Abstract
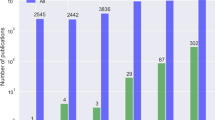
A number of large technology companies have created code-free cloud-based platforms that allow researchers and clinicians without coding experience to create deep learning algorithms. In this study, we comprehensively analyse the performance and featureset of six platforms, using four representative cross-sectional and en-face medical imaging datasets to create image classification models. The mean (s.d.) F1 scores across platforms for all model–dataset pairs were as follows: Amazon, 93.9 (5.4); Apple, 72.0 (13.6); Clarifai, 74.2 (7.1); Google, 92.0 (5.4); MedicMind, 90.7 (9.6); Microsoft, 88.6 (5.3). The platforms demonstrated uniformly higher classification performance with the optical coherence tomography modality. Potential use cases given proper validation include research dataset curation, mobile ‘edge models’ for regions without internet access, and baseline models against which to compare and iterate bespoke deep learning approaches.
Similar content being viewed by others
Main
Clinical decision making increasingly benefits from the supplementary data afforded by modern medical imaging techniques, and many non-invasive modalities are now routinely incorporated into patient evaluation pathways. Ophthalmology and retinal medicine is an exemplar specialty with an exceptionally high use of in-office imaging1. Some institutions report a >10-fold increase in the annual generation of imaging data over the last decade2. The most ubiquitous imaging modalities in ophthalmology are fundus photography and optical coherence tomography (OCT). First reported in 1886 but now available increasingly in primary care and even smartphone-based settings3, fundus photography provides a two-dimensional (2D) colour image typically encompassing the central retina, major blood vessels and optic nerve. Major applications of fundus photography include screening for two leading causes of global blindness in diabetic eye disease and glaucoma4,5,6,7. OCT, in contrast, leverages near-infrared light and interferometry to depict volumetric (that is, 3D) data of the retina with axial resolutions of less than 10 μm (ref. 8). Many diseases of the retina have been redefined by its advent. Indeed, OCT-based parameters (such as the thickness of the central retina) are now well-established biomarkers of disease activity and clinical trial endpoints9,10,11,12.
One form of artificial intelligence, deep learning, has demonstrated compelling results in the imaging classification of numerous ophthalmic diseases13,14,15,16. Modelled on the concept of biological neural networks, deep learning employs hidden layers of nodes, whose collective interplay can map an output through weights derived by a training process from input data17. Convolutional neural networks (CNNs) have shown encouraging results across a range of medical image classification tasks18. CNNs modelled on fundus photography have diagnostic accuracy comparable to that of many international screening programmes in diabetic retinopathy (DR)13,14,19. Similarly, CNNs in OCT have shown performance comparable to retinal specialists with decades of experience15,16. However, the development of such deep learning-based models demands substantial resources, including (1) well-curated and labelled data in a computationally tractable form, (2) sufficient computer hardware, often in the form of expensive graphics processing units (GPUs) for model development, and (3) deep learning expertise20.
With limited resources and concentrated artificial intelligence (AI) talent pools, coordinating the aforementioned requirements is difficult for clinical research groups, and more so for individual clinicians21. One promising solution to facilitate all mentioned provisions is automated machine learning (AutoML). AutoML describes a set of tools and techniques for streamlining model development by automating the selection of optimal network architectures, pre-processing methods and hyperparameter optimization. As these platforms mature, the automation of these processes may diminish the necessity for the programming experience required to design such models. A number of services offering AutoML additionally provide the prerequisite hardware through cloud-based GPUs or tensor processing units (TPUs). Some platforms offer a code-free deep learning (CFDL) approach, which is even more accessible to a clinician or researcher without coding expertise.
Previously, we reported on the feasibility of using Google Cloud AutoML Vision to design medical image classifiers across a range of modalities including chest X-ray, dermatoscopy, fundus photography and OCT. However, this exploratory study was limited to a single application programming interface, provided by Google Inc20. Since that report, the field of AutoML has matured substantially, with several vendors now providing platforms for code-free design of deep learning models. Here, on publicly available datasets of retinal fundus photography and OCT scans, we evaluate the diagnostic accuracy and features of six CFDL platforms: Amazon Rekognition Custom Labels (Amazon), Apple Create ML (Apple), Clarifai Train (Clarifai), Google Cloud AutoML Vision (Google), MedicMind Deep Learning Training Platform (MedicMind) and Microsoft Azure Custom Vision (Microsoft).
Model and platform evaluation
Deep learning model performance
The mean (s.d.) F1 scores representing the harmonic mean of the precision and recall across platforms for all model–dataset pairs were as follows: Amazon, 93.9 (5.4); Apple, 72.0 (13.6); Clarifai, 74.2 (7.1); Google, 92.0 (5.4); MedicMind, 90.7 (9.6); Microsoft, 88.6 (5.3) (Fig. 1).
As large datasets could not be processed by the Clarifai and MedicMind platforms, missing values prevented an analysis of variance (ANOVA) analysis of the F1 scores across all platforms and datasets. Therefore, we split our analysis into platforms that were able and unable to process large datasets.
When comparing platforms able to process large datasets (Amazon, Apple, Google and Microsoft), post hoc two-way ANOVA analysis of F1 scores with Bonferroni’s multiple comparison correction (Supplementary Table 1) showed a significant difference only for Amazon versus Apple, with a mean difference (95% CI) of 21.9(1.3, 42.5). Post hoc analysis comparing platforms within each dataset (Supplementary Table 2) yielded significant differences in F1 scores of models generated on the Kermany dataset of Google versus Apple 45.8(4.6, 87.0) and Amazon versus Apple 47.2(6.0, 88.4). A platform performance comparison on small datasets yielded significantly poorer performance for Apple and Clarifai platforms.
