Abstract
Most research on human pancreatic islets is conducted on samples obtained from normoglycaemic or diseased brain-dead donors and thus cannot accurately describe the molecular changes of pancreatic islet beta cells as they progress towards a state of deficient insulin secretion in type 2 diabetes (T2D). Here, we conduct a comprehensive multi-omics analysis of pancreatic islets obtained from metabolically profiled pancreatectomized living human donors stratified along the glycemic continuum, from normoglycemia to T2D. We find that islet pools isolated from surgical samples by laser-capture microdissection display remarkably more heterogeneous transcriptomic and proteomic profiles in patients with diabetes than in non-diabetic controls. The differential regulation of islet gene expression is already observed in prediabetic individuals with impaired glucose tolerance. Our findings demonstrate a progressive, but disharmonic, remodelling of mature beta cells, challenging current hypotheses of linear trajectories toward precursor or transdifferentiation stages in T2D. Furthermore, through integration of islet transcriptomics with preoperative blood plasma lipidomics, we define the relative importance of gene coexpression modules and lipids that are positively or negatively associated with HbA1c levels, pointing to potential prognostic markers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq data were deposited in the NCBI Gene Expression Omnibus with GEO accession number GSE164416. Human genome reference assembly GRCh38 is publicly available.
The proteomics raw data sets and the MaxQuant output files generated and analysed throughout this study were deposited at the ProteomeXchange Consortium via the PRIDE partner repository with the project accession number PXD022561 (https://www.ebi.ac.uk/pride/archive/). Lipidomics data were deposited in the Zenodo database (zenodo.org, https://doi.org/10.5281/zenodo.4716063). Source data are provided with this paper.
References
Saeedi, P. et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas, 9th edition. Diabetes Res. Clin. Pract. 157, 107843 (2019).
Mizera, M. et al. Type 2 diabetes remission 5 years after laparoscopic sleeve Ggastrectomy: multicenter cohort study. Obes. Surg. 31, 980–986 (2020).
Lim, E. L. et al. Reversal of type 2 diabetes: normalisation of beta cell function in association with decreased pancreas and liver triacylglycerol. Diabetologia 54, 2506–2514 (2011).
Talchai, C., Xuan, S., Lin, H. V., Sussel, L. & Accili, D. Pancreatic β cell dedifferentiation as a mechanism of diabetic β cell failure. Cell 150, 1223–1234 (2012).
Wang, Z., York, N. W., Nichols, C. G. & Remedi, M. S. Pancreatic β cell dedifferentiation in diabetes and redifferentiation following insulin therapy. Cell Metab. 19, 872–882 (2014).
Cinti, F. et al. Evidence of β-cell dedifferentiation in human type 2 diabetes. J. Clin. Endocrinol. Metab. 101, 1044–1054 (2016).
American Diabetes Association. Classification and diagnosis of diabetes: standards of medical care in diabetes — 2020. Diabetes Care 43, S14–S31 (2020).
Barovic, M. et al. Metabolically phenotyped pancreatectomized patients as living donors for the study of islets in health and diabetes. Mol. Metab. 27, s1–s6 (2019).
Poitout, V. et al. A call for improved reporting of human islet characteristics in research articles. Diabetes 68, 209–211 (2019).
Ebrahimi, A. et al. Evidence of stress in β cells obtained with laser capture microdissection from pancreases of brain dead donors. Islets 9, 19–29 (2017).
Toyama, H., Takada, M., Suzuki, Y. & Kuroda, Y. Activation of macrophage-associated molecules after brain death in islets. Cell Transplant. 12, 27–32 (2003).
Negi, S. et al. Analysis of betacell gene expression reveals inflammatory signaling and evidence of dedifferentiation following human islet isolation and culture. PLoS ONE 7, 1–11 (2012).
Weir, G. C. Glucolipotoxicity, β-cells, and diabetes: the emperor has no clothes. Diabetes 69, 273–278 (2020).
Solimena, M. et al. Systems biology of the IMIDIA biobank from organ donors and pancreatectomised patients defines a novel transcriptomic signature of islets from individuals with type 2 diabetes. Diabetologia 61, 641–657 (2018).
