Abstract
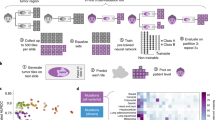
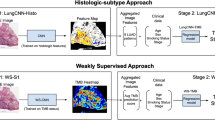
Tumour mutational burden (TMB) is an important biomarker for predicting the response to immunotherapy in patients with cancer. Gold-standard measurement of TMB is performed using whole exome sequencing (WES), which is not available at most hospitals because of its high cost, operational complexity and long turnover times. We have developed a machine learning algorithm, Image2TMB, which can predict TMB from readily available lung adenocarcinoma histopathological images. Image2TMB integrates the predictions of three deep learning models that operate at different resolution scales (×5, ×10 and ×20 magnification) to determine if the TMB of a cancer is high or low. On a held-out set of patients, Image2TMB achieves an area under the precision recall curve of 0.92, an average precision of 0.89, and has the predictive power of a targeted sequencing panel of ~100 genes. This study demonstrates that it is possible to infer genomic features from histopathology images, and potentially opens avenues for exploring genotype–phenotype relationships.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data used in this study are publicly available via the TCGA Research Network (https://www.cancer.gov/tcga).
Code availability
Our full approach, including data download, preprocessing and Image2TMB, is publicly available from Github (https://github.com/msj3/Image2TMB).
References
Garon, E. et al. Pembrolizumab for the treatment of non-small cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Pardoll, D. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Rizvi, N. et al. Activity and safety of nivolumab and an anti-PD-1 immune checkpoint inhibitor and for patients with advanced and refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol. 16, 257–265 (2015).
Chan, T. et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann. Oncol. 30, 44–56 (2018).
Gandara, D. et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat. Med. 24, 1441–1448 (2018).
Goodman, M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
Samstein, R. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Hellmann, M. et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 378, 2093–2104 (2018).
Steuer, C. & Ramalingam, S. Tumor mutation burden: leading immunotherapy to the era of precision medicine? J. Clin. Oncol. 36, 631–632 (2018).
Chalmers, Z. et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 9, 34 (2017).
Buchhalter, I. et al. Size matters: dissecting key parameters for panel-based tumor mutational burden analysis. Int. J. Cancer 144, 848–858 (2019).
Carpenter, A. et al. Cellprofiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
McQuin, C. et al. Cellprofiler 3.0: next-generation image processing for biology. Genome Biol. 16, e2005970 (2018).
Yu, K. et al. Association of omics features with histopathology patterns in lung adenocarcinoma. Cell Syst. 5, 620–627 (2017).
Yu, K. et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat. Commun. 7, 12474 (2016).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Ehteshami, B. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. Genome Biol. 318, 2199–2210 (2017).
Araújo, T. et al. Classification of breast cancer histology images using convolutional neural networks. PLoS ONE 12, e0177544 (2017).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Khosravi, P., Kazemi, E., Imielinski, M., Elemento, O. & Hajirasouliha, I. Deep convolutional neural networks enable discrimination of heterogeneous digital pathology images. EBioMedicine 27, 317–328 (2018).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (IEEE, 2016); preprint at https://arxiv.org/abs/1512.00567.
Russakovsky, O. et al. ImageNet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252 (2015).
Hellmann, M. et al. Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell 33, 843–852 (2018).
Kim, J., Kim, H., Kim, B., Lee, J. & Jang, H. Prognostic value of c-Met overexpression in pancreatic adenocarcinoma: a metaanalysis. Oncotarget 8, 73098–73104 (2017).
Offin, M. et al. Tumor mutation burden and efficacy of EGFR-tyrosine kinase inhibitors in patients with EGFR-mutant lung cancers. Clin. Cancer Res. 25, 1063–1069 (2019).
Koboldt, D. et al. Varscan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Acknowledgements
The presented results are based on data generated by the TCGA Research Network (https://www.cancer.gov/tcga). We thank R. Joshi, N. Neishaboori and E. Nohr for feedback and comments on the manuscript.
Author information
Authors and Affiliations
Contributions
M.S.J. conceived and performed the study and experiments. T.F.M. supervised the experiments. M.S.J. and T.F.M. wrote the manuscript. Both authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Tables 1–2, methods and references.
Rights and permissions
About this article
Cite this article
Jain, M.S., Massoud, T.F. Predicting tumour mutational burden from histopathological images using multiscale deep learning. Nat Mach Intell 2, 356–362 (2020). https://doi.org/10.1038/s42256-020-0190-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-020-0190-5
This article is cited by
-
Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
Nature Communications (2024)
-
Multi-omics and artificial intelligence predict clinical outcomes of immunotherapy in non-small cell lung cancer patients
Clinical and Experimental Medicine (2024)
-
Advancement in Machine Learning: A Strategic Lookout from Cancer Identification to Treatment
Archives of Computational Methods in Engineering (2023)
-
A Comprehensive Review of Markov Random Field and Conditional Random Field Approaches in Pathology Image Analysis
Archives of Computational Methods in Engineering (2022)
-
Review of deep learning: concepts, CNN architectures, challenges, applications, future directions
Journal of Big Data (2021)