BCR trajectories
During somatic hypermutations in the germinal centres (GCs), BCRs of B cells keep mutating until one or more BCRs with high affinity to the target antigens have been created, supposedly creating a family of continuously evolving BCRs with different antigen-targeting efficiencies (Fig. 2a). In the Benisse analyses, such networks of related BCRs are detected as a part of the algorithm. We calculated pseudotimes for the B cells using Monocle30, and we picked, in each BCR network, the BCR clonotypes that are most newly created according to pseudotimes. These recently generated BCR clonotypes probably represent the BCRs that are the ‘best’ in terms of antigen targeting, and therefore activation of B cells, within each of their networks (Fig. 2a). We term these BCR clonotypes the ‘prime’ clonotypes of their respective networks.
a, A schematic showing the proposed model of a linear and continuous pattern of BCR evolution. A phylogenetic tree-like pattern of BCR evolution is shown for one BCR network, where the BCRs are created through VDJ recombination and somatic hypermutation to become better and better at targeting the antigen. The x axis shows the direction of BCR evolution. The y axis shows the B-cell activity levels of the BCR clonotypes on the tree. The coloured bars on the Y axis refer to the levels of B cell activation in successive stages of BCR maturation as a result of SHM (marked by different colours). The dotted circles denote the prime clonotypes assigned by our model. SHMs, somatic hypermutations. b, An example dataset showing how the phenotypes of the B cells vary as a function of the BCR distance from the prime clonotypes (x axis). The y axis shows the expression level of the activation signature in the B cells. Clone sizes of the BCR clonotypes and class-switching status are also shown for each BCR clonotype. NSCLC, non-small-cell lung carcinoma. c, Boxplots showing the pseudotimes of the B cells in the BCR clonotypes that are grouped into groups 1, 2 and 3. Only BCR networks with at least three BCR clonotypes were included. B cells in group 1 have BCRs that are closest to those of the prime BCR clonotypes (and include prime BCR clonotypes themselves). Group 3 is most dissimilar from prime BCRs. d, Expression of the B-cell activation signature of the B cells belonging to groups 1, 2 and 3. e, GSEA plots of the pathways enriched in the genes that showed a monotonic increase or decrease of expression from the B cells of group 1 to group 3 BCR clonotypes. The COVID U41 dataset is shown as an example. f, The clonal sizes of the B cells belonging to groups 1, 2 and 3. g,h, The proportions of the IgD, M, G, A and E BCRs of all the B cells in the 10x melanoma dataset (g) or all datasets (h) in each of the three groups.
To evaluate our hypothesized paradigm of BCR evolution, we calculated the similarity of each BCR to its network’s prime BCR, in terms of Euclidean distance in the latent space learned by Benisse (x axis of Fig. 2b). We divided BCR clonotypes of each BCR network into three groups on the basis of this BCR distance. The first group of BCRs (including the prime BCR clonotypes themselves) has the largest similarity to the prime BCR clonotypes, and the third group of BCRs has the least similarity. We then investigated how the phenotypes of the B cells vary as a function of the distances from the prime clonotypes on the BCR trajectories. We first examined the pseudotimes of all the BCR clonotypes in each network, and observed that group 1 BCR clonotypes have the smallest pseudotimes, and more importantly that these three groups of BCR clonotypes formed a gradient with group 2 clonotypes having larger pseudotimes than group 1 clonotypes and group 3 clonotypes having larger pseudotimes than group 2 (Fig. 2c).
We also delineated specifically which part of the transcriptome is associated with the trajectories of the BCR networks, by examining a gene signature of B-cell activation (Methods). We again observed a gradient of B-cell activation among the three groups, with group 1 BCR clonotypes’ B cells having the highest level of activation (Fig. 2d). On the other hand, we performed an unbiased search, using gene set enrichment analyses (GSEAs)31, for the pathways enriched in the genes that showed a monotonic increase or decrease of expression from group 1 to group 3 in each dataset (Fig. 2e). We observed strong enrichment of pathways relevant for B-cell activation (for example, ‘Naive_vs_Memory_Bcell_DN’). We made similar observations with all other datasets that we investigated (Extended Data Fig. 4). Concomitant with the variation in transcriptomic phenotypes, group 1 BCR clonotypes also have larger clonal sizes than group 2 clonotypes, which in turn have larger clonal sizes than group 3 (Fig. 2f).