Evaluation by platform and modality
OCT
Microsoft does not provide image-level results in the graphical user interface (GUI), so we were unable to calculate the specificity, negative predictive value (NPV) and accuracy of this platform, and those metrics were reported as not aplicable (NA). Deep learning models trained on the relatively smaller Waterloo OCT dataset exhibited uniformly high classification performance (Extended Data Fig. 1) with F1;(sensitivity, specificity, positive predictive value (PPV), accuracy) as follows: Amazon, 97.8;(97.4, 99.6, 98.2, 99.1); Apple, 78.8;(78.8, 94.7, 78.8, 91.5); Clarifai, 79.2;(73.0, 96.5, 86.6, 90.9); Google, 93.8;(93.8, 98.5, 93.8, 97.5); MedicMind, 97.4;(97.4, 99.3, 97.4, 98.9); Microsoft, 94.8;(94.8, NA, 94.8, NA) (Fig. 2). The MedicMind and Clarifai models were both unable to be trained on the much larger Kermany OCT dataset due to GUI crashes during training and dataset upload, respectively. This was attempted a minimum of two times on each platform. Platforms were made aware of this in February 2020 and their response elucidated upload limits of 128 and 1,000 images, respectively. Classification models on platforms that were successfully able to train deep learning models demonstrated the following classification performance: Amazon, 99.2;(99.3, 99.7, 99.1, 99.6); Apple, 52.0;(51.5, 84.5, 52.6 76.3); Google, 97.8;(97.8, 99.3, 97.8, 98.9); Microsoft, 91.1;(90.6, NA, 91.7, NA).
Fundus photography
Classification models trained for referable diabetic retinopathy (RDR) and non-referable diabetic retinopathy (NRDR) classification on the relatively smaller fundus photograph Messidor dataset demonstrated uniformly moderate performance with F1;(sensitivity, specificity, PPV, accuracy) as follows: Amazon, 88.5;(88.5, 88.5, 88.5, 88.5); Apple, 75.1;(75.1, 75.1, 75.1, 75.1); Clarifai, 69.2;(69.2, 69.2, 69.2, 69.2); Google, 84.8;(84.8, 84.8, 84.8, 84.8); MedicMind, 83.9;(83.9, 83.9%, 83.9, 83.9); Microsoft, 84.8;(84.8, NA, 84.8, NA). Class-pooled calculation results in identical values for these metrics, because platform limitation required that the binary RDR versus NRDR task was trained as two independent classes; thus, a false positive for RDR is also a false negative for NRDR. The MedicMind and Clarifai models were both similarly unable to be trained on the much larger EyePACS fundus dataset due to GUI crashes during training and dataset upload, respectively. This was attempted a minimum of two times on each platform. Classification models on platforms that were successfully able to train models demonstrated moderately high classification performance: Amazon, 90.0;(92.7, 86.7, 87.5, 89.7); Apple, 82.1;(82.1, 82.1, 82.1, 82.1); Google, 91.6;(91.6, 91.6, 91.6, 91.6); Microsoft, 83.7;(83.7, NA, 83.7, NA).
External validation
External validation of the fundus models whose platforms supported this feature (Google, MedicMind) was performed with the IDRiD diabetic retinopathy evaluation set22. The Google EyePACS, Google Messidor and MedicMind Messidor models demonstrated F1;(sensitivity, specificity, PPV, accuracy) of 85.3%;(90.6%, 64.1%, 80.6%, 80.6%), 81.3%;(98.4%, 28.2%, 69.2%, 71.8%) and 83.3%;(93.8%, 48.7%, 75.0%, 76.7%), respectively (Supplementary Table 3). MedicMind failed to train a model on the large Kermany dataset and thus could not be validated externally. An OCT dataset for external validation containing image disease labels matching the Kermany and Waterloo datasets was not located after an extensive literature review utilizing Google Dataset Search. To ensure study reproducibility, we intentionally limited our investigation to using public datasets. As researchers design future models with these AutoML platforms, proper external validation will be necessary for each model before implementation, ensuring ethics approvals are obtained for patient-derived validation datasets.
Repeatability
We trained three models of a representative dataset (Waterloo OCT) on each platform. The model F1 (s.d.), range values were Amazon, 97.8 (0.50), 1.00; Apple, 72.0 (1.26), 2.4; Clarifai, 79.8 (0.90), 1.52; Google, 94.1 (1.35), 2.66; MedicMind, 95.9 (2.57), 4.45; Microsoft, 91.6 (4.80), 2.74. The standard deviations were relatively small, demonstrating reasonable repeatability, probably due to varying random seeds for AutoML training23.
Usability, features and cost
For the application of CFDL to diagnostic classification problems, we identified the following as useful features: custom test/train splits, batch prediction, cross-validation, data augmentation, .csv file upload, saliency maps, threshold adjustment and confusion matrices. These features were variably present in the platforms (Table 1).
Select features were found to be especially useful when considering ease, reproducibility and model explainability. For data management, these include the ability to designate test/train splits (Amazon, Apple, Google, MedicMind), the ability to perform k-fold cross-validation (Microsoft, Clarifai) and the ability to perform data augmentation, to assist with generalizability (Apple). The Apple platform also ran locally, which had the simultaneous advantages of cloud cost savings and limitations of locally available compute power. Researchers also highlighted the efficiency of local data manipulation and subsequent upload via .csv files, supported by Google and MedicMind, which was singled out as a crucial platform feature.
