Abstract
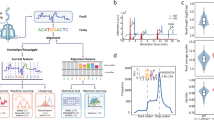
Research on DNA methylation on N6-adenine (6mA) in eukaryotes has received much recent attention. Recent studies have generated a large amount of 6mA genomic data, yet the role of DNA 6mA in eukaryotes remains elusive, or even controversial. We argue that the sparsity of DNA 6mA in eukaryotes, the limitations of current biotechnologies for 6mA detection and the sophistication of the 6mA regulatory mechanism together pose great challenges for elucidation of DNA 6mA. To exploit existing 6mA genomic data and address this challenge, here we develop a deep-learning-based algorithm for predicting potential DNA 6mA sites de novo from sequence at single-nucleotide resolution, with application to three representative model organisms, Arabidopsis thaliana, Drosophila melanogaster and Escherichia coli. Extensive experiments demonstrate the accuracy of our algorithm and its superior performance compared with conventional k-mer-based approaches. Furthermore, our saliency maps-based context analysis protocol reveals interesting cis-regulatory patterns around the 6mA sites that are missed by conventional motif analysis. Our proposed analytical tools and findings will help to elucidate the regulatory mechanisms of 6mA and benefit the in-depth exploration of their functional effects. Finally, we offer a complete catalogue of potential 6mA sites based on in silico whole-genome prediction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The SMART-seq data that support the findings of this study are available from GitHub (https://github.com/tanfei2007/DeepM6A/tree/master/Data). The sequencing data for A. thaliana are available on the GEO database under accession no. GSE149060. The data for different stages of D. melanogaster embryos are available on the GEO database under accession no. GSE86795. The raw data for 6mA-DNA-IP-Seq of D. melanogaster are available from https://trace.ddbj.nig.ac.jp/DRASearch/study?acc=SRP055483.
Code availability
The custom computer code is available from GitHub (https://github.com/tanfei2007/DeepM6A/tree/master/Code) under https://doi.org/10.5281/zenodo.3887349.
References
Heyn, H. & Esteller, M. An adenine code for DNA: a second life for N 6-methyladenine. Cell 161, 710–713 (2015).
Luo, G.-Z., Blanco, M. A., Greer, E. L., He, C. & Shi, Y. DNA N 6-methyladenine: a new epigenetic mark in eukaryotes? Nat. Rev. Mol. Cell Biol. 16, 705–710 (2015).
Zeng, H. & Gifford, D. K. Predicting the impact of non-coding variants on DNA methylation. Nucleic Acids Res 45, e99 (2017).
Feng, S. et al. Conservation and divergence of methylation patterning in plants and animals. Proc. Natl Acad. Sci. USA 107, 8689–8694 (2010).
Wu, T. P. et al. DNA methylation on N 6-adenine in mammalian embryonic stem cells. Nature 532, 329–333 (2016).
Fu, Y. et al. N 6-methyldeoxyadenosine marks active transcription start sites in Chlamydomonas. Cell 161, 879–892 (2015).
Greer, E. L. et al. DNA methylation on N 6-adenine in C. elegans. Cell 161, 868–878 (2015).
Liu, J. et al. Abundant DNA 6mA methylation during early embryogenesis of zebrafish and pig. Nat. Commun. 7, 13052 (2016).
Zhang, G. et al. N 6-methyladenine DNA modification in Drosophila. Cell 161, 893–906 (2015).
Barras, F. & Marinus, M. G. The great GATC: DNA methylation in E. coli. Trends Genet. 5, 139–143 (1989).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. In Proceedings of the 25th International Conference on Neural Information Processing Systems 1097–1105 (NIPS, 2012).
Lee, D. et al. A method to predict the impact of regulatory variants from DNA sequence. Nat. Genet. 47, 955–961 (2015).
Ji, G., Wu, X., Shen, Y., Huang, J. & Li, Q. Q. A classification-based prediction model of messenger RNA polyadenylation sites. J. Theor. Biol. 265, 287–296 (2010).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Rosenblatt, F. Principles of Neurodynamics. Perceptrons and the Theory of Brain Mechanisms (Cornell Aeronautical Lab, 1961).
Maaten, Lvd & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
He, S. et al. 6mA-DNA-binding factor Jumu controls maternal-to-zygotic transition upstream of Zelda. Nat. Commun. 10, 2219 (2019).
D’haeseleer, P. What are DNA sequence motifs? Nat. Biotechnol. 24, 423–425 (2006).
Bailey, T. L. & Elkan, C. Unsupervised learning of multiple motifs in biopolymers using expectation maximization. Mach. Learn. 21, 51–80 (1995).
Liang, Z. et al. DNA N 6-adenine methylation in Arabidopsis thaliana. Dev. Cell 45, 406–416 (2018).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, Z., Zhao, P. & XiaQ. Epigenetic methylations on N 6-adenine and N 6-adenosine with the same input but different output. Int. J. Mol. Sci. 20, 2931 (2019).
Musheev, M. U., Baumgartner, A., Krebs, L. & Niehrs, C. The origin of genomic N6-methyl-deoxyadenosine in mammalian cells. Nat. Chem. Biol. 16, 630–634 (2020).
He, K., Zhang, X., Ren, S. & Sun, J. Delving deep into rectifiers: surpassing human-level performance on ImageNet classification. In 2015 International Conference on Computer Vision 1026–1034 (IEEE, 2015)
Srivastava, N., Hinton, G. E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Simonyan, K., Vedaldi, A. & Zisserman, A. Deep inside convolutional networks: visualising image classification models and saliency maps. Workshop at International Conference on Learning Representations (2014).
Acknowledgements
We thank H. Liu for the partial data preprocessing. This study was supported by The Children’s Hospital of Philadelphia Endowed Chair in Genomic Research to H.H. and an Institutional Development Award to the Center for Applied Genomics from The Children’s Hospital of Philadelphia. This work was supported by Extreme Science and Engineering Discovery Environment (XSEDE) through allocation CIE160021 and CIE170034 (supported by National Science Foundation grant no. ACI-1548562).
Author information
Authors and Affiliations
Contributions
Z.W. and H.H. conceived and supervised the project. F.T., T.T. and X.H. designed the methods and conducted the experiments with input from L.G. T.T., X.Y., B.D.G., F.M. and F.T. conducted the validation experiments. F.T., T.T., Z.W. and H.H. wrote the manuscript. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–19, Discussion and Tables 11–13.
Supplementary Tables
Supplementary Tables 1–10 and 14–21.
Rights and permissions
About this article
Cite this article
Tan, F., Tian, T., Hou, X. et al. Elucidation of DNA methylation on N6-adenine with deep learning. Nat Mach Intell 2, 466–475 (2020). https://doi.org/10.1038/s42256-020-0211-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-020-0211-4