In the GCs, a process called class switching happens, which is independent of but related to BCR somatic hypermutations32,33,34. Class switching changes a B cell’s production of immunoglobulin from IgM and IgD to IgG, IgA and IgE during B-cell maturation. In Fig. 2g (the 10x melanoma dataset), we calculated the probability of class switching for the B cells belonging to each of groups 1–3. Interestingly, we observed that the group 1 BCRs have the lowest proportions of IgM/D and highest rate of class switching, followed by group 2 BCRs and then group 3 BCRs. We performed this analysis for all scRNA-seq datasets (Fig. 2h), and observed that group 1 BCRs do indeed have the highest rates of class switching (t-test P value of IgD + M% = 0.0034 for group 1 versus group 2, 0.00035 for 1 versus 3, 0.35 for 2 versus 3). Overall, we showed that Benisse is capable of defining functionally relevant trajectories of BCR evolution, which revealed how somatic hypermutations generate increasingly ‘better’ BCRs that confer stronger activation and maturation signals to their B cells.
Benisse reveals tighter coupling of BCR and B-cell expression during COVID-19
To demonstrate the capability of Benisse to reveal novel biological insights, we deployed Benisse to analyse a set of single-B-cell RNA-sequencing datasets with matched BCR sequencing from COVID-19 patients35. These data consist of 16,066 B cells from 11 patients and two healthy donors. We applied Benisse to the expression and BCR data of each patient’s B cells, constructed the BCR networks and detected the prime BCR clonotypes (Fig. 3a and Extended Data Fig. 5). We first validated that the detection of the BCR networks and the prime/non-prime BCR clonotypes is biologically meaningful, by leveraging the data of validated SARS-CoV-2 antibodies36 (N = 2,037). As we hypothesize that the prime BCRs are more likely to be antigen specific and will more strongly activate the B cells, we should expect the prime BCRs to be more likely to be the same as or at least very similar to these validated SARS-CoV-2 antibodies. Indeed, the prime BCRs are much more likely to be the same as one of the 2,037 validated antibodies or only differ by a small number of amino acids (from <2 to 5) from them, compared with the non-prime BCRs (Fig. 3b).
a, The BCR networks constructed by Benisse for patient Coc012. Principal component analyses were performed to reduce the dimensions of the embedding output by Benisse to two for visualization. The nodes in the same BCR networks were drawn in the same colours. b, The percentages of prime BCRs and non-prime BCRs, detected by Benisse, that are the same as one of the validated SARS-CoV-2 antibodies (edit distance <1) or differ only by a small number of amino-acid residues (edit distance <2 to <5). c, The number of BCR clonotypes found in the scRNA-seq data of each sample of each patient. d,e, The degrees of connection of the BCR clonotypes (number of other clonotypes in the same BCR network for each BCR clonotype) of B cells found in each stage of COVID-19: d, all BCR clonotypes; e, only prime BCR clonotypes of the networks. f, The probability of each BCR clonotype belonging to a BCR network of at least two member clonotypes, for B-cell clonotypes found in each stage of COVID-19. g, The correlation between B-cell gene expression and BCR embeddings in the latent space, for B cells belonging to the severe/recovery/cured phases of the COVID-19 patients and the healthy donors. h, Circos plot showing the enriched pathways in the genes whose expression was highly correlated with the BCRs in each stage of COVID-19. The top 30 pathways in each category are shown. However, the false discovery rates for these 30 pathways are all <1 × 10−9. The curves in the centre connect pathways that share genes, with darker colouring of the curves referring to more shared genes. The sizes of the dots on the circle refer to the number of highly correlated genes in each pathway divided by all genes in the pathway. The colouring of these dots refers to a pathway activation score calculated by the mean of expression of all genes in each pathway. mRNA, messenger RNA.
We next examined the topology of the BCR networks constructed by Benisse, which could reflect the activity of the somatic mutation events during BCR clonotype generation. For the diseased patients, the B cells were collected from one or more of the severe, recovery and cured phases of COVID-19 (definition in Methods, Fig. 3c). We counted the number of connections (degree) each BCR clonotype makes within the networks (Fig. 3d), in each of three disease phases and the healthy donors. We also narrowed down the investigation to the prime BCR clonotypes of each network and examined the degrees of connection of only these BCR clonotypes (Fig. 3e). Interestingly, we observed that the B-cell clonotypes from all phases of the COVID-19 patients demonstrate more connections than the B-cell clonotypes from the two healthy controls (Fig. 3d,e), indicating more active affinity maturation of BCRs during COVID-19. Next, examining the different phases of COVID-19, we observed that, overall, B cells from the severe phases demonstrate the highest level of connections, followed by the recovery and cured phases (Fig. 3d,e). To further confirm this observation, we also calculated another index, by examining each B-cell clonotype from each patient sample and calculating the probability that it is part of a BCR network with at least one other member clonotype. Again, this index is highest in the severe phase of COVID-19, followed by the recovery and cured phases, which are all higher than the control samples (Fig. 3f). The maturation of the BCR repertoire should be characterized by successive iterations of somatic hypermutations that generate well connected networks of BCRs.
We sought to further investigate whether the expression of the B cells demonstrates any concordant changes with BCRs during COVID-19. We calculated the correlations between BCR distances of all pairwise B-cell clonotype pairs and the expressional differences of the same clonotype pairs from the Benisse networks. Again we observed that this coupling between expression and BCR is stronger in all phases of COVID-19 compared with the healthy controls (Fig. 3g). We also again observed that the correlation between BCR repertoire and B-cell gene expression is strongest in the severe phase, followed by the recovery phase and then the cured phase (Fig. 3g). We also calculated the correlation between BCR and gene expression for each individual gene. We selected the top 2% of all investigated genes (N = 452) with the highest average correlation, from B cells of all datasets of the severe phases of COVID-19. We removed from this set the 177 genes that are also highly correlated with BCRs in B cells of the healthy samples (same criterion). We also did the same for the genes from the recovery and cured phases, and performed Gene Ontology (GO) analyses for remaining genes. For all three phases, we observed many GO terms associated with immune-cell functions, especially the activation of humoral immunity (Fig. 3h). The severe phase has the highest level of pathway activation, followed by the recovering phase and then the cured phase. The pathways in the severe phase also have more genes in common with one another. These observations indicate the highest level of concerted action of the B cells during the severe phase of COVID-19.
Taken together, our analyses indicate that the BCR signalling pathway is most activated and induces the strongest BCR rearrangement events in earlier severe phases of COVID-19, and weakens when the patients are on the pathway to recovery.
Discussion
In this work, we developed the Benisse model to build the link between expression of B-cell genes and their BCRs. The fields of BCR sequencing and scRNA-seq have been developing as independent disciplines. Our work models the two types of data together, providing new opportunities to mechanistically dissect the roles of B cells and BCRs in normal development and disease progression. Methodologically, Benisse is built upon mathematically innovative techniques that enable learning a sparse weighted graph in the latent space probabilistically from the high-dimensional data of BCRs, under the supervision of gene expression37. We applied Benisse to two human diseases, COVID-19 (Fig. 3) and ulcerative colitis (Supplementary Note 2), which revealed interesting biological insights.
Zhang et al.38 and Yost et al.39 discovered convergent VDJ (D, diversity) recombinations for T-cell receptors (TCRs). These works found that T cells tend to generate different but similar TCRs through VDJ recombination against the same antigen. The TCRs that are most similar to the ‘average/centre’ of these clustered TCRs are probably the most efficient in antigen targeting. In an interesting contrast to T cells, we observed that centre BCRs do not have higher activation signals or larger clonal expansion than non-centre BCRs (Supplementary Note 2). In other words, the prime BCR clonotypes that we identified above in each BCR network are not the centre of their BCR networks. Rather, BCRs of B cells continuously evolve, via somatic hypermutation, to obtain higher specificity in antigen targeting, forming a directional trajectory. This is most obvious in Extended Data Fig. 3, where the phylogenetic tree of BCRs has only one major branch of evolution, pointing towards the best antibody, placed at the tip of the tree branch. This is different from T cells, which have very limited somatic hypermutation ability40,41, so they tend to converge to the best solution from different independent trials of VDJ recombinations.
We observed the coupling between BCRs and B-cell expression in various biomedical contexts. There are two possible mechanisms responsible for this coupling effect. First, in the GCs, the somatic mutational processes keep producing BCRs of various qualities in a continuous process. BCRs that create higher-affinity antibodies for the target antigens will induce stronger proliferation signals for the B cells (the well known positive selection process). Similar BCRs will probably induce similar proliferative signals for the B cells. Second, some plasma cells still express functional BCRs that can positively impact the survival and activation of plasma cells42,43. Therefore, even in the stage of plasma cells, BCRs can still constitutively modulate B-cell phenotypes depending on the qualities of the BCRs, thereby explaining this coupling effect.
Overall, the BCR repertoire should not be considered alone as a marker of clonality, and the interrogation of how the BCRs interact with and impact the transcriptomic status of the B cells will facilitate the discovery of vital insights. We expect Benisse and similar works to propell our understanding of the function of B cells in various physiological processes.
Methods
The BCR embedding algorithm
We created an algorithm based on deep contrastive learning for embedding the BCR CDR3H amino-acid sequences with numeric vectors. We first encoded the BCR CDR3H sequences using the Atchley factors, which represent each amino acid with five numeric values, and can comprehensively characterize the biochemical properties of each amino acid22,38. In our recent work on TCRs38, we also created a similar embedding model for TCR CDR3β (complementarity-determining region 3β) sequences with Atchley factors and demonstrated the appropriateness of Atchley factors for numerically embedding immune receptors. Next, we leveraged contrastive multiview coding (CMC44) to build a short numerical vector embedding of BCR CDR3H sequences in the space of Atchley matrices for easy manipulations in the following model. To achieve this, we leverage the fact that, for most CDR3H protein sequences we obtained from various public sources, the nucleotide sequences for the BCR sequences are also available. We treat the protein sequences (in the space of Atchley factor matrices) as one view and their corresponding nucleotide sequences (A/T/G/C are one-hot encoded) as another view—both matching the same underlying CDR3H sequence. Then we developed a CMC model that is capable of pairing CDR3H nucleotide sequences with the correct CDR3H protein sequences from a pool, and also vice versa: namely, the loss function considers the loss in both directions. In this process, the CMC model learns a latent embedding of CDR3H in the form of a short numeric vector.
This approach is different from a simplistic protein translation between DNA and protein sequences from several perspectives. First, the codon table is not given as a model input, but rather we require the model to implicitly learn this matching relationship from the given numerical vectors. Thus the model also has to learn more information regarding the key inherent properties of the nucleotide and protein sequences of CDR3H at the same time. Second, the goal of this CMC model is to obtain the numeric embedding, which is a short fixed-length vector. The correct pairing between CDR3H protein sequences and nucleotide sequences forces the CMC model to capture the key structural information of BCRs through this short vector, rather than only learning a simple matching relationship between individual amino acids and triplets of nucleotides (the codon table).
The core Benisse model
The core Benisse model deals with a pool of B cells where, for each cell we have (1) expression data, as a numeric vector, and (2) BCR data, already encoded into a numeric vector as described above. In the pool of B cells we analyse, there are groups of B cells that are in the same BCR clonotype (same V, same J and same CDR3 for the H chain), and their expression is averaged to the clonotype level. Benisse is tasked with detecting BCR clonotypes that are related (similar sequences, probably generated for the same antigen) and building a graph of BCR networks, with each network composed of related BCRs. Within each network, the similarity between different BCR clonotypes should be influenced by the similarity of their clonotype-level expression profiles. To achieve this, Benisse finds a lower-dimensional embedding based upon the original BCR embeddings that also satisfies the coupling between BCRs and expression. Moreover, Benisse concomitantly detects the graph of BCR networks in this lower-dimensional space.
Technically, the core Benisse model is based on a probabilistic supervised learning model of sparse graph structure for BCR data, informed by the expression of the B cells. The graph is derived from a unified model of density estimation and pairwise distance preservation, where latent embeddings are assumed to be random variables following an unknown density function to be learned, and pairwise distances are then calculated as the expectations over the density for the BCR clonotypes. The problem is solved using the alternating direction method of multipliers45 algorithm in two main alternating steps: (1) spectral projection for solving the latent space embedding of BCRs supervised by gene expression and (2) L-BFGS-B46 for solving the graph of BCR networks. Mathematical details of the Benisse model are described in Supplementary Note 1, along with simulation analyses.
Acquisition and preprocessing of the BCR-seq and scRNA-seq data
The processed BCR-seq data used for the training and validation of the BCR encoder are obtained from bulk BCR sequencing and also from scRNA sequencing of B cells with paired scBCR-seq. For the training/validation of the BCR encoder, we included data from several sources, including IEDB (https://www.iedb.org/database_export_v3.php), AdaptiveBiotech47,48,49,50,51,52,53,54,55,56,57,58,59 and Rizzetto et al.60. We also included all BCRs from all the scBCR-seq datasets used in this study, including 10x Genomics, and refs. 24,35,61,62,63,64,65. For the 10x single-cell data, we only included BCR records with ‘is_cell’, ‘high_confidence’, ‘full_length’ and ‘productive’ all being true. The class-switching status of each B cell in the scRNA-seq data was inferred using the standard 10x Cell Ranger software.
For the analyses involving the Benisse model, we only investigated cells from the scRNA-seq data that have both expression information and also the BCR heavy-chain information. The scRNA-seq data were handled by the Seurat R package (v4.0.5)66. For the single-cell expression matrices, we aggregated the expression of the transcripts to the gene levels (HUGO gene symbols) via the annotation files provided with the R biomaRt package (v2.48.0). We normalized by library size of each cell and performed log(x + 1) transformation before downstream analyses.
Implementation of bcRep and Lindenbaum cosine similarity
bcRep was implemented using the bcRep R package (v1.3.6), based on the cosine dissimilarity method, which was showcased in Fig. 6 of bcRep’s original publication. For the Lindenbaum et al method, we calculated the tf–idf (term frequency–inverse document frequency) representations (the TfidfVectorizer function from the Python sklearn package, v1.0) and cosine similarities between BCR sequences, as described in their original paper. The cosine dissimilarity used to form the tree structures and waterfall plots in Extended Data Fig. 3e,f was calculated by subtracting the min max normalized cosine similarity score (MinMaxScaler from the Python sklearn package, v1.0) from 1.
Defining the different phases of COVID-19 for the Bernardes et al. dataset
We assigned our four disease groups on the basis of the classification of Bernardes et al.35, but slightly consolidated their different categories of disease trajectories. This is done so that we have fewer groups but more samples in each of the four groups (for more robust analysis results), and we also have more balanced numbers of B cells in each category. Their ‘uninfected (control)’ samples were denoted as ‘healthy’ in our study. Their ‘incremental’ and ‘critical’ samples were denoted as ‘severe’, their ‘complicated’ and ‘early/moderate/late convalescence’ samples were denoted as ‘recovering’ and their ‘long-term follow-up’ samples were denoted as ‘cured’. In particular, all the incremental samples were taken within two days of the patients entering the critical phase, including one patient who died. Therefore, we grouped incremental and critical samples together.
Statistical analyses
All computations are performed in the R (v4.02) and Python (v3.7) languages. The BCR embedding is created using the PyTorch package. PyTorch version, 1.10.0; pandas version, 1.3.4; NumPy version, 1.21.3; sklearn version, 1.0. UMAP was performed using the R umap package (v0.2.7.0) and t-SNE was performed using the R Rtsne package (v0.15). Pseudotime inference was performed using Monocle230. The predicted pseudotimes were reordered in each dataset so that the latest-appearing cells have the smallest pseudotimes. For all boxplots appearing in this Article, box boundaries represent interquartile ranges, whiskers extend to the most extreme data point, which is no more than 1.5 times the interquartile range, and the line in the middle of the box represents the median. We assembled a B-cell activation gene signature (Fig. 2d) on the basis of literature review and availability of the genes in the scRNA-seq datasets of this study (Extended Data Table 2). GSEA analyses were performed using clusterProfiler v3.1467. The GOrilla webserver (v2013Mar8) was used to detect enriched GO pathways68. The CytoSig analyses were performed with the CytoSig software downloaded from https://cytosig.ccr.cancer.gov/, using all default settings. Canonical correlation analysis was performed using the R CCA package (v1.2.1). All statistical tests were two tailed, unless otherwise specified. All correlations are Pearson correlations unless otherwise specified.
Data availability
The data used for the training of the BCR embedding algorithm and the single-cell sequencing data used for Benisse analyses are all publicly available. Their accession methods are shown in Extended Data Table 124,35,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65. The LIBRA-seq data are publicly available from Setliff et al.24. The SARS-CoV-2 antibody data are from Raybould et al.36. Our own in-house BCR data can be found in our GitHub repository at https://github.com/wooyongc/Benisse.
Code availability
The Benisse software, including the BCR embedding algorithm, is publicly available under the MIT License at https://github.com/wooyongc/Benisse and in the Supplementary Software. The software is also available via Zenodo69.
References
Berzofsky, J. A. T–B reciprocity. An Ia-restricted epitope-specific circuit regulating T cell–B cell interaction and antibody specificity. Surv. Immunol. Res. 2, 223–229 (1983).
Ozaki, S. & Berzofsky, J. A. Antibody conjugates mimic specific B cell presentation of antigen: relationship between T and B cell specificity. J. Immunol. 138, 4133–4142 (1987).
Sabhnani, L. et al. Developing subunit immunogens using B and T cell epitopes and their constructs derived from the F1 antigen of Yersinia pestis using novel delivery vehicles. FEMS Immunol. Med. Microbiol. 38, 215–229 (2003).
Zhang, J. et al. Modulation of nonneutralizing HIV-1 gp41 responses by an MHC-restricted TH epitope overlapping those of membrane proximal external region broadly neutralizing antibodies. J. Immunol. 192, 1693–1706 (2014).
Zhu, J. et al. BepiTBR: T–B reciprocity enhances B cell epitope prediction. iScience 25, 103764 (2022).
Greaney, A. J. et al. Comprehensive mapping of mutations to the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human plasma antibodies. Cell Host Microbe 29, 463–476 (2021).
Oh, H.-L. J., Gan, S. K.-E., Bertoletti, A. & Tan, Y.-J. Understanding the T cell immune response in SARS coronavirus infection. Emerg. Microbes Infect. 1, e23 (2012).
Wang, S.-S. et al. Tumor-infiltrating B cells: their role and application in anti-tumor immunity in lung cancer. Cell. Mol. Immunol. 16, 6–18 (2019).
Garaud, S. et al. Tumor infiltrating B-cells signal functional humoral immune responses in breast cancer. JCI Insight 4, e129641 (2019).
Chiaruttini, G. et al. B cells and the humoral response in melanoma: the overlooked players of the tumor microenvironment. Oncoimmunology 6, e1294296 (2017).
Lechner, A. et al. Tumor-associated B cells and humoral immune response in head and neck squamous cell carcinoma. Oncoimmunology 8, 1535293 (2019).
Carmi, Y. et al. Allogeneic IgG combined with dendritic cell stimuli induce antitumour T-cell immunity. Nature 521, 99–104 (2015).
Burger, J. A. & Wiestner, A. Targeting B cell receptor signalling in cancer: preclinical and clinical advances. Nat. Rev. Cancer 18, 148–167 (2018).
Leong, T. L. & Bryant, V. L. B cells in lung cancer—not just a bystander cell: a literature review. Transl. Lung Cancer Res. 10, 2830–2841 (2021).
Zhang, L. et al. Long-term sculpting of the B-cell repertoire following cancer immunotherapy in patients treated with sipuleucel-T. Cancer Immunol. Res. 8, 1496–1507 (2020).
Chen, H. et al. BCR selection and affinity maturation in Peyer’s patch germinal centres. Nature 582, 421–425 (2020).
Miho, E., Roškar, R., Greiff, V. & Reddy, S. T. Large-scale network analysis reveals the sequence space architecture of antibody repertoires. Nat. Commun. 10, 1321 (2019).
Wu, X. et al. Maturation and diversity of the VRC01-antibody lineage over 15 years of chronic HIV-1 infection. Cell 161, 470–485 (2015).
Bashford-Rogers, R. J. M. et al. Analysis of the B cell receptor repertoire in six immune-mediated diseases. Nature 574, 122–126 (2019).
Khavrutskii, I. V. et al. Quantitative analysis of repertoire-scale immunoglobulin properties in vaccine-induced B-cell responses. Front. Immunol. 8, 910 (2017).
Bolen, C. R., Rubelt, F., Vander Heiden, J. A. & Davis, M. M. The Repertoire Dissimilarity Index as a method to compare lymphocyte receptor repertoires. BMC Bioinform. 18, 155 (2017).
Atchley, W. R., Zhao, J., Fernandes, A. D. & Drüke, T. Solving the protein sequence metric problem. Proc. Natl Acad. Sci. USA 102, 6395–6400 (2005).
Chen, T., Kornblith, S., Norouzi, M. & Hinton, G. A simple framework for contrastive learning of visual representations. In Proc. 37th International Conference on Machine Learning Vol. 119 (eds Hal Daumé, I. I. I. & Singh, A.) 1597–1607, 2020 (PMLR).
Setliff, I. et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell 179, 1636–1646.e15 (2019).
Lindenbaum, O., Nouri, N., Kluger, Y. & Kleinstein, S. H. Alignment free identification of clones in B cell receptor repertoires. Nucleic Acids Res. 49, e21 (2021).
Bischof, J. & Ibrahim, S. M. bcRep: R package for comprehensive analysis of B cell receptor repertoire data. PLoS ONE 11, e0161569 (2016).
Liao, H.-X. et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496, 469–476 (2013).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Neuberger, M. S. & Milstein, C. Somatic hypermutation. Curr. Opin. Immunol. 7, 248–254 (1995).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Nagumo, H. et al. The different process of class switching and somatic hypermutation; a novel analysis by CD27− naive B cells. Blood 99, 567–575 (2002).
Petrova, V. N. et al. Combined influence of B-cell receptor rearrangement and somatic hypermutation on B-cell class-switch fate in health and in chronic lymphocytic leukemia. Front. Immunol. 9, 1784 (2018).
Chi, X., Li, Y. & Qiu, X. V(D)J recombination, somatic hypermutation and class switch recombination of immunoglobulins: mechanism and regulation. Immunology 160, 233–247 (2020).
Bernardes, J. P. et al. Longitudinal multi-omics analyses identify responses of megakaryocytes, erythroid cells, and plasmablasts as hallmarks of severe COVID-19. Immunity 53, 1296–1314.e9 (2020).
Raybould, M. I. J., Kovaltsuk, A., Marks, C. & Deane, C. M. CoV-AbDab: the coronavirus antibody database. Bioinformatics 37, 734–735 (2021).
Wang, L., Chan, R. & Zeng, T. Probabilistic semi-supervised learning via sparse graph structure learning. IEEE Trans. Neural Netw. Learn. Syst. 32, 853–867 (2021).
Zhang, Z., Xiong, D., Wang, X., Liu, H. & Wang, T. Mapping the functional landscape of T cell receptor repertoires by single-T cell transcriptomics. Nat. Methods 18, 92–99 (2021).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
Ott, J. A. et al. Somatic hypermutation of T cell receptor α chain contributes to selection in nurse shark thymus. eLife 7, e28477 (2018).
Bilal, S., Lie, K. K., Sæle, Ø. & Hordvik, I. T cell receptor alpha chain genes in the teleost Ballan wrasse (Labrus bergylta) are subjected to somatic hypermutation. Front. Immunol. 9, 1101 (2018).
Pinto, D. et al. A functional BCR in human IgA and IgM plasma cells. Blood 121, 4110–4114 (2013).
Blanc, P. et al. Mature IgM-expressing plasma cells sense antigen and develop competence for cytokine production upon antigenic challenge. Nat. Commun. 7, 13600 (2016).
Tian, Y., Krishnan, D. & Isola, P. Contrastive Multiview Coding in Computer Vision – ECCV 2020: 16th European Conference, Glasgow, UK, August 23–28, 2020, Proceedings, Part XI 776–794 (Springer-Verlag, Glasgow, United Kingdom, 2020).
Boyd, S., Parikh, N., Chu, E., Peleato, B. & Eckstein, J. Distributed optimization and statistical learning via the alternating direction method of multipliers. Found. Trends Mach. Learning 3, 1–122 (2010).
Byrd, R. H., Lu, P., Nocedal, J. & Zhu, C. A limited memory algorithm for bound constrained optimization. SIAM J. Sci. Comput. 16, 1190–1208 (1995).
Bertoli, D. et al. Lack of specific T- and B-cell clonal expansions in multiple sclerosis patients with progressive multifocal leukoencephalopathy. Sci. Rep. 9, 16605 (2019).
Cui, Y. et al. Combined immunodeficiency caused by a loss-of-function mutation in DNA polymerase delta 1. J. Allergy Clin. Immunol. 145, 391–401.e8 (2020).
Ahmed, R. et al. A public BCR present in a unique dual-receptor-expressing lymphocyte from type 1 diabetes patients encodes a potent T cell autoantigen. Cell 177, 1583–1599.e16 (2019).
Ruella, M. et al. Induction of resistance to chimeric antigen receptor T cell therapy by transduction of a single leukemic B cell. Nat. Med. 24, 1499–1503 (2018).
Fraietta, J. A. et al. Disruption of TET2 promotes the therapeutic efficacy of CD19-targeted T cells. Nature 558, 307–312 (2018).
Allenspach, E. J. et al. Absence of functional fetal regulatory T cells in humans causes in utero organ-specific autoimmunity. J. Allergy Clin. Immunol. 140, 616–619.e7 (2017).
Lombardo, K. A. et al. High-throughput sequencing of the B-cell receptor in African Burkitt lymphoma reveals clues to pathogenesis. Blood Adv. 1, 535–544 (2017).
Seay, H. R. et al. Tissue distribution and clonal diversity of the T and B cell repertoire in type 1 diabetes. JCI Insight 1, e88242 (2016).
Boisvert, M. et al. Novel E2 glycoprotein tetramer detects hepatitis C virus-specific memory B cells. J. Immunol. 197, 4848–4858 (2016).
DeWitt, W. S. et al. A public database of memory and naive B-cell receptor sequences. PLoS ONE 11, e0160853 (2016).
Kanakry, C. G. et al. Origin and evolution of the T cell repertoire after posttransplantation cyclophosphamide. JCI Insight 1, e86252 (2016).
Mattoo, H. et al. De novo oligoclonal expansions of circulating plasmablasts in active and relapsing IgG4-related disease. J. Allergy Clin. Immunol. 134, 679–687 (2014).
Kolhatkar, N. S. et al. Altered BCR and TLR signals promote enhanced positive selection of autoreactive transitional B cells in Wiskott–Aldrich syndrome. J. Exp. Med. 212, 1663–1677 (2015).
Rizzetto, S. et al. B-cell receptor reconstruction from single-cell RNA-seq with VDJPuzzle. Bioinformatics 34, 2846–2847 (2018).
Lu, B. et al. IL-17 production by tissue-resident MAIT cells is locally induced in children with pneumonia. Mucosal Immunol. 13, 824–835 (2020).
Attaf, N. et al. FB5P-seq: FACS-based 5-prime end single-cell RNA-seq for integrative analysis of transcriptome and antigen receptor repertoire in B and T cells. Front. Immunol. 11, 216 (2020).
Durante, M. A. et al. Single-cell analysis reveals new evolutionary complexity in uveal melanoma. Nat. Commun. 11, 496 (2020).
Boland, B. S. et al. Heterogeneity and clonal relationships of adaptive immune cells in ulcerative colitis revealed by single-cell analyses. Sci. Immunol. 5, eabb4432 (2020).
Wang, T. et al. An empirical approach leveraging tumorgrafts to dissect the tumor microenvironment in renal cell carcinoma identifies missing link to prognostic inflammatory factors. Cancer Discov. 8, 1142–1155 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinform. 10, 48 (2009).
Chang, W. Y. & Wang, T. wooyongc/Benisse: v1.0.0. Zenodo https://doi.org/10.5281/zenodo.6113441 (2022).
Acknowledgements
This paper is dedicated to Z.Z. (also known as Briseis), and the method named ‘Benisse’ in her honour.
Funding
This study was supported by the National Institutes of Health (NIH) (5P30CA142543/TW, R01CA258584/TW, XW, U01AI156189/TW, 1DP2AI154450/CY), Cancer Prevention Research Institute of Texas (RP190208/TW, XW, RR210035/CY) and National Science Foundation (DMS-2009689/LW).
Author information
Authors and Affiliations
Contributions
Z.Z. and W.Y.C. performed all analyses. T.Wang supervised the study. L.W. contributed the mathematical model. K.W., Y.Y. and X.W. provided input on the modelling and downstream analyses. All authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Lili Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic diagram of the contrastive deep learning model to embed the BCR CDR3H sequences into numeric vectors.
The model is tasked to distinguish n different CDR3H protein sequences (n = 2 under our setting) to identify the correct one corresponding to the given BCR DNA sequence, and also vice versa. This way, the key features of the CDR3H amino acid sequences are captured on a short numeric vector of 20 elements. “Affinity score” refers to how close the pool of BCR CDR3H protein sequences are to the correct nucleotide sequence in the deep learning model’s latent space, and vice versa.
Extended Data Fig. 2
Scatterplots of these two distances for BCR distances determined by our contrastive learning model.
Extended Data Fig. 3 Tracking clonal evolution of BCRs in an HIV-infected patient.
BCRs were embedded by (a,b) the Benisse BCR encoder (the deep learning only part); (c,d) bcRep; and (e,f) the Lindenbaum method. (a,c,e) A phylogenetic tree of BCRs was constructed from the distance matrices of BCRs calculated according to the BCR embeddings. The trees were rooted at the Unmutated Common Ancestor (UCA) of the BCRs. (b,d,f) A waterfall plot of BCR Euclidean distances between the UCA BCR and all the other BCRs, colored by their collection times.
Extended Data Fig. 4 GSEA of the pathways enriched in the genes that showed a monotonic increase (a) or decrease (b) of expression from group 1 to group 3 BCR clonotypes’ B cells.
Pathways enriched in the genes with “increasing” or “decreasing” trends were shown separately. The results for all datasets are shown, with the top 5 pathways of the smallest False Discovery Rates (FDRs) shown for each dataset. The red circles denote the FDR cutoffs of 5% and 10%.
Extended Data Fig. 5 The BCR networks constructed by Benisse for patient Coc003.
Principal Component Analyses were performed to reduce the dimensions of the embedding output by Benisse to 2 dimensions for visualization.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2.
Supplementary Software
The Benisse software.
Rights and permissions
About this article
Cite this article
Zhang, Z., Chang, W.Y., Wang, K. et al. Interpreting the B-cell receptor repertoire with single-cell gene expression using Benisse. Nat Mach Intell 4, 596–604 (2022). https://doi.org/10.1038/s42256-022-00492-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-022-00492-6
This article is cited by
-
Integrating single-cell multi-omics and prior biological knowledge for a functional characterization of the immune system
Nature Immunology (2024)
-
Adaptive immune receptor repertoire analysis
Nature Reviews Methods Primers (2024)
-
Single-cell immune repertoire analysis
Nature Methods (2024)
-
Best practices for single-cell analysis across modalities
Nature Reviews Genetics (2023)