For model evaluation, useful features include saliency maps (MedicMind) (Fig. 3) and deeper model evaluation via TensorBoard, which have value for model explainability24,25,26. A similarly important feature for performance evaluation is threshold adjustment and live reclassification (Clarifai, Google). This allowed researchers to perform real-time threshold operating point selection, a necessary feature for decision curve analysis and real-world model deployment27,28. Beyond precision (PPV) and recall (sensitivity), confusion matrix generation (Apple, Clarifai, Google, MedicMind) is useful to generate clinically meaningful specificity and NPV metrics, without which it becomes difficult to accurately infer model performance at population levels. We contacted platforms that did not report confusion matrices to request the feature.
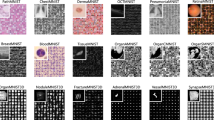
Input test images (left) and resulting saliency maps (right). a, Fundus photo of proliferative diabetic retinopathy with subhyaloid haemorrhage: the saliency map highlights vessels, temporal fibrosis and inferior subhyaloid haemorrhage. b, Fundus photo of macular oedema (hard exudates): the saliency map largely highlights hard exudates in the temporal macula. c, OCT of macular hole: the saliency map highlights the central macula and retinal hole. d, OCT of central serous retinopathy: the saliency map highlights the central macula and subretinal fluid.
Although the Apple and MedicMind platforms were free to use and the remaining platforms have free tiers, costs may mount for those utilizing these systems. Free tiers have cloud training hour limits, and models trained from large datasets may quickly exceed them. Model training is charged per cloud compute hour (Amazon, Google, Microsoft) from US$1 to US$19 or per number of images (Clarifai). Of the models we developed utilizing paid tiers (Microsoft), none exceeded US$100 for training. Platforms additionally charge for cloud model deployment and inference. Google allows training of an edge model, which is optimized for mobile devices and can be downloaded locally, enabling unlimited free prediction.
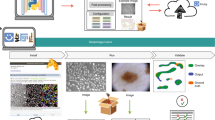
Among the CFDL platforms, GUIs consistently comprised three segments: data upload, data visualization and labelling, and model evaluation (Supplementary Video). These are split by panes or across web pages in their respective user interfaces (Extended Data Fig. 2). The three researchers (E.K., D.F., Z.G.) who evaluated the models were sent five-question surveys, which enquired about the user interface experience and ease of use of each of the aforementioned segments, along with overall platform experience (Supplementary Table 4). The latter question represents how likely they are to use those platforms in the future. In terms of overall experience, all users selected ‘satisfied’ (or above) with the Amazon and Google platforms, and all users of Google selected ‘very satisfied’.
Discussion
We believe that CFDL platforms have the potential to improve access to deep learning for both clinicians and biomedical researchers, and represent another step towards the democratization and industrialization of AI. In this study, we evaluated the diagnostic accuracy and user interface (UX) features of six CFDL platforms on publicly available medical datasets of multiple modalities. We specifically focused our evaluation on both objective and subjective metrics of each platform. To ensure fair comparison, we utilized identical test/train data splits across platforms and the maximum allowable training hours. Although differing reporting metrics among platforms prevented analyses across certain model performance metrics, we manually created contingency matrices (Table 2) to calculate relevant clinical criteria, including sensitivity and specificity.
Our evaluation yielded a split between platforms that were able to handle large imaging datasets (n > 35,000) to train deep learning models (Amazon, Apple, Google and Microsoft) and those that could not (Clarifai and MedicMind). Among the former platforms, we found high classification performance, with only Apple performing significantly worse when compared to the highest performing Amazon platform. Although this may be a result of computational limitations of training a model locally with the Apple platform as compared to a scaled cloud approach, the automated nature of these platforms makes it difficult to find the definitive reason. When comparing on smaller datasets across all six platforms, all platforms except Clarifai and Apple similarly demonstrated robust model performance. OCT classification models uniformly performed better than fundus photography models, which is probably a result of the higher dimensionality of the latter modality in each 2D image—that is, there are more variables (colour channels and regions of interest) in each colour fundus photograph than in an OCT image.
Our evaluation did not show significant performance differences among the leading platforms (Amazon, Google and Microsoft). However, these platforms differed significantly in terms of the critically important evaluation features available, such as providing threshold adjustments, precision–recall curves and confusion matrices through their respective GUIs. Amazon provided none of these, Google provided all of these, and Microsoft provided only threshold adjustment. Of these three platforms, only Google has batch prediction capability, which enables external validation at scale. Furthermore, because our evaluation did not yield significant performance differences among the majority of capable platforms, subjective feature evaluation becomes increasingly important. For the three clinicians who performed both model training and UX evaluation, the top preferred platforms were Amazon, Google and Microsoft. Although platform cost is in flux as a result of rapid iterations, performance per dollar will be another key metric for budget-constrained researchers choosing a platform. Furthermore, although cloud computing is infinitely more scalable, researchers must consider its cost paradigms as compared with traditional fixed-outlay local resources (it may be simpler to budget for the latter).
Comparison to published bespoke algorithms
Using a similar development dataset of fundus photographs from the EyePACS screening service, Gulshan et al. achieved a better sensitivity of 90.3% (95% CI, 87.5–92.7%) at a similar specificity of 98.1% (95% CI, 97.8–98.5%) as compared with Amazon, Apple and Google CFDL modes, although the former study utilized a larger development dataset, consisting of 128,175 fundus photographs14. On a similar-sized EyePACS dataset development dataset to ours, Voets et al. reported a (sensitivity; specificity) of (93.6%; 92.0%) at a high-specificity operating point, and (90.6%; 84.7%) at a high-sensitivity operating point29. The Google and Amazon CFDL EyePACS models both demonstrated higher performance compared to the bespoke referenced model. Other published examples from this dataset predict either individual diabetic retinopathy grades or perform binarization of referable versus non-referable in a different manner, making direct performance comparison difficult30,31,32,33. Although the Messidor dataset was used for validation of a number of published models, we were unable to locate bespoke models that used it for development, preventing direct comparison29,34,35.
