Abstract
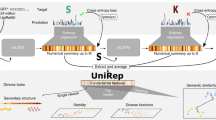
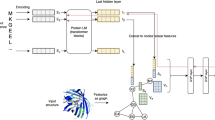
Proteins are nature’s most versatile molecular machines. Deep neural networks trained on large protein datasets have recently been used to tackle the unmet complexity of protein sequence–function relationships. The implicit knowledge contained in these networks represents a powerful, but thus far inaccessible, resource for understanding protein biology. Here, we show that occlusion-based sensitivity analysis can leverage the knowledge present in deep-neural-network-based protein sequence classifiers to identify functionally relevant parts of proteins. We first validated our approach by successfully predicting positions that mediate small molecule binding or catalytic activity across different protein classes. Next, we inferred the impact of point mutations on the activity of ERK and HRas, signalling factors frequently deregulated in cancer. Finally, we used our approach to identify engineering hotspots in CRISPR–Cas9 and anti-CRISPR protein AcrIIA4. Our work demonstrates how implicit knowledge in neural networks can be harnessed for protein functional dissection and protein engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Sensitivity analysis data for all presented proteins, including the ~800 proteins used to calculate spatial homogeneity of sphere variances, as well as weights for DeeProtein classifier, are available on Zenodo (https://doi.org/10.5281/zenodo.2577920 and https://doi.org/10.5281/zenodo.2574979). AcrIIA4–LOV2 expression vectors can be obtained from the corresponding authors on reasonable request.
Code availability
The code for DeeProtein, including scripts employed for sensitivity analysis, and code for mapping sensitivities to protein 3D structures in PyMol, is available on GitHub under MIT License (https://github.com/juzb/DeeProtein, https://doi.org/10.5281/zenodo.2619339). A stand-alone compute capsule covering central functions of DeeProtein is available on Code Ocean (https://doi.org/10.24433/CO.1473214.v1)65.
Change history
21 May 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper
References
Kulmanov, M., Khan, M. A., Hoehndorf, R. & Wren, J. DeepGO: predicting protein functions from sequence and interactions using a deep ontology-aware classifier. Bioinformatics 34, 660–668 (2018).
Jensen, L. J., Gupta, R., Staerfeldt, H. H. & Brunak, S. Prediction of human protein function according to Gene Ontology categories. Bioinformatics 19, 635–642 (2003).
You, R. et al. GOLabeler: improving sequence-based large-scale protein function prediction by learning to rank. Bioinformatics 34, 2465–2473 (2018).
Frasca, M. & Cesa Bianchi, N. Combining cost-sensitive classification with negative selection for protein function prediction. Preprint at https://arxiv.org/abs/1805.07331 (2018).
Szalkai, B. & Grolmusz, V. Near perfect protein multi-label classification with deep neural networks. Methods 132, 50–56 (2018).
Sinai, S., Kelsic, E., Church, G. M. & Nowak, M. A. Variational auto-encoding of protein sequences. Preprint at https://arxiv.org/abs/1712.03346 (2017).
Riesselman, A. J., Ingraham, J. B. & Marks, D. S. Deep generative models of genetic variation capture the effects of mutations. Nat. Methods 15, 816–822 (2018).
Fowler, D. M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Biswas, S. et al. Toward machine-guided design of proteins. Preprint at https://doi.org/10.1101/337154 (2018).
Fong, R. & Vedaldi, A. Interpretable explanations of black boxes by meaningful perturbation. Preprint at https://arxiv.org/abs/1704.03296(2017).
Kindermans, P.-J. et al. Learning how to explain neural networks: PatternNet and PatternAttribution. Preprint at https://arxiv.org/abs/1705.05598 (2017).
Grégoire, M., Samek, W. & Müller, K.-R. Methods for interpreting and understanding deep neural networks. Dig. Sig. Process. 73, 1–15 (2018)..
Radivojac, P. et al. A large-scale evaluation of computational protein function prediction. Nat. Methods 10, 221–227 (2013).
Arras, L., Horn, F., Montavon, G., Müller, K.-R. & Wojciech, S. “What is relevant in a text document?”: an interpretable machine learning approach. PLoS ONE 12, e0181142 (2017).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In IEEE Conference on Computer Vision and Pattern Recognition 770–778 (IEEE, 2016).
