Abstract
Molecular simulations are a powerful tool to complement and interpret ambiguous experimental data on biomolecules to obtain structural models. Such data-assisted simulations often rely on parameters, the choice of which is highly non-trivial and crucial to performance. The key challenge is weighting experimental information with respect to the underlying physical model. We introduce FLAPS, a self-adapting variant of dynamic particle swarm optimization, to overcome this parameter selection problem. FLAPS is suited for the optimization of composite objective functions that depend on both the optimization parameters and additional, a priori unknown weighting parameters, which substantially influence the search-space topology. These weighting parameters are learned at runtime, yielding a dynamically evolving and iteratively refined search-space topology. As a practical example, we show how FLAPS can be used to find functional parameters for small-angle X-ray scattering-guided protein simulations.
Similar content being viewed by others
Main
Proteins are the molecular workhouses of biological cells, with a myriad of tasks including oxygen transport, cellular communication and energy balance. As a protein’s function is linked to its structure and dynamics, its understanding requires resolving the protein’s three-dimensional shape. Misfolded proteins are associated with several neurodegenerative diseases1, and deciphering the structure–function paradigm paves the way to developing treatments for Alzheimer’s and Parkinson’s diseases2, amyloidosis3, type-2 diabetes4, Creutzfeldt–Jakob disease5, among others.
Proteins are nanoscale and can only be observed indirectly. Hence, experimental data are often ambiguous, incomplete or of such low resolution that they require interpretation to access their information content. A practical example is small-angle X-ray scattering (SAXS), where dissolved biomolecules are irradiated by X-rays and the scattering intensity is recorded. However, the desired molecular electron density is the Fourier transform of the experimentally inaccessible complex-valued scattering amplitude. Recovering structural models from an intensity, that is, the absolute amplitude squared, is an ill-posed inverse problem. The sparse information in the SAXS data is insufficient to determine all degrees of freedom in a molecular structure. Protein structure determination thus depends on combining experimental results and computational methods, and recent studies highlight the potential of such hybrid approaches6,7,8,9.
One effective approach is to complement the experimental data with molecular dynamics (MD)10,11. MD simulations provide a physics-based description of molecular motion and give in-depth insight into biomolecular function. Atomistic trajectories are derived by integrating Newton’s equations of motion for a system of interacting particles, and forces are calculated from empirical interatomic potentials (force fields). In data-assisted MD, an energetic restraint on the target data is added to the force field to favour conformations consistent with the data. This bias potential is proportional to the least-squares deviation of theoretical data from simulated structures and the experimental data7,10,12,13,14. Derived forces are assumed to drive the protein towards conformations reproducing the target data. As a molecular system seeks to minimize its free energy, the bias effectively determines a cost for disregarding the data in the simulation. The better the simulated structures align to the data, the smaller is the energetic penalty in the form of the bias15.
An inherent issue with data-assisted simulations is their reliance on MD parameters7,8,9. Selecting adequate values is non-trivial and crucial for simulation performance, and determining the bias potential’s weight is the key challenge. This selection determines how experimental and theoretical information is balanced. The bias weight is an empirical MD parameter expressing the confidence in the experimental data versus the physics-based force field8. In Bayesian methods, the right weighting is derived from a statistical treatment13,14. However, such sophisticated approaches are practically inapplicable for users with a primarily experimental background. It is still common practice to manually determine an ‘optimal’ bias weight via grid search, that is, an exhaustive search through a fixed subset of the parameter space7,9,12. Adopting concepts from computational intelligence, we introduce FLAPS (‘flexible self-adapting particle swarm’ optimization), a self-learning metaheuristic based on particle swarms, to resolve this parameter selection problem. Our contributions include the following:
-
A new type of flexible objective function (OF) to assess a data-assisted simulation’s plausibility in terms of simulated structures and thus the suitability of the MD parameters used.
-
A self-adapting particle swarm optimizer for dynamically evolving environments resulting from multiple quality features of different scales in the flexible OF.
-
Fully integrated and automated MD parameter selection for data-assisted biomolecular simulations.
As a proof of concept, we apply FLAPS to the selection of relevant MD parameters in SAXS-guided structure-based protein simulations7.
Particle swarm optimization
Particle swarm optimization (PSO) is a bio-inspired computational-intelligence technique to handle computationally hard problems based on the emergent behaviour of swarms16. Swarm intelligence is the collective behaviour of decentralized, self-organized systems. Such systems comprise a population of cooperating particles. Although there is no supervising control, this leads to ‘intelligent’ global behaviour that is hidden to the individual particles.
PSO iteratively improves a candidate solution with respect to a quality-gauging OF. The optimization problem is approached by considering a swarm of particles, each corresponding to a particular position in parameter space. Across multiple search rounds (generations), the particles ‘jump’ around in the search space according to their positions and velocities. Particles ‘remember’ their personal best location and the global best location in their social network, which act as attractors in the search space. To propagate a particle to the next generation, a velocity is added to its current position. The velocity has a cognitive component towards the particle’s current personal best and a social component towards the current best in its communication network. Each component is weighted by a random number in the range \(\left[0,{\phi }_{i}\right]\), where the acceleration coefficients ϕi balance exploration versus exploitation in the optimization. Eventual convergence is achieved through progressive particle flight contraction by mechanisms such as velocity clamping17, inertia weight18 or constriction19. Altogether, this is assumed to move the swarm towards the best parameter combination.