Bespoke OCT models developed on the Kermany dataset by the original authors demonstrated a (sensitivity; specificity) of (97.8%; 97.4%)16. Amazon and Google CFDL models demonstrated superior performances of (99.3%; 99.7%) and (97.8%; 99.2%), respectively, when utilizing this dataset. The Kaggle data science community has produced reports of similarly high bespoke model performance, although these are not peer-reviewed36. OCT models developed on the Waterloo dataset by Aggarwal et al. demonstrated a (sensitivity; specificity) of (86.0%; 96.5%) which improved to (94.0%; 98.5%) with data augmentation37. Amazon, Google, MedicMind and Microsoft CFDL platforms were able to produce models with comparable or superior results without manual data augmentation. Factors that may have led to differing performance between CFDL platforms and bespoke published models include CFDL’s lack of task-specific image augmentation pre-processing, inability to specify task-specific base models for transfer learning approaches, and the inherent performance variations resulting from bespoke model construction and tuning.
Limitations
Limitations are expected when comparing platforms with differing featuresets and reporting metrics. Although we attempted to report clinically meaningful metrics by generating contingency tables to calculate specificity and NPV, Microsoft did not provide a confusion matrix. Thus, our objective comparison focused on PPV and sensitivity, and the resulting F1 score, as these were the only metrics that could be generated from all platforms. Across all platforms, model explainability was deficient. Although this is not unique to CFDL, due to its automated nature CFDL has the potential to further reduce machine learning explainability. When one is not manually setting model parameters, it becomes increasingly difficult to discern which underlying model architectures and hyperparameters lead to differing performances. The platforms lacked important evaluation features such as image-level results for the validation set, which precludes post hoc analyses of additional image metadata such as source International Classification of Diabetic Retinopathy (ICDR) grades. Datasets were limited in the patient-level data they contained, so we were unable to ensure patient-level splits on all but the Kermany datasets. This leaves the potential for data leakage and falsely elevated performance metrics.
External validation is a critical step in the evaluation of AI models prior to implementation38,39. Varying levels of platform support for batch prediction precluded the ability to perform external validation with all but the Google and MedicMind platforms (Table 1). The importance of this capability cannot be understated, and the authors are unable to recommend platforms that do not have this feature. As of 27 August 2020, Google supports batch prediction through a command line interface, limiting its use by those without the relevant expertise. External validation performance demonstrated decreased specificity as compared with the internal evaluation datasets, generating increased false-positive RDR classification. Such models may need site-specific threshold tuning to local populations. Although we utilized varying modalities and datasets, the ability to generalize to similar datasets for validation is limited due to the unique labels and disease grading guidelines of each dataset. We were unable to locate a dataset that contained the same OCT labels as Kermany and Waterloo, and thus were unable to externally validate the respective OCT models. Dataset upload speed did not vary widely among platforms and was limited by the client internet connection upload speed; however, this was not systematically or quantitatively evaluated.
Although saliency maps offer some potential to provide clinical interpretability, their utility in this regard has yet to be proven. Plausible saliency maps are often provided in the clinical AI literature, but such maps may be prone to cherry picking. Even in representative cases, their interpretation is subjective and they do not provide semantic explanations. There is a need for more systematic clinical evaluation of these maps before they can be used in direct patient care40,41. For example, saliency maps in Fig. 3c,d erroneously highlight the B scan slice key as an important area for prediction.
Platform evaluation tasks and surveys were subjective in nature. As a result of time constraints, we were limited to three clinicians—one (Z.G.) a final-year medical student—performing this evaluation and survey. Meaningful statistical evaluation was both not possible and likely to contain bias influenced by technology brand preferences. The overall user experience was positive, so platform choice will probably be driven by feature availability.
Potential CFDL use cases
We believe our findings demonstrate the potential of CFDL for clinicians and researchers across a multitude of medical imaging modalities and tasks. Although the representative datasets in this study were ophthalmic in nature, due to their dimensionality, this demonstration of CFDL has the potential to scale significantly. OCT is an exemplar of cross-sectional imaging, with models discerning features and edges among monochromatic pixels—a similar task to X-ray, computed tomography (CT) and magnetic resonance imaging (MRI). Fundus photography tasks entail en-face hue, luminance and contract pattern detection, often discerning subtle pathology at the single-pixel level—a task that is comparable to dermatology and pathology image classification.
The use cases for CFDL are broad, and candidate low-risk tasks include dataset curation for researchers. Currently, a major pain-point in medical image analysis is data collation and cleaning. CFDL may prove to be a rapid and reliable method for differentiating images between left and right, gradable and ungradable, proper field of view, and the like, potentially becoming a big time saver for researchers. Models may be trained in the standard supervised fashion utilizing labelled data (for example, to label eye images as gradable or ungradable). This trained model can then be utilized as a research tool, deployed on new datasets or on prospectively curated data, to collate images that fit a criterion (for example, if wanting to curate gradable images)42. Similarly, clinicians may train CFDL models representative of subtle phenotypic variations in their local populations. Edge models, running locally on a device without requiring an internet connection, may be used as screening tools in rural and underserved areas after proper validation, and may entail simpler information governance structures. Use of CFDL is not limited to those without coding expertise, as computer engineers may rapidly train CFDL models as a baseline against which bespoke deep learning models could be iterated and tuned on. These potential use cases are not exhaustive, and more will be elucidated as clinicians and researchers gain an understanding of ML principles through the exploration of CFDL.