Gerst, F. et al. The expression of aldolase B in islets is negatively associated with insulin secretion in humans. J. Clin. Endocrinol. Metab. 103, 4373–4383 (2018).
Khamis, A. et al. Laser capture microdissection of human pancreatic islets reveals novel eQTLs associated with type 2 diabetes. Mol. Metab. 24, 98–107 (2019).
Cohrs, C. M. et al. Dysfunction of persisting β cells is a key feature of early type 2 diabetes pathogenesis. Cell Rep. 31, 107469 (2020).
Viñuela, A. et al. Genetic variant effects on gene expression in human pancreatic islets and their implications for T2D. Nat. Commun. 11, 4912 (2020).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Taneera, J. et al. Identification of novel genes for glucose metabolism based upon expression pattern in human islets and effect on insulin secretion and glycemia. Hum. Mol. Genet. 24, 1945–1955 (2014).
Carrat, G. R. et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am. J. Hum. Genet. 100, 238–256 (2017).
Xin, Y. et al. RNA sequencing of single human islet cells reveals type 2 diabetes genes. Cell Metab. 24, 608–615 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Haythorne, E. et al. Diabetes causes marked inhibition of mitochondrial metabolism in pancreatic β-cells. Nat. Commun. 10, 2474 (2019).
Meier, F. et al. Online parallel accumulation–serial fragmentation (PASEF) with a novel trapped ion mobility mass spectrometer. Mol. Cell. Proteom. 17, 2534–2545 (2018).
Thorens, B. GLUT2, glucose sensing and glucose homeostasis. Diabetologia 58, 221–232 (2015).
Pipatpolkai, T., Usher, S., Stansfeld, P. J. & Ashcroft, F. M. New insights into KATP channel gene mutations and neonatal diabetes mellitus. Nat. Rev. Endocrinol. 16, 378–393 (2020).
Brunner, A. D. et al. Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation. Preprint at bioRxiv https://doi.org/10.1101/2020.12.22.423933 (2020).
Wiśniewski, J. R., Hein, M. Y., Cox, J. & Mann, M. A ‘proteomic ruler’ for protein copy number and concentration estimation without spike-in standards. Mol. Cell. Proteom. 13, 3497–3506 (2014).
Cox, J. & Mann, M. 1D and 2D annotation enrichment: a statistical method integrating quantitative proteomics with complementary high-throughput data. BMC Bioinformatics 13, S12 (2012).
Zitomer, N. C. et al. Ceramide synthase inhibition by fumonisin B1 causes accumulation of 1-deoxysphinganine. A novel category of bioactive 1-deoxysphingoid bases and 1-deoxydihydroceramides biosynthesized by mammalian cell lines and animals. J. Biol. Chem. 284, 4786–4795 (2009).
Boccard, J. & Rutledge, D. N. A consensus orthogonal partial least squares discriminant analysis (OPLS-DA) strategy for multiblock Omics data fusion. Anal. Chim. Acta 769, 30–39 (2013).
Boccard, J. & Rutledge, D. N. Iterative weighting of multiblock data in the orthogonal partial least squares framework. Anal. Chim. Acta 813, 25–34 (2014).
Campbell-Thompson, M. et al. Network for pancreatic organ donors with diabetes (nPOD): developing a tissue biobank for type 1 diabetes. Diabetes Metab. Res. Rev. 28, 608–617 (2012).
Kaestner, K. H., Powers, A. C., Naji, A. & Atkinson, M. A. NIH initiative to improve understanding of the pancreas, islet, and autoimmunity in type 1 diabetes: The Human Pancreas Analysis Program (HPAP). Diabetes 68, 1394–1402 (2019).
Ebrahimi, A. G. et al. Beta cell identity changes with mild hyperglycemia: implications for function, growth, and vulnerability. Mol. Metab. 35, 100959 (2020).
van der Meulen, T. et al. Virgin beta cells persist throughout life at a neogenic niche within pancreatic islets. Cell Metab. 25, 911–926.e6 (2017).
Lawlor, N. et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type–specific expression changes in type 2 diabetes. Genome Res. 27, 208–222 (2017).
Avrahami, D. et al. Single-cell transcriptomics of human islet ontogeny defines the molecular basis of β-cell dedifferentiation in T2D. Mol. Metab. 42, 101057 (2020).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Huynh, K. et al. High-throughput plasma lipidomics: detailed mapping of the associations with cardiometabolic risk factors. Cell Chem. Biol. 26, 71–84(2019).