PSO has been demonstrated to work well for various static problems20. However, realistic problems are often dynamic, and the global optimum can shift with time. Application to such problems has been explored extensively, revealing that the algorithm needs to be enhanced by concepts such as repulsion, dynamic network topologies or multi-swarms21,22.
Maintaining a population of diverse solutions enables enormous exploration and paves the way to large-scale parallelization. Such algorithms can be scaled easily to exploit the full potential of modern supercomputers23. PSO has several hyperparameters affecting its behaviour and efficiency, and selecting them has been researched extensively24,25. Strategies include using meta-optimizers26,27 or refining them during the optimization28. Reference 29 provides an overview of practical PSO applications, while refs.22 and30 give comprehensive reviews of PSO with a focus on dynamic environments and hybridization perspectives, respectively.
Multi-response problems
Real-life optimization problems are often determined by multiple incomparable or conflicting quality features (responses). To obtain compatible solutions, these responses must be taken into account simultaneously. This is often accomplished by combining individual contributions within one composite OF. PSO has been applied to multi-response optimization in many fields31,32,33,34. Commonly, the set of responses is reduced via multiplication by manually chosen weights, henceforth referred to as OF parameters. Choosing OF parameters is non-trivial, yet strongly impacts global optimization performance by skewing the OF. FLAPS builds on a flexible OF that automatically and interdependently balances different responses. OF parameters are learned at runtime through iterative refinement, yielding a dynamically evolving OF landscape. In this way, FLAPS can cope with various responses of different scales.
A flexible self-adapting objective function
Typically, the set of responses is mapped to a scalar score by calculating the scalar product with fixed weights. These OF parameters supposedly reflect relative importance, implicitly encoding arbitrary prior beliefs. Instead, we set up a ‘maximum-entropy’ OF with the fewest possible assumptions:
where z is the set of OF parameters, μj is the mean and σj is the standard deviation of response Rj for a particle at position x. All responses are considered equally important but can have different ranges and units. To make them comparable on a shared scale, we standardize each response’s set of values gathered over previous generations. This strategy imitates the concept of rolling batch normalization35. Each layer’s inputs are recentred and rescaled with the aim to improve a neural network’s speed, performance and stability. Initially proposed to mitigate internal covariate shift, batch normalization is believed to introduce a regularizing and smoothing effect and promote robustness with respect to different initialization schemes. The OF in equation (1) depends not only on the parameters of interest, x, in our case the MD parameters, but also on a priori unknown, context-providing OF parameters, \({\bf{z}}=({\{\mu ,\sigma \}}_{j})\), from the standardization. Their values cannot be deduced from individual OF evaluations, yet fundamentally control OF performance and hence the optimization process.
Algorithm 1 FLAPS algorithm
Initialize population pop with swarm size S particles at random positions \({{\bf{x}}}_{p}\,\left(p=1,...,S\right)\) between upper and lower bounds of the search space bup and blo, respectively.
for g ← 1 to maximum generations G do
for particle in pop do
Evaluate responses at particle.position = xp:
particle.fargs = [responsej(xp)]j
end
Append current generation pop to history histp: histp.append(pop)
Update OF parameters zg based on current knowledge state of responses in histp: zg = updateParams(histp)
for particle in histp do
(Re-)evaluate objective function using most recent zg:
particle.fitness = f(xp; zg)
end
for generation in histp do
for particle in generation do
Determine personal best \({p}_{{\rm{best}}}^{p}\) and update global best gbest accordingly.
end
end
for particle in pop do
Update velocity and position:
\(\begin{array}{l} particle.speed+={\rm{rand}}\left(0,{\phi }_{1}\right)\left({p}_{{\rm{best}}}^{p}-particle.position\right)+\\{\rm{rand}}\left(0,{\phi }_{2}\right)\left({g}_{{\rm{best}}}-particle.position\right)\end{array}\)
Regulate velocity via \({{\bf{s}}}_{\max }=0.7\ {G}^{-1}\left({{\bf{b}}}_{{\rm{up}}}-{{\bf{b}}}_{{\rm{lo}}}\right)\):
if particle.speed > smax then
particle.speed = smax
end
if particle.speed < −smax then
particle.speed = −smax
end
particle.position+ = particle.speed
end
end
Result: gbest
Our self-adapting PSO variant, FLAPS, solves this problem. Provided with a comprehensive history of all previous particles and their responses, OF parameters are learned on the fly. They are continuously refined according to the current state of the optimization, yielding a dynamically evolving and increasingly distinct OF topology. This environmental dynamism may cause convergence problems if the OF fails to approach a stable topology. As more particles are evaluated, the ranges and distributions of individual responses become better understood. Therefore, OF parameters become more accurate, improving OF performance in assessing the suitability of the actual parameters of interest, x. After each generation, the values \({\bf{z}}=({\{\mu ,\sigma \}}_{j})\) are used to reevaluate the OF for all particles in the history. Personal best positions, \({p}_{\,\text{best}\,}^{p}\), and the swarm’s global best position, gbest, are updated accordingly for propagating particles to the next generation. FLAPS uses a traditional PSO velocity formulation16. Strategies to prevent diverging velocities include introducing an inertia weight18 or a constriction factor19. We regulate the velocities by means of a maximum value at each particle update17. FLAPS’s pseudo code is shown in Algorithm 1. Inspired by the ‘simplifying PSO’ paradigm, it builds on a slim standard PSO core and can easily be complemented by concepts such as inertia weight18 and swarm constriction19 or diversity increasing mechanisms. Its time complexity is similar to that of a standard PSO with \({\mathcal{O}}\left(\frac{S}{P}\cdot G\cdot {\rm{Sim}}+S\cdot {\rm{Opt}}\right)={\mathcal{O}}\left(\frac{S}{P}\cdot G\cdot {\rm{Sim}}+{S}^{2}\cdot G\right)\) in Landau notation, where P is the number of simulation processors, ‘Sim’ the maximal simulation time, and all other variables as defined in Algorithm 1.