AI fundamentals are not taught in medical schools or prerequisite statistics courses, and most clinicians’ understanding of AI principles is understandably limited. Although obviating the need for coding expertise, CFDL platforms still require proper data stewardship, employing careful dataset curation, class balancing, representative patient-level splits, external validation and continued monitoring to detect model deterioration43. As CFDL exposes more clinicians and researchers to machine learning, their exploration of the benefits and pitfalls of these techniques will lead to a broader understanding of responsible and safe AI. Clinicians and researchers should be aware of the falsely increased performance that may occur from data leakage of patients from development to validation sets. They should ensure that validation set disease prevalence approximates that of their real-world use-case population. Furthermore, models evaluated and utilized on populations with differing demographics, image acquisition techniques and artefacts from the distribution of the initial validation dataset may demonstrate widely varying real-world performance. CFDL is but one of the educational tools for AI available to clinicians, who, in their patients’ interest, must evaluate the safety of AI-based medical devices coming to market.
CFDL is a robust framework, with the potential to democratize ML access for clinicians and researchers. The evaluation performed herein has the potential for application across a range of medical image classification tasks. Although some platforms struggle with large datasets, and explainability remains an issue, we have discovered high image classification performance across most platforms. Thus, platform selection will probably be driven by select highlighted features for efficient dataset management and comprehensive model evaluation. Although use cases are broad, the increased exposure to machine learning that CFDL provides to those without coding expertise will drive exploration of responsible AI practices.
Methods
Datasets and study design
We utilized four open-source de-identified ophthalmic imaging datasets to train deep learning models on six AutoML platforms for a total of 24 deep learning models. A search was performed for candidate publicly available datasets. Datasets were chosen that represented common ophthalmic diseases and representative clinical classifications. Convenience sampling was used, and both prevalence of prior community contributions (Kaggle) and citations were considered. Four datasets were selected, including two retinal fundus photograph datasets (Messidor-2, n = 1,744; EyePACS, n = 35,108) and two OCT datasets (Waterloo, n = 572; Kermany, n = 101,418) representing small and large dataset sizes for each respective modality16,44,45,46,47. Patient demographics and inclusion criteria for each of these datasets are published in accordance with the source datasets. Where patient-level statistics are not reported, they were not provided with source datasets.
Dataset details
Fundus datasets were re-categorized to the binary labels of RDR (comprising DR grades of moderate, severe, proliferative and/or the presence of diabetic macular oedema (DMO) and NRDR (the absence of RDR) to represent a clinically meaningful task performed in screening programmes and by regulatory-approved models13,48,49,50,51,52,53.
Messidor-2 consists of 1,744 fundus photographs in .png format. DR and DMO labels adjudicated by retina specialists were applied from the Kaggle adjudicated dataset44,45. Source DR grades were assigned according to the ICDR protocol and DMO was defined by hard exudates within one disc diameter of the fovea. Images were labelled as NRDR (n = 1,279) and RDR (n = 465). This dataset hereafter will be referred to as Messidor. EyePACS comprises 35,108 fundus photographs in .jpeg format from the resized EyePACS Kaggle dataset46. Source DR grades were assigned according to the ICDR protocol. Images were labelled as NRDR (n = 28,240) and RDR (n = 6,868). The Waterloo dataset comprises 572 OCT images in .jpeg format47. Source labels include age-related macular degeneration (AMD; n = 55), central serous retinopathy (CSR; n = 102), macular hole (MH; n = 102), DR (n = 107) and normal (n = 206). The Kermany dataset contains 101,418 OCT images in .jpeg format from 5,761 patients. Labels include choroidal neovascularization (CNV; n = 31,882), DMO (n = 11,165), drusen (n = 8,061) and normal (n = 50,310). The datasets are summarized in Table 3.
AutoML platform selection
Thirteen AutoML platforms were located based on searches from the Google search engine performed in the period September–December 2019. Candidate platforms included Amazon, Apple, Clarifai, Google, MedicMind, Microsoft, IBM Watson Visual Recognition, Baidu, Platform.ai, Datarobot, ProductAI/Malong, DeepCognition and Uber Ludwig. Candidate platforms were evaluated for characteristics including the lack of a coding requirement, availability of usage within the region (UK), English language and for including a free trial period. Reasons for exclusion are detailed in Supplementary Table 5. Three researchers (E.K., D.F. and Z.G.) with minimal to no coding experience spent a minimum of 4 h exploring each platform. Time was spent on user interface exploration, testing and reading documentation for each of the platforms. Six platforms (Amazon, Apple, Clarifai, Google, MedicMind and Microsoft) were selected for this study. The initial exploration was performed in September 2019 with review in August 2020, and does not consider more recent updates, which may have altered the features and performance of candidate platforms. MedicMind and Apple are free platforms. Where available, we utilized the free tiers of paid platforms. Paid tiers were used when free credits expired, and if paid tier allowed for longer model training (Microsoft).