Suvitaival, T. et al. Lipidome as a predictive tool in progression to type 2 diabetes in Finnish men. Metabolism 78, 1–12 (2018).
Wigger, L. et al. Plasma dihydroceramides are diabetes susceptibility biomarker candidates in mice and humans. Cell Rep. 18, 2269–2279 (2017).
Mamtani, M. et al. Lipidomic risk score independently and cost-effectively predicts risk of future type 2 diabetes: results from diverse cohorts. Lipids Health Dis. 15, 67 (2016).
Sturm, D. et al. Improved protocol for laser microdissection of human pancreatic islets from surgical specimens. J. Vis. Exp. 50231 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal https://doi.org/10.14806/ej.17.1.200 (2011).
Wingett, S. W. & Andrews, S. FastQ Screen: a tool for multi-genome mapping and quality control. F1000Research 7, 1338 (2018).
Davis, M. P. A., van Dongen, S., Abreu-Goodger, C., Bartonicek, N. & Enright, A. J. Kraken: a set of tools for quality control and analysis of high-throughput sequence data. Methods 63, 41–49 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Smyth, G. K. et al. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Research 5, ISCB Comm J-1408 (2018).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omi. A J. Integr. Biol. 16, 284–287 (2012).
Surma, M. A. et al. An automated shotgun lipidomics platform for high throughput, comprehensive, and quantitative analysis of blood plasma intact lipids. Eur. J. Lipid Sci. Technol. 117, 1540–1549 (2015).
Smilde, A. K., Kiers, H. A. L., Bijlsma, S., Rubingh, C. M. & Van Erk, M. J. Matrix correlations for high-dimensional data: the modified RV-coefficient. Bioinformatics 25, 401–405 (2009).
Bylesjö, M., Rantalainen, M., Nicholson, J. K., Holmes, E. & Trygg, J. K-OPLS package: Kernel-based orthogonal projections to latent structures for prediction and interpretation in feature space. BMC Bioinformatics 9, 106 (2008).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Prianichnikov, N. et al. Maxquant software for ion mobility enhanced shotgun proteomics. Mol. Cell. Proteom. 19, 1058–1069 (2020).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteom. 13, 2513–2526 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Hu, J., Ge, H., Newman, M. & Liu, K. OSA: A fast and accurate alignment tool for RNA-seq. Bioinformatics 28, 1933–1934 (2012).
Li, B., Ruotti, V., Stewart, R. M., Thomson, J. A. & Dewey, C. N. RNA-seq gene expression estimation with read mapping uncertainty. Bioinformatics 26, 493–500 (2009).
Acknowledgements
We wish to thank L. Groop, E. Ahlqvist, S. Speier, T. Chavakis, R. Scharfmann and A. Schevchenko for discussion; S. Pradervand for advice on RNA-seq processing and QC; C. Rocher for Bioinformatics analysis support; K. Pfriem for administrative assistance. This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement no. 115881 (RHAPSODY). This joint undertaking receives support from the European Union’s Horizon 2020 research and innovation program and EFPIA. This work is further supported by the Swiss State Secretariat for Education‚ Research and Innovation (SERI) under contract number 16.0097-2. Work in the Solimena lab is also supported with funds from the German Ministry of Education and Research to the German Center for Diabetes Research (DZD). The opinions expressed and arguments employed herein do not necessarily reflect the official views of these funding bodies.
Author information
Authors and Affiliations
Contributions
J. W. and M. D., participant recruitment and surgery, provision of clinical data; E. S., N. K. and D. F., sample collection and processing, data entry; D. A., pathology; M .B., N. K. and E. S., patient database management and selection; A.-D. B. and M. M., proteomics; M. L., A. D., RNA-seq, C. L. Q., P. B. S. H, P. D., C. Klose, M. G., K. S., lipidomics and sphingolipidomics; L. W., M. B., A.-D. B., F. Marzetta., F. Mehl, F. B. and C. Kessler., analysis and integration of multi-omics data; E. B., autoantibody test; A. S., data in mouse tissue and cell lines; M.B., immunofluorescence stainings and antibody validation; B. T., D. A., J. W., A. S., M. M., M. I. and M. S., conceptual insights and provision of funds; L. W., M. B., A.-D. B., F. Marzetta, F. Mehl, A. S., M. I., M. M. and M. S., writing of the manuscript. All authors read, revised and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K. S. is CEO of Lipotype. K. S. and C. K. are shareholders of Lipotype. M. J. G. is an employee of Lipotype. P. B. S. H. and P. D. are employees of Servier. A. S. is an employee of Sanofi-Aventis Deutschland. The other authors declare no conflict of interest.