Application to data-assisted protein simulations
We applied FLAPS to the optimization of MD parameters in SAXS-guided protein simulations7. SAXS data are integrated into computationally efficient structure-based models, which probe dynamics arising from a protein’s native geometry36,37. To assess the utility of different MD parameter sets, we need a metric for simulation quality in terms of physically reasonable structures matching the data. Designing such an OF in advance is non-trivial and has two major aspects: (1) physical plausibility of a simulated ensemble of protein structures and (2) its agreement with the target data, that is, how well the data are reproduced by simulated structures. To represent these aspects, we use the Rosetta energy function 2015 (REF15)38,39 and the least-squares deviation χ2 of simulated data from the target data7.
Protein structure determination relies on quick and reliable scoring of many models to select those closest to the native state. Structures are rated by energetic scores associated with their conformational state. REF15 is a weighted sum of energy terms efficiently approximating the energy of a biomolecular conformation as a function of geometric degrees of freedom and chemical identities38. With a protein’s native fold corresponding to the state with minimal free energy, a lower-scoring structure is expected to be more native-like. Because the scores do not have a direct conversion to physical energies, REF15 and structural stability are not correlated across different proteins. Similarly, χ2 values without context are inconclusive and must be compared for each protein system. Both REF15 and χ2 are available from a simulation, yielding a molecular system’s atom positions over time.
For SAXS-guided structure-based simulations, two MD parameters are particularly important: bias weight kχ and temperature T. kχ balances information in the SAXS data with the physical model, and T is a measure of available thermal energy and controls the system’s conformational flexibility. Thus, a particle corresponds to a simulation using a particular MD parameter set, \({\bf{x}}=\left({k}_{\chi },\ T\right)\). The OF is set up as
The first response evaluates the average physical stability of simulated structures, the second is the median χ2 deviation of simulated data from the target data. Owing to the ill-posed nature of the SAXS inverse problem, globally distinct protein structures can possess the same scattering intensity. As shown in Fig. 1, this can lead to a pronounced ambiguity in χ2. To resolve the resulting non-injectivity in the OF, we introduce a third response, the inverse average χ2 deviation. This acts as a regularizer, rewarding deviations from the target data and thus preventing possible overfitting. Combining these responses yields a surrogate model of a simulated ensemble’s similarity to the desired target structure. The smaller the OF, the more physical, data-consistent and (likely) similar to the target state the simulated structures are.
The ill-posedness of the SAXS inverse problem results in a pronounced ambiguity in the χ2 deviation of simulated data from the target data. This manifests in a non-injective ‘two-branch’ behaviour of the second response as a function of similarity of simulated structures to the desired target state, here quantified by the GDT. GDTsys, GDT between initial and target protein structure; kχ, bias weight; ε, energy scale of the structure-based model. Data from all lysine-, arginine-, ornithine-binding protein holo-to-apo runs are combined.
In physico-empirical structure-based models, different combinations of bias weight and temperature can equally yield useful results. There is no MD parameter ground truth for this type of simulation, so a purely evidence-based evaluation according to the similarity of simulated protein structures to the target is to be applied.
We use the global distance test (GDT)40 to quantify differences between two conformations of a protein (Methods section ‘Root-mean-square deviation’). To estimate how similar two superimposed structures are, the displacement of each alpha carbon is compared to various distance cutoffs. Percentages, Px, of alpha carbons with displacements below cutoffs of x Å are used to calculate the total score:
Higher GDT values indicate a stronger similarity between two models. Structures with GDT > 50 are considered topologically accurate41. The GDT is used to validate the OF as a surrogate model of an ensemble’s similarity to the target structure.
Results
We optimized MD parameters of SAXS-guided structure-based simulations for two well-characterized proteins: lysine-, arginine-, ornithine-binding (LAO) protein and adenylate kinase (ADK).