Data processing
Training supervised deep learning models entails splitting datasets into training, validation and test sets. For the Kermany dataset, in which test and train splits were already performed by the source dataset publishers, this split was preserved when training the CFDL platforms that allowed manual setting of splits (Table 1). This ensured equitable comparison to published bespoke models developed from the same dataset20. The smaller Waterloo and Messidor datasets were randomly split into training and test (80% and 20%, respectively), while the larger EyePACS dataset was randomly split into 90% and 10%, respectively, for large dataset split ratios consistent with the Kermany dataset (test n = 1,000). For platforms that allowed manual setting of validation sets, we further subsampled from the training set by splitting training and validation into 90% and 10%, respectively. Equal proportions of diagnostic labels were preserved in each split to ensure the smaller datasets were not class imbalanced between splits. No patient-level data were provided for the Messidor, EyePACS and Waterloo datasets, so we were unable to ensure that patient-level splits were maintained. Duplicate images were automatically detected and excluded by the Microsoft and Google platforms. All deep learning models were trained for the maximum compute hours allowable on each platform. Platform early stopping features were employed, which automatically terminated training when no further model improvement was noted.
Data upload and labelling
Apple allows local data processing, but the remaining platforms required data upload, some allowing multiple methods depending on use-case (Table 1). The methods range from direct GUI upload via a cloud bucket interface or via shell scripting with prerequisite installation of a cloud software developer kit (SDK). All selected platforms offer a GUI-based upload for ease of use, and none requires programming skill. A variety of methods were utilized based on platform and dataset size.
Labelling was performed via folder upload with folders split by label (Amazon, Microsoft, Clarifai, MedicMind), via .csv files containing labels and cloud bucket locations (Google) or via local folders split by label (Apple).
Model training
Models were trained on all selected CFDL platforms (Amazon, Apple, Clarifai, Google, MedicMind, Microsoft) by clinicians E.K., Z.G. and D.F. One model was trained per dataset–platform pair. There were no computer system requirements for the usage of cloud-based platforms, as they trained and evaluated on cloud-hosted GPUs. The Apple platform is run locally, and requires MacOS with the XCode developer program installed. RDR versus NRDR is a binary classification, but, except for MedicMind, the evaluated CFDL platforms do not support training binary classification algorithms. The RDR versus NRDR task in the Messidor and EyePACS datasets was therefore trained as two independent classifications, that is, two distinct labels—RDR and NRDR.
Result metrics and statistical analysis
Graphpad Prism version 7 was used for statistical analysis. The CFDL platforms provide various model metrics including recall (sensitivity), non-weighted average precision (PPV) for given model thresholds, along with the area under the precision–recall curve (AUPRC) and F1 scores. Confusion matrices are provided by Apple, Clarifai, Google and MedicMind. We extracted label data and calculated F1 scores (Extended Data Fig. 1). Where possible, contingency tables were manually constructed to calculate clinical metrics including specificity (Table 2). Clarifai reports a confusion matrix for one fold of its k-fold cross-validation. MedicMind label specificity and sensitivity reports did not match the evaluation spreadsheet classifications, which were used to derive our evaluation metrics. We surmise that the former statistics are performed on the training set and not the test set. In February 2020, we made MedicMind aware of the confusion this may cause, and their advice was to use the evaluation spreadsheet download function, which we followed. Google and Microsoft report AUCPRC, while Clarifai reports AUC under the receiver operating characteristic curve (AUROC), making direct comparison of reported AUCs not possible. Models that allow threshold selection (Google and Clarifai) were evaluated with the default threshold of 0.5. Although points along the precision–recall curves may be mapped across a variety of thresholds, variations among platform confusion matrices and levels of reporting prevented us from directly comparing AUPRCs. The only platform to generate a graphical precision–recall curve was Google, against which each individual model’s precision and recall were plotted (Fig. 2). We adhered to the typical clinical accuracy terminology of sensitivity, specificity, PPV, NPV and accuracy. Qualitative platform surveys were scored on a five-point scale from 1 (very dissatisfied) to 5 (extremely satisfied) (Supplementary Table 4).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
All datasets utilized in this study were downloaded from publicly available sources and were not modified. Datasets may be accessed according to the references and the following DOIs: Kermany OCT, https://doi.org/10.17632/rscbjbr9sj.3; Waterloo OCT, https://doi.org/10.5683/SP2/W43PFI; Messidor 2, https://doi.org/10.1001/jamaophthalmol.2013.1743. All other data supporting the findings of this study are available within the paper and its Supplementary Information files.
Code availability
The code for the six utilized platforms is not made publicly available by the respective companies responsible for its development. However, the links to platforms evaluated are provided in Supplementary Table 5. Replication of results may be attempted on all platforms evaluated, which are explicitly free of charge, although updates to the respective backends can occur at any time.
References
Keane, P. A. & Sadda, S. R. Retinal imaging in the twenty-first century: state of the art and future directions. Ophthalmology 121, 2489–2500 (2014).
Pontikos, N. et al. Correspondence: trends in retina specialist imaging utilization from 2012 to 2016 in the United States Medicare fee-for-service population. Am. J. Ophthalmol. https://doi.org/10.1016/j.ajo.2019.09.021 (2019).
Panwar, N. et al. Fundus photography in the 21st century—a review of recent technological advances and their implications for worldwide healthcare. Telemed. J. E. Health 22, 198–208 (2016).
DCCT/EDIC Research Group et al. Frequency of evidence-based screening for retinopathy in type 1 diabetes. New Engl. J. Med. 376, 1507–1516 (2017).
Scanlon, P. H. The systematic DR screening in England for two million people with diabetes. Digital Teleretinal Screening https://doi.org/10.1007/978-3-642-25810-7_12 (2012).
DeVience, E., McMillan, B. D. & Gross, R. L. Screening for primary open-angle glaucoma (POAG). Int. Ophthalmol. Clin. 58, 1–9 (2018).
Tan, N. Y. Q., Friedman, D. S., Stalmans, I., Ahmed, I. I. K. & Sng, C. C. A. Glaucoma screening: where are we and where do we need to go? Curr. Opin. Ophthalmol. 31, 91–100 (2020).