Additional information
Peer review information Nature Metabolism thanks Anna Gloyn, Huiyong Yin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
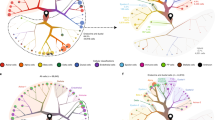
Extended Data Fig. 1 Deconvolution of cell types based on RNA-Seq data.
a, Cell-type proportions by sample, as estimated with DeconRNASeq. Beta cells, orange; alpha cells, green; delta cells, blue; gamma cells, purple. b, Sample distribution across each cell type proportion. Highlighted are samples presenting a cell type specific gene being the most expressed. Marker genes were GCG and TTR for alpha cells, INS for beta cells, SST for delta cells, and PPY for gamma cells.
Extended Data Fig. 2 Differential gene expression analysis between glycemic groups in the entire cohort.
a-b, Gene expression profile (a) and GSEA analysis (b) of DE genes between IGT, T3cD or T2D and ND PPP. Results are similar to those shown in Fig. 2bc, but obtained from the entire cohort of 133 PPP.
Extended Data Fig. 3 Validation experiments pertaining ALDOB.
a-b, ALDOB expression (RNAseq, Illumina) in (a) islets from 13-week-old male db/db, db/+ mice and C57Bl6 mice (3 animals/strain, 3 independent RNA extractions) or (b) mouse αTC1 clone 6 alpha and Min6s4 beta cell lines (4 independent RNA extractions/cell line). Boxplot spans from 25th until 75th percentile, with centerline at median, whiskers extend to the most extreme data point which is no more than 1.5 times the length of the box away from the box. c, Western blot of MinIN6 single cell-derived clones with antibodies against ALDOB and ALDOA. Framed lanes mark ALDOB knockout clones as verified by site-specific sequencing.
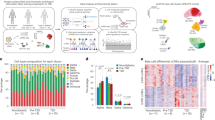
Extended Data Fig. 4 Extended analysis of proteomic data.
a, Hierarchical clustering of protein expression correlations in all biological replicates highlighting the technical and biological reproducibility of our proteome data set. b, Distribution of differentially expressed proteins in islets of T2D and ND PPP across chromosomes. c, Ranked ABCC8 protein expression levels across T2D and ND subjects. T2D are highlighted in orange, ND are highlighted in blue. Patient 118 was treated with Glimepiride; Patient 87 was treated with Mitiglinide; Patient 197 was treated with Glibenclamide. d, Transcriptome to Proteome abundance correlation (Pearson correlation coefficient = 0.3). e, Differential expression analysis of pancreatic islet transcriptomes of ND vs T2D PPP. (Orange: Significantly upregulated in diseased; Blue: Significantly downregulated in diseased).
Supplementary information
Supplementary Information
Supplementary Tables 1, 3, 8–12
Supplementary Tables
Supplementary Tables 2, 4–7, 13–16
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Wigger, L., Barovic, M., Brunner, AD. et al. Multi-omics profiling of living human pancreatic islet donors reveals heterogeneous beta cell trajectories towards type 2 diabetes. Nat Metab 3, 1017–1031 (2021). https://doi.org/10.1038/s42255-021-00420-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00420-9
This article is cited by
-
Gene expression analysis reveals diabetes-related gene signatures
Human Genomics (2024)
-
Web-based multi-omics integration using the Analyst software suite
Nature Protocols (2024)
-
Integration of polygenic and gut metagenomic risk prediction for common diseases
Nature Aging (2024)
-
Inter-organ crosstalk during development and progression of type 2 diabetes mellitus
Nature Reviews Endocrinology (2024)
-
Genomic insights into the comorbidity between type 2 diabetes and schizophrenia
Schizophrenia (2024)