Small ligands such as sugars and amino acids are actively transported into bacteria across cell membranes42. Dedicated transport systems comprise a receptor (that is, the binding protein) and a membrane-bound protein complex. Interactions of the ligated binding protein with the membrane components induce conformational changes in the latter, forming an entry pathway for the ligand. We study LAO protein (Fig. 2a), which undergoes a conformational change from an apo (unligated; Protein Data Bank43 (PDB) code 2LAO44) to a holo (ligated; PDB code 1LST44) state upon ligand binding. The structures have a GDT of 39.39. SAXS-guided simulations started from the holo state and aimed at the apo state.
a, Lysine-, arginine-, ornithine-binding protein in its holo (coloured) and apo (grey) state, with a GDT of 39.39. b, Adenylate kinase in its open (coloured) and closed (grey) state, with a GDT of 33.06. The colouring indicates the displacement of each alpha carbon in the initial structure with respect to the target state. The average alpha-carbon displacement in each coloured structure with respect to each grey structure is given. Structures are visualized with PyMOL58.
Adenosine triphosphate (ATP) is the universal energy source in cells and is vital to processes such as muscle contraction and nerve impulse propagation. By continuously checking ATP levels, ADK provides the cell with a mechanism to monitor energetic levels and metabolic processes. The transition between an open (PDB code 4AKE45) and closed (PDB code 1AKE46) state is quintessential to its catalytic function47. The structures have a GDT of 33.06. SAXS-guided simulations started from the open conformation and aimed at the closed one.
Artificial target data were calculated from known structures with CRYSOL48. Statistical uncertainties were modelled following ref. 49.
We performed seven FLAPS runs with different initial conditions for each protein. Swarm-based metaheuristics such as FLAPS have hyperparameters influencing the optimization behaviour, and their efficacy can only be demonstrated empirically by a finite number of computational experiments. We present results for a swarm of 10 particles and 15 generations as a workable trade-off between optimization performance and compute time for the considered application. This set-up was found to be sufficient for convergence in preceding trial runs. Calculations were performed on 1,000 cores of a supercomputer. One run cost ~40,000 core hours. The results of the three best runs are listed in Table 1 (complete results are provided in Supplementary Tables 1 and 2).
As shown for LAO protein in Fig. 3, the OF consistently converged to a stable topology (Supplementary section ‘Analyzing swarm convergence’).
For each simulation, we calculated the median GDT with respect to the target state from all structures in the trajectory. To validate the OF, we state its Pearson correlation ρ with the median GDT as a measure of linear correlation. Because minimizing the OF should be equivalent to maximizing the GDT, negative correlations, ideally −1, are expected. The OF’s suitability is confirmed for both LAO protein and ADK with correlations up to −0.94 and −0.85, respectively. As the Pearson correlation only reflects linearity, we also studied the exact relations between the OF and the median GDT (Fig. 4). To identify the best MD parameter combinations, the OF must have low values for large GDTs, irrespective of the actual relationship’s complexity. This is the case for both test systems.
Global best positions, gbest, returned functional MD parameter combinations throughout. For LAO protein, gbest median GDTs were consistently of the order of 70 and correspond well to the best values reached. This means that for half of the simulated structures, at least 70% of all alpha carbons lie within a small radius from their positions in the target state. These results indicate the structural accuracy of the simulated ensemble for gbest and thus convergence to the target state and successful refinement against the data. The same is true for ADK, where gbest and the best median GDTs are around 63 and more similar than those of LAO protein. Example structures from gbest simulations, shown in Fig. 5, are in nearly perfect accordance with the target states.
a,b, LAO protein (seed 1790954) (a) and ADK (seed 1795691) (b). Structures with maximum GDT are shown (coloured) and are almost identical to the respective target states (grey). The colouring indicates the displacement of each alpha carbon in the simulated structure with respect to the target state. The average alpha-carbon displacement in each coloured structure with respect to each grey structure is given. Structures are visualized with PyMOL58.
Additionally, we considered the reversed conformational transitions, that is, from apo to holo state for LAO protein and closed to open state for ADK (Supplementary Tables 3 and 4, respectively). With ρ between −0.56 and −0.88, FLAPS could also identify functional MD parameters for SAXS-guided apo-to-holo simulations of LAO protein. gbest and the best median GDTs were consistently slightly below 70, indicating high similarity of the simulated structures and the desired target state for the MD parameter combinations found.
By comparison, Pearson correlations up to −0.53 and gbest median GDTs of ~45 indicate only average structural accuracy for the closed-to-open transition of ADK. However, overall best median GDTs were around 50. With only half of all alpha carbons within a small radius from their target positions, the observed behaviour is not a problem of FLAPS, but is due to the limits of the underlying simulation method7. The information in the coarse-grained structure-based model and the low-resolution SAXS data seems to be insufficient to determine the molecular structure with the same accuracy as for the other test cases. However, even under these circumstances, FLAPS was capable of determining acceptable parameters.
To evaluate FLAPS’s efficiency, we performed comparative grid-search optimizations where we found superior performance of FLAPS for all considered protein systems (Supplementary section ‘Comparison to grid search’).
Discussion
The inverse problem of reconstructing molecular structures from low-resolution SAXS data is still unsolved. Biomolecular simulations are among the most powerful tools for eliminating the arising ambiguity and access the valuable structural information content of such data. However, data-assisted simulations rely on MD parameters, where, most importantly, experimental information must be weighted accurately with respect to the physical model.