Fujimoto, J. & Swanson, E. The development, commercialization, and impact of optical coherence tomography. Invest. Ophthalmol. Vis. Sci. 57, OCT1–OCT13 (2016).
Heier, J. S. et al. Intravitreal aflibercept for diabetic macular edema: 148-week results from the VISTA and VIVID studies. Ophthalmology 123, 2376–2385 (2016).
Campochiaro, P. A. et al. Ranibizumab for macular edema following branch retinal vein occlusion: six-month primary end point results of a phase III study. Ophthalmology 117, 1102–1112 (2010).
Liakopoulos, S. et al. ORCA study: real-world versus reading centre assessment of disease activity of neovascular age-related macular degeneration (nAMD). Br. J. Ophthalmol. https://doi.org/10.1136/bjophthalmol-2019-315717 (2020).
Castillo, M. M. et al. Optical coherence tomography for the monitoring of neovascular age-related macular degeneration: a systematic review. Ophthalmology 122, 399–406 (2015).
Ting, D. S. W. et al. Development and validation of a deep learning system for diabetic retinopathy and related eye diseases using retinal images from multiethnic populations with diabetes. JAMA 318, 2211–2223 (2017).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316, 2402–2410 (2016).
Fauw, J. D. et al. Clinically applicable deep learning for diagnosis and referral in retinal disease. Nat. Med. 24, 1342–1350 (2018).
Kermany, D. S. et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172, 1122–1131 (2018).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Liu, X. et al. A comparison of deep learning performance against health-care professionals in detecting diseases from medical imaging: a systematic review and meta-analysis. Lancet Digit. Health 1, e271–e297 (2019).
Gargeya, R. & Leng, T. Automated identification of diabetic retinopathy using deep learning. Ophthalmology 124, 962–969 (2017).
Faes, L. et al. Automated deep learning design for medical image classification by health-care professionals with no coding experience: a feasibility study. Lancet Digit. Health 1, e232–e242 (2019).
Perrault, R. The AI Index 2019 Annual Report (AI Index Steering Committee, Human-Centered AI Institute, Stanford University, 2019).
Porwal, P. et al. Indian Diabetic Retinopathy Image Dataset (IDRiD): a database for diabetic retinopathy screening research. Brown Univ. Dig. Addict. Theory Appl. 3, 25 (2018).
D’Amour, A. et al. Underspecification presents challenges for credibility in modern machine learning. Preprint at https://doi.org/10.1136/bjophthalmol-2019-315717 (2020).
Choo, J. & Liu, S. Visual analytics for explainable deep learning. IEEE Comput. Graph. Appl. 38, 84–92 (2018).
Ribeiro, M. T., Singh, S. & Guestrin, C. ‘Why should I trust you?’: explaining the predictions of any classifier. Preprint at https://arxiv.org/pdf/1602.04938.pdf (2016).
Wongsuphasawat, K. et al. Visualizing dataflow graphs of deep learning models in TensorFlow. IEEE Trans. Vis. Comput. Graph. 24, 1–12 (2018).
Vickers, A. J., Cronin, A. M., Elkin, E. B. & Gonen, M. Extensions to decision curve analysis, a novel method for evaluating diagnostic tests, prediction models and molecular markers. BMC Med. Inform. Decis. Mak. 8, 53 (2008).
Shah, N. H., Milstein, A. & Bagley, S. C. Making machine learning models clinically useful. JAMA https://doi.org/10.1001/jama.2019.10306 (2019).
Voets, M., Møllersen, K. & Bongo, L. A. Reproduction study using public data of: development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. PLoS ONE 14, e0217541 (2019).
Xie, L., Yang, S., Squirrell, D. & Vaghefi, E. Towards implementation of AI in New Zealand national diabetic screening program: cloud-based, robust, and bespoke. PLoS ONE https://doi.org/10.1371/journal.pone.0225015.t009 (2020).
Raju, M. et al. Development of a deep learning algorithm for automatic diagnosis of diabetic retinopathy. Stud. Health Technol. Inform. 245, 559–563 (2017).
Kwasigroch, A., Jarzembinski, B. & Grochowski, M. Deep CNN based decision support system for detection and assessing the stage of diabetic retinopathy. In 2018 International Interdisciplinary PhD Workshop (IIPhDW) 111–116 (IEEE, 2018).
Pratt, H., Coenen, F., Broadbent, D. M., Harding, S. P. & Zheng, Y. Convolutional neural networks for diabetic retinopathy. Procedia Comput. Sci. 90, 200–205 (2016).
Ramachandran, N., Hong, S. C., Sime, M. J. & Wilson, G. A. Diabetic retinopathy screening using deep neural network. Clin. Exp. Ophthalmol. 46, 412–416 (2018).
Sahlsten, J. et al. Deep learning fundus image analysis for diabetic retinopathy and macular edema grading. Sci. Rep. 9, 10750 (2019).
Mooney, P. Retinal OCT images (optical coherence tomography). Kaggle https://www.kaggle.com/paultimothymooney/kermany2018/code (2018).
Aggarwal, P. Machine learning of retinal pathology in optical coherence tomography images. J. Med. Artif. Intell. 2, 20 (2019).
Collins, G. S. et al. External validation of multivariable prediction models: a systematic review of methodological conduct and reporting. BMC Med. Res. Methodol. 14, 40 (2014).
Steyerberg, E. W. & Harrell, F. E. Jr Prediction models need appropriate internal, internal–external, and external validation. J. Clin. Epidemiol. 69, 245–247 (2016).