Boutet, E., Lieberherr, D., Tognolli, M., Schneider, M. & Bairoch, A. UniProtKB/Swiss-Prot. Methods Mol. Biol. 406, 89–112 (2007).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331–D338 (2017).
Cozzetto, D., Minneci, F., Currant, H. & Jones, D. T. FFPred 3: feature-based function prediction for all Gene Ontology domains. Sci. Rep. 6, 31865 (2016).
Gong, Q., Ning, W. & Tian, W. GoFDR: a sequence alignment based method for predicting protein functions. Methods 93, 3–14 (2016).
Zeiler, M. D. & Fergus, R. Visualizing and understanding convolutional networks. In European Conference on Computer Vision (eds Fleet, D., Pajdla, T., Schiele, B. & Tuytelaars, T.) 818–833 (Springer, 2014).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Zhang, L. et al. Functional role of histidine in the conserved His–x–Asp motif in the catalytic core of protein kinases. Sci. Rep. 5, 10115 (2015).
Samatar, A. A. & Poulikakos, P. I. Targeting RAS-ERK signalling in cancer: promises and challenges. Nat. Rev. Drug Discov. 13, 928–942 (2014).
Roskoski, R. Jr. ERK1/2 MAP kinases: structure, function, and regulation. Pharmacol. Res. 66, 105–143 (2012).
Kornev, A. P., Taylor, S. S. & Ten Eyck, L. F. A helix scaffold for the assembly of active protein kinases. Proc. Natl Acad. Sci. USA 105, 14377–14382 (2008).
Brenan, L. et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 17, 1171–1183 (2016).
Bandaru, P. et al. Deconstruction of the Ras switching cycle through saturation mutagenesis. eLife https://doi.org/10.7554/eLife.27810 (2017).
Richter, F. et al. Switchable Cas9. Curr. Opin. Biotechnol. 48, 119–126 (2017).
Ha, J. H. & Loh, S. N. Protein conformational switches: from nature to design. Chemistry 18, 7984–7999 (2012).
Stein, V. & Alexandrov, K. Synthetic protein switches: design principles and applications. Trends Biotechnol. 33, 101–110 (2015).
Hoffmann, M. D., Bubeck, F., Eils, R. & Niopek, D. Controlling cells with light and LOV. Adv. Biosyst. https://doi.org/10.1002/adbi.201800098 (2018).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Liu, J. J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Oakes, B. L. et al. Profiling of engineering hotspots identifies an allosteric CRISPR–Cas9 switch. Nat. Biotechnol. 34, 646–651 (2016).
Rauch, B. J. et al. Inhibition of CRISPR-Cas9 with bacteriophage proteins. Cell 168, 150–158 (2017).
Bubeck, F. et al. Engineered anti-CRISPR proteins for optogenetic control of CRISPR–Cas9. Nat. Methods 15, 924–927 (2018).
Basgall, E. M. et al. Gene drive inhibition by the anti-CRISPR proteins AcrIIA2 and AcrIIA4 in Saccharomyces cerevisiae. Microbiology 164, 464–474 (2018).
Dong, D. et al. Structural basis of CRISPR-SpyCas9 inhibition by an anti-CRISPR protein. Nature 546, 436–439 (2017).
Yang, H. & Patel, D. J. Inhibition mechanism of an anti-CRISPR suppressor AcrIIA4 targeting SpyCas9. Mol. Cell 67, 117–127 e115 (2017).
Shin, J. et al. Disabling Cas9 by an anti-CRISPR DNA mimic. Sci. Adv. 3, e1701620 (2017).
McReynolds, A. C. et al. Phosphorylation or mutation of the ERK2 activation loop alters oligonucleotide binding. Biochemistry 55, 1909–1917 (2016).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. Preprint at https://arxiv.org/abs/1703.01365 (2017).
Bach, S. et al. On pixel-wise explanations for non-linear classifier decisions by layer-wise relevance propagation. PLoS ONE 10, e0130140 (2015).