Here, we have shown how computational intelligence can be used to systematically explore MD parameter spaces and optimize the performance of complex physics-based simulation techniques. We introduced FLAPS, a data-driven solution for a fully automatic and reproducible parameter search based on particle swarms.
To identify the best MD parameters for SAXS-guided protein simulations, we designed an OF as an accurate surrogate of simulation quality in terms of physical structures matching the target data. A suitable OF will typically depend on multiple quality features of different scales to equally reflect a data-assisted simulation’s physical plausibility and its agreement with the data. To handle multiple responses in classical PSO, they need to be mapped to a scalar score via multiplication by fixed weights. These additional OF parameters must either be chosen manually (and probably suboptimally) in advance or be absorbed into the search space, resulting in a massive increase in dimensionality. FLAPS solves this problem by intelligently learning OF parameters in the optimization process, avoiding the need to set them as ‘magic numbers’, while reducing the search-space dimensionality to a minimum. Various responses are automatically balanced with respect to each other to enable a meaningful and unbiased comparison on a shared scale. Implemented in FLAPS, our conceptual OF reliably identified useful MD parameters for two different proteins, where we observed convergence of simulated structures to the target state. Owing to the dedicated in situ processing, the algorithm leverages available computing resources and transparently scales from a laptop to current supercomputers.
In previous studies, the bias weight is conveniently chosen as the smallest value yielding satisfactory χ2 (refs. 7,8). This purely data-based selection criterion neglects the physical information provided by the molecular simulations, risking the selection of physically dysfunctional values due to the ill-posedness of the SAXS inverse problem. The flexible composite OF in FLAPS allows us to include multiple selection criteria, yielding a direct surrogate of simulation quality in terms of not only data conformity but also the physical plausibility of simulated structures. In contrast to grid search, the optimization does not rely on a predefined list of manually chosen and a priori fixed candidates. This circumvents either missing an optimum, spanning large search spaces due to fine-grained grids, or both. Instead, a directional search led by the most current best solutions provides guidance in the selection process and a valid context for meaningful interpretation of multi-layer criteria. Additionally, the compute time to find useful parameters is decreased by reducing the dimensionality.
FLAPS can be transferred easily to optimize other life-sciences applications, such as simulation methods incorporating data from various experimental sources11. In a more general context, it solves the problem of weighting different contributions of composite OFs in multi-response optimization. Such problems often occur in industrial manufacturing, processing and design, where manually chosen weights are commonly used32,33. In FLAPS, OF parameters can be learned in a self-improving manner according to any desired scheme, for example, standardization, rescaling, mean normalization or a relative weighting after one of the aforementioned steps. FLAPS shows how computational-intelligence concepts can be harnessed successfully for practical optimization problems at the forefront of life sciences.
Methods
SAXS-guided protein simulations
SAXS-guided protein simulations were set up as described in ref. 7 and run with the MD engine GROMACS 5, including the scattering-guided MD extension12. Simulation input files are available on GitHub (github.com/FLAPS-NMI/FLAPS-sim_setups). We used the popular CRYSOL software48 in default mode to compute artificial SAXS intensities from known initial and target structures. Each intensity contained 700 equidistant data points up to a momentum transfer of q = 0.35 Å−1. During a simulation, the SAXS intensities of simulated structures are calculated internally in GROMACS7,12 using the Debye equation50 with amino-acid scattering factors corrected for displaced solvent51. The CRYSOL intensities were rescaled such that the extrapolation of the forward scattering matches the internally calculated GROMACS value. Uncertainties were modelled as described in ref. 49. Target SAXS data were calculated as the difference of rescaled CRYSOL intensities, including uncertainties, of known initial and target structures. The initial structure’s rescaled CRYSOL intensity, including uncertainties, served as the absolute reference scattering in the simulations7. We included 17 data points selected as the difference curve’s local extrema and interjacent points centred between each two extrema. For both proteins, this corresponds well to the number of independent Shannon channels giving the number of independent data points in a SAXS curve10.
Root-mean-square deviation
In addition to the GDT analysis presented in the main text, we considered the more common root-mean-square deviation (RMSD) as a structural similarity measure. The results are presented in Supplementary Tables 1–4.
The RMSD is the minimal mass-weighted average distance between N atoms (usually backbone or alpha carbon) of two superimposed structures over all possible spatial translations and rotations,
where \(M=\mathop{\sum }\nolimits_{i = 1}^{N}{m}_{i}\) and mi is the mass of atom i. ri and ri,0 are the positions of atom i in the mobile and reference structure, respectively. Holo/apo LAO protein and open/closed ADK have alpha-carbon RMSD values of 4.7 Å and 7.1 Å, respectively.
A disadvantage is that RMSD correlates strongly with the largest displacement between two structures, and small numbers of displaced atoms induce large changes. We use GDT40,52,53 as the main target metric as it more accurately accounts for local misalignments.
