Abstract
Global histone acetylation varies with changes in the nutrient and cell cycle phases; however, the mechanisms connecting these variations are not fully understood. Herein, we report that nutrient-related and cell-cycle-regulated nuclear acetate regulates global histone acetylation. Histone deacetylation-generated acetate accumulates in the nucleus and induces histone hyperacetylation. The nuclear acetate levels were controlled by glycolytic enzyme triosephosphate isomerase 1 (TPI1). Cyclin-dependent kinase 2 (CDK2), which is phosphorylated