Adebayo, J. et al. Sanity checks for saliency maps. In Advances in Neural Information Processing Systems 31 (eds Bengio, S. et al.) 9505–9515 (Curran Associates, 2018).
Alqaraawi, A., Schuessler, M., Weiß, P., Costanza, E. & Berthouze, N. Evaluating saliency map explanations for convolutional neural networks: a user study. Preprint at https://arxiv.org/pdf/2002.00772.pdf (2020).
Nguyen, Q., Valizadegan, H. & Hauskrecht, M. Learning classification models with soft-label information. J. Am. Med. Inform. Assoc. 21, 501–508 (2014).
Pollard, T. J. et al. Turning the crank for machine learning: ease, at what expense? Lancet Digit. Health 1, e198–e199 (2019).
Decencière, E. et al. Feedback on a publicly distributed image database: the Messidor database. Image Anal. Stereol. 33, 231–234 (2014).
Krause, J. et al. Grader variability and the importance of reference standards for evaluating machine learning models for diabetic retinopathy. Ophthalmology 125, 1264–1272 (2018).
ilovescience Diabetic retinopathy (resized). Kaggle https://www.kaggle.com/tanlikesmath/diabetic-retinopathy-resized (2019).
Gholami, P., Roy, P., Parthasarathy, M. K. & Lakshminarayanan, V. OCTID: Optical Coherence Tomography Image Database. Preprint at https://arxiv.org/pdf/1812.07056.pdf (2018).
Bursell, S. E. et al. Stereo nonmydriatic digital-video color retinal imaging compared with early treatment diabetic retinopathy study seven standard field 35-mm stereo color photos for determining level of diabetic retinopathy. Ophthalmology 108, 572–585 (2001).
Scanlon, P. H. The English national screening programme for diabetic retinopathy 2003–2016. Acta Diabetol. 54, 515–525 (2017).
Abràmoff, M. D., Lavin, P. T., Birch, M., Shah, N. & Folk, J. C. Pivotal trial of an autonomous AI-based diagnostic system for detection of diabetic retinopathy in primary care offices. npj Digit. Med. 1, 39 (2018).
Bellemo, V. et al. Artificial intelligence using deep learning to screen for referable and vision-threatening diabetic retinopathy in Africa: a clinical validation study. Lancet Digit. Health 1, e35–e44 (2019).
Tufail, A. et al. Automated diabetic retinopathy image assessment software: diagnostic accuracy and cost-effectiveness compared with human graders. Ophthalmology 124, 343–351 (2017).
Ipp, E., Shah, V. N., Bode, B. W. & Sadda, S. R. 599-P: diabetic retinopathy (DR) screening performance of general ophthalmologists, retina specialists, and artificial intelligence (AI): analysis from a pivotal multicenter prospective clinical trial. Diabetes https://doi.org/10.2337/db19-599-P (2019).
Acknowledgements
This work was supported by a Springboard Grant from the Moorfields Eye Charity (E.K.) and a UK National Institute for Health Research (NIHR) Clinician Scientist Award (NIHR-CS–2014-12-023; P.A.K.). The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health. The funder of the study had no role in study design, data collection, data analysis, data interpretation or writing of the report. The corresponding author had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Author information
Authors and Affiliations
Contributions
E.K. and P.A.K. designed the study. E.K., D.F., Z.G. and G.Z. acquired data. E.K., D.F., Z.G., G.Z., N.P., X.L. and L.F. analysed data. E.K. wrote the first draft of the manuscript. S.K.W., P.A.K., A.K.D., S.G.F., H.K., G.M. and K.B. contributed to the writing and approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
E.K. has been a consultant for Google Health, the parent company of which (Google) has created one of the platforms evaluated in this study. P.A.K. has been a consultant for the artificial intelligence company, DeepMind, the parent company of which (Google) has created one of the platforms evaluated in this study. P.A.K. has received speaker fees from Heidelberg Engineering, Topcon, Carl Zeiss Meditec, Haag-Streit, Allergan, Novartis and Bayer. He has served on advisory boards for Novartis and Bayer and has been an external consultant for DeepMind and Optos. All other authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 F1 Scores of Each Model.
Scores are grouped by dataset (columns), and in alphabetical order by platform (rows), best performing model from each dataset bolded. 2 way ANOVA analysis demonstrated significant difference between Amazon vs Apple F1 scores, mean difference [95% CI]: 21.9[1.3,42.5]. Post-hoc analysis comparing platforms within each dataset showed significant differences between models generated from the Kermany dataset by Google vs Apple 45.8[4.6,87.0] and Amazon vs Apple 47.2[6.0,88.4].
Extended Data Fig. 2 AutoML Graphical User Interface.
Typical AutoML platform user interface components. Each component is represented by a figure pane.
Supplementary information
Supplementary Information
Supplementary Tables 1–5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Korot, E., Guan, Z., Ferraz, D. et al. Code-free deep learning for multi-modality medical image classification. Nat Mach Intell 3, 288–298 (2021). https://doi.org/10.1038/s42256-021-00305-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-021-00305-2
This article is cited by
-
Comparing code-free and bespoke deep learning approaches in ophthalmology
Graefe's Archive for Clinical and Experimental Ophthalmology (2024)
-
A survey on deep learning tools dealing with data scarcity: definitions, challenges, solutions, tips, and applications
Journal of Big Data (2023)
-
OCT-based deep-learning models for the identification of retinal key signs
Scientific Reports (2023)
-
Potato diseases detection and classification using deep learning methods
Multimedia Tools and Applications (2023)
-
Diabetic retinopathy classification for supervised machine learning algorithms
International Journal of Retina and Vitreous (2022)