Shrikumar, A., Greenside, P. & Kundaje, A. Learning important features through propagating activation differences. In Proceedings of the 34th International Conference on Machine Learning 3145–3153 (PMLR, 2017).
Martín Abadi, A. A., et al. TensorFlow: large-scale machine learning on heterogeneous systems. Preprint at https://arxiv.org/abs/1603.04467 (2015).
Dong, H. et al. TensorLayer: a versatile library for efficient deep learning development. In Proceedings of the 25th ACM international conference on Multimedia 1201–1204 (ACM, 2017).
Glorot, X., Bordes, A. & Bengio, Y. Deep sparse rectifier neural networks. Proc. 14th International Conference on Artificial Intelligence and Statistics. Vol. 15, 35–323 (2011).
He, K., Zhang, X., Ren, S. & Sun, J. Delving deep into rectifiers: surpassing human-level performance on ImageNet classification. Preprint at https://arxiv.org/abs/1502.01852 (2015).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. ICML (2015).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
The UniProt consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. In International Conference on Learning Representations (ICLR, 2015).
Oliphant, E., Peterson, P. et al. SciPy: Open source scientific tools for Python, 2001–2019. SciPy http://www.scipy.org/ (2019).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J . Mol. Biol. 215, 403–410 (1990).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Hamelryck, T. & Manderick, B. PDB file parser and structure class implemented in Python. Bioinformatics 19, 2308–2310 (2003).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Chojnacki, S., Cowley, A., Lee, J., Foix, A. & Lopez, R. Programmatic access to bioinformatics tools from EMBL-EBI update: 2017. Nucleic Acids Res. 45, W550–W553 (2017).
Touw, W. G. et al. A series of PDB-related databanks for everyday needs. Nucleic Acids Res. 43, D364–D368 (2015).
The PyMOL Molecular Graphics System Version 2.0 (Schrödinger, 2019).
Upmeier zu Belzen, J. et al. Leveraging implicit knowledge in neural networks for functional dissection and engineering of proteins. Code Ocean https://doi.org/10.24433/CO.1473214.v1 (2019).
Acknowledgements
This work was funded by the Klaus Tschira foundation, the German Research Foundation (DFG) and the Federal Ministry of Education and Research. We thank J. Quittek and M. Niepert (both at NEC), T. Wollmann (IPMB, BioQuant and the German Cancer Research Center (DKFZ)) for helpful discussions and M. Hemberger (BioQuant) for support with IT and GPU cluster use. J.U.z.B., T.B., S.H., L.A., C.G., M.K., J.M., P.P., L.P., M.P., M.S., D.H., M.D.H., M.J., C.S., M.W., I.L., D.N. and R.E. represent the iGEM Team Heidelberg 2017.
Author information
Authors and Affiliations
Contributions
All members of the iGEM Team Heidelberg 2017 conceived the initial idea and J.U.z.B, T.B., S.H., I.L., D.N. and R.E. refined it. T.B., J.U.z.B. and S.H. implemented DeeProtein. J.U.z.B. performed sensitivity analysis. F.B. cloned AcrIIA4–LOV2 fusions and performed luciferase assays. J.U.z.B., T.B., S.H., F.B., D.N. and R.E. interpreted data. D.N. and R.E. jointly supervised the work. J.U.z.B., D.N. and R.E. wrote the paper with support from T.B. and S.H. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.B., M.D.H., D.N. and R.E. have filed a European Patent application (17196813.4) for the AcrIIA4–LOV2 constructs.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, Supplementary Tables 3–6, Supplementary Notes 1–4, Supplementary references,
Supplementary Table 1
Ligand binding sensitivity
Supplementary Table 2
Sensitivity for catalytic activity
Rights and permissions
About this article
Cite this article
Upmeier zu Belzen, J., Bürgel, T., Holderbach, S. et al. Leveraging implicit knowledge in neural networks for functional dissection and engineering of proteins. Nat Mach Intell 1, 225–235 (2019). https://doi.org/10.1038/s42256-019-0049-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-019-0049-9
This article is cited by
-
A hybrid deep learning model for classification of plant transcription factor proteins
Signal, Image and Video Processing (2023)
-
ECNet is an evolutionary context-integrated deep learning framework for protein engineering
Nature Communications (2021)