Implementation
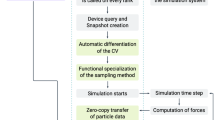
FLAPS is implemented as a stand-alone solver in Hyppopy, a Python-based hyperparameter optimization package available at https://github.com/MIC-DKFZ/Hyppopy. Hyppopy provides tools for blackbox optimization. It has a simple, unified application programming interface (API) that can be used to access a collection of solver libraries. Our implementation of FLAPS is available on GitHub54,55. We implemented a Message-Passing-Interface (MPI)-parallel version of the code using a sophisticated parallelization architecture as described in Supplementary Fig. 13. Available compute nodes comprising a given number of processors are divided into blocks, each of which corresponds to one particle in the swarm. Within one block, the simulation itself runs on a single core, while all the other cores process the generated frames in the trajectory on the fly. This results in a massive reduction in runtime.
The experiments were run on the ForHLR II cluster system located at the Steinbuch Centre for Computing at Karlsruhe Institute of Technology. The system comprises 1,152 thin, that is, solely central processing unit (CPU)-based, compute nodes. Each node is equipped with two 10-core Intel Xeon E5-2660 v3 Haswell CPUs at 3.3 GHz, 64 GB of DDR3 main memory and 4x Mellanox 100-Gbit EDR InfiniBand links. The software packages used were a RHEL Linux with kernel version 4.18.0 and Python 3.6.8.
Each run used 51 compute nodes (1,020 cores in total). Owing to the magnitude of metadata and I/O operations, we used a private on-demand file system (BeeGFS On-Demand) with a stripe count of 1, where one node was reserved for the metadata server56. Each block in the underlying simulator-worker scheme consisted of five nodes, that is, 100 cores (one simulator, 99 workers). Each run cost ~40,000 CPU hours. For the presented application, we used cognitive acceleration coefficient ϕ1 = 2.0 and social acceleration coefficient ϕ2 = 1.5 in the particle update (Algorithm 1). The complete set-up, including all PSO hyperparameters used, is available on GitHub57.
Data availability
The software for the metaheuristic molecular dynamics parameter optimization of SAXS-guided structure-based protein simulations used in this work54,55 is publicly available at https://github.com/FLAPS-NMI?tab=repositories. A minimal dataset to reproduce the presented results57 is publicly available at https://github.com/FLAPS-NMI/FLAPS-sim_setups/releases/tag/v1.0 and published under the Creative Commons Attribution 4.0 International Public License.
Code availability
All code used in this work is publicly available at https://github.com/FLAPS-NMI?tab=repositories and published under the New BSD Licence. The MPI-parallelized FLAPS solver is implemented in Optunity, a hyperparameter tuning package for Python. Our extended version of Optunity55 is available at https://github.com/FLAPS-NMI/FLAPS-optunity/releases/tag/v1.0 and integrated into Hyppopy, a Python-based toolbox for blackbox optimization. Our extended version of Hyppopy54 is available at https://github.com/FLAPS-NMI/FLAPS-Hyppopy/releases/tag/v1.0.
Change history
27 October 2021
In the version of this article initially published online, the following metadata was omitted and has now been included: “Open access funding provided by Forschungszentrum Jülich GmbH (4205)”.
References
Selkoe, D. Folding proteins in fatal ways. Nature 426, 900–904 (2003).
Selkoe, D. J. Cell biology of protein misfolding: the examples of Alzheimer’s and Parkinson’s diseases. Nat. Cell Biol. 6, 1054–1061 (2004).
Soto, C. Unfolding the role of protein misfolding in neurodegenerative diseases. Nat. Rev. Neurosci. 4, 49–60 (2003).
Mukherjee, A., Morales-Scheihing, D., Butler, P. C. & Soto, C. Type 2 diabetes as a protein misfolding disease. Trends Mol. Med. 21, 439–449 (2015).
Dobson, C. M. Protein-misfolding diseases: getting out of shape. Nature 418, 729–730 (2002).
Karaca, E., Rodrigues, J. P., Graziadei, A., Bonvin, A. M. & Carlomagno, T. M3: an integrative framework for structure determination of molecular machines. Nat. Methods 14, 897–902 (2017).
Weiel, M., Reinartz, I. & Schug, A. Rapid interpretation of small-angle X-ray scattering data. PLoS Comput. Biol. 15, e1006900 (2019).
Hermann, M. R. & Hub, J. S. SAXS-restrained ensemble simulations of intrinsically disordered proteins with commitment to the principle of maximum entropy. J. Chem. Theory Comput. 15, 5103–5115 (2019).
Chen, P. et al. Combined small-angle X-ray and neutron scattering restraints in molecular dynamics simulations. J. Chem. Theory Comput. 15, 4687–4698 (2019).
Chen, P. & Hub, J. S. Interpretation of solution X-ray scattering by explicit-solvent molecular dynamics. Biophys. J. 108, 2573–2584 (2015).
Whitford, P. C. et al. Excited states of ribosome translocation revealed through integrative molecular modeling. Proc. Natl Acad. Sci. USA 108, 18943–18948 (2011).
Björling, A., Niebling, S., Marcellini, M., van der Spoel, D. & Westenhoff, S. Deciphering solution scattering data with experimentally guided molecular dynamics simulations. J. Chem. Theory Comput. 11, 780–787 (2015).
Shevchuk, R. & Hub, J. S. Bayesian refinement of protein structures and ensembles against SAXS data using molecular dynamics. PLoS Comput. Biol. 13, e1005800 (2017).
Hummer, G. & Köfinger, J. Bayesian ensemble refinement by replica simulations and reweighting. J. Chem. Phys. 143, 243150 (2015).
Hub, J. S. Interpreting solution X-ray scattering data using molecular simulations. Curr. Opin. Struct. Biol. 49, 18–26 (2018).
Kennedy, J. & Eberhart, R. Particle swarm optimization. In Proc. ICNN’95—International Conference on Neural Networks Vol. 4, 1942–1948 (IEEE, 1995); https://doi.org/10.1109/ICNN.1995.488968
Kennedy, J. The particle swarm: social adaptation of knowledge. In Proc. 1997 IEEE International Conference on Evolutionary Computation (ICEC’97) 303–308 (IEEE, 1997); https://doi.org/10.1109/ICEC.1997.592326
Shi, Y. & Eberhart, R. A modified particle swarm optimizer. In Proc. 1998 IEEE International Conference on Evolutionary Computation. IEEE World Congress on Computational Intelligence (cat. no. 98TH8360) 69–73 (IEEE, 1998); https://doi.org/10.1109/ICEC.1998.699146
Clerc, M. & Kennedy, J. The particle swarm—explosion, stability and convergence in a multidimensional complex space. IEEE Trans. Evol. Comput. 6, 58–73 (2002).
Parsopoulos, K. E. & Vrahatis, M. N. Recent approaches to global optimization problems through particle swarm optimization. Nat. Comput. 1, 235–306 (2002).
Jordehi, A. R. Particle swarm optimisation for dynamic optimisation problems: a review. Neural Comput. Appl. 25, 1507–1516 (2014).
Blackwell, T. in Evolutionary Computation in Dynamic and Uncertain Environments (eds Yang, S. et al.) 29–49 (Springer, 2007); https://doi.org/10.1007/978-3-540-49774-5_2
Stanley, K. O., Clune, J., Lehman, J. & Miikkulainen, R. Designing neural networks through neuroevolution. Nat. Mach. Intell. 1, 24–35 (2019).
Taherkhani, M. & Safabakhsh, R. A novel stability-based adaptive inertia weight for particle swarm optimization. Appl. Soft Comput. 38, 281–295 (2016).
Eberhart, R. C. & Shi, Y. Comparing inertia weights and constriction factors in particle swarm optimization. In Proc. 2000 Congress on Evolutionary Computation. CEC00 (cat. no. 00TH8512) Vol. 1, 84–88 (IEEE, 2000); https://doi.org/10.1109/CEC.2000.870279
Pedersen, M. E. H. & Chipperfield, A. J. Simplifying particle swarm optimization. Appl. Soft Comput. 10, 618–628 (2010).
Meissner, M., Schmuker, M. & Schneider, G. Optimized particle swarm optimization (OPSO) and its application to artificial neural network training. BMC Bioinformatics 7, 125 (2006).
Nobile, M. S. et al. Fuzzy self-tuning PSO: a settings-free algorithm for global optimization. Swarm Evol. Comput. 39, 70–85 (2018).
Poli, R. Analysis of the publications on the applications of particle swarm optimisation. J. Artif. Evol. Appl. 2008, 685175 (2008).
Sengupta, S., Basak, S. & Peters, R. A. Particle swarm optimization: a survey of historical and recent developments with hybridization perspectives. Mach. Learn. Knowledge Extraction 1, 157–191 (2019).
Navalertporn, T. & Afzulpurkar, N. V. Optimization of tile manufacturing process using particle swarm optimization. Swarm Evol. Comput. 1, 97–109 (2011).
Pawar, P., Rao, R. & Davim, J. Multiobjective optimization of grinding process parameters using particle swarm optimization algorithm. Mater. Manuf. Process. 25, 424–431 (2010).
Ma, C. & Qu, L. Multiobjective optimization of switched reluctance motors based on design of experiments and particle swarm optimization. IEEE Trans. Energy Conver. 30, 1144–1153 (2015).
Zhang, C., Chen, Z., Mei, Q. & Duan, J. Application of particle swarm optimization combined with response surface methodology to transverse flux permanent magnet motor optimization. IEEE Trans. Magn. 53, 8113107 (2017).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. In Proc. 32nd International Conference on Machine Learning Vol. 37 (eds. Bach, F. & Blei, D.) 448–456 (PMLR, 2015).
Onuchic, J. N. & Wolynes, P. G. Theory of protein folding. Curr. Opin. Struct. Biol. 14, 70–75 (2004).
Schug, A. & Onuchic, J. N. From protein folding to protein function and biomolecular binding by energy landscape theory. Curr. Opin. Pharmacol. 10, 709–714 (2010).
Alford, R. F. et al. The Rosetta all-atom energy function for macromolecular modeling and design. J. Chem. Theory Comput. 13, 3031–3048 (2017).
Leman, J. K. et al. Macromolecular modeling and design in Rosetta: recent methods and frameworks. Nat. Methods 17, 665–680 (2020).
Zemla, A. LGA: a method for finding 3D similarities in protein structures. Nucleic Acids Res. 31, 3370–3374 (2003).
Moult, J., Fidelis, K., Kryshtafovych, A., Schwede, T. & Tramontano, A. Critical assessment of methods of protein structure prediction (CASP)–round XII. Proteins 86, 7–15 (2018).
Ames, G. F.-L. Bacterial periplasmic transport systems: structure, mechanism and evolution. Annu. Rev. Biochem. 55, 397–425 (1986).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Oh, B.-H. et al. Three-dimensional structures of the periplasmic lysine/arginine/ornithine-binding protein with and without a ligand. J. Biol. Chem. 268, 11348–11355 (1993).
Müller, C., Schlauderer, G., Reinstein, J. & Schulz, G. E. Adenylate kinase motions during catalysis: an energetic counterweight balancing substrate binding. Structure 4, 147–156 (1996).
Müller, C. W. & Schulz, G. E. Structure of the complex between adenylate kinase from Escherichia coli and the inhibitor Ap5A refined at 1.9-Å resolution: a model for a catalytic transition state. J. Mol. Biol. 224, 159–177 (1992).
Whitford, P. C., Miyashita, O., Levy, Y. & Onuchic, J. N. Conformational transitions of adenylate kinase: switching by cracking. J. Mol. Biol. 366, 1661–1671 (2007).
Svergun, D., Barberato, C. & Koch, M. H. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Sedlak, S. M., Bruetzel, L. K. & Lipfert, J. Quantitative evaluation of statistical errors in small-angle X-ray scattering measurements. J. Appl. Crystallogr. 50, 621–630 (2017).
Debye, P. Zerstreuung von Röntgenstrahlen. Annal. Phys. 351, 809–823 (1915).
Yang, S., Park, S., Makowski, L. & Roux, B. A rapid coarse residue-based computational method for X-ray solution scattering characterization of protein folds and multiple conformational states of large protein complexes. Biophys. J. 96, 4449–4463 (2009).
Kufareva, I. & Abagyan, R. in Homology Modeling (eds Orry, A. J. W. & Abagyan, R.) 231–257 (Springer, 2011).
Modi, V., Xu, Q., Adhikari, S. & Dunbrack Jr, R. L. Assessment of template-based modeling of protein structure in CASP11. Proteins 84, 200–220 (2016).
Weiel, M. et al. FLAPS Hyppopy code repository (FLAPS-NMI@Github/FLAPS-Hyppopy, 2021); https://doi.org/10.5281/zenodo.4773970, https://github.com/FLAPS-NMI/FLAPS-Hyppopy/releases/tag/v1.0
Weiel, M. et al. FLAPS Optunity code repository (FLAPS-NMI@Github/FLAPS-optunity, 2021); https://doi.org/10.5281/zenodo.4773992, https://github.com/FLAPS-NMI/FLAPS-optunity/releases/tag/v1.0
Soysal, M. et al. Using on-demand file systems in HPC environments. In 2019 International Conference on High Performance Computing & Simulation (HPCS) 390–398 (IEEE, 2019); https://doi.org/10.1109/HPCS48598.2019.9188216
Weiel, M. et al. Minimal dataset repository for reproduction of presented results (FLAPS-NMI@Github/FLAPS-sim_setups, 2021); https://doi.org/10.5281/zenodo.4773999, https://github.com/FLAPS-NMI/FLAPS-sim_setups/releases/tag/v1.0
The PyMOL Molecular Graphics System, Version 1.8 (Schrödinger, 2015).
Acknowledgements
We are thankful for computing time on the ForHLR II computer cluster, funded by the Ministry of Science, Research and Arts Baden-Württemberg and by the Federal Ministry of Education and Research. M.W., M.G., A.K. and D.C. acknowledge support by the Helmholtz Association’s Initiative and Networking Funds under project no. ZT-I-0003. M.G. and D.C. acknowledge support by the Helmholtz AI platform grant. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. M.W. gratefully acknowledges support by the DFG Research Training Group 2450.
Funding
Open access funding provided by Forschungszentrum Jülich GmbH (4205).
Author information
Authors and Affiliations
Contributions
M.W., M.G. and A.S. conceived the study. M.W., M.G., A.K. and R.F. developed the methodology. M.W., A.K. and M.G. implemented the FLAPS solver in Hyppopy. M.W. conducted the optimization runs under the supervision of M.G. and A.S. All authors discussed the results. M.W., M.G. and D.C. wrote the manuscript. M.W. designed and produced figures. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13, discussion on swarm convergence and comparison to grid search and Tables 1–5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Weiel, M., Götz, M., Klein, A. et al. Dynamic particle swarm optimization of biomolecular simulation parameters with flexible objective functions. Nat Mach Intell 3, 727–734 (2021). https://doi.org/10.1038/s42256-021-00366-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-021-00366-3
This article is cited by
-
RNA contact prediction by data efficient deep learning
Communications Biology (2023)
-
Machine learning-enabled globally guaranteed evolutionary computation
Nature Machine Intelligence (2023)
-
Bird’s Eye View feature selection for high-dimensional data
Scientific Reports (2023)
-
Identification of nonlinear bolted lap joint parameters using instantaneous power flow balance-based substructure approach
International Journal of Dynamics and Control (2023)
-
A consistency and consensus-driven approach for granulating linguistic information in GDM with distributed linguistic preference relations
Artificial Intelligence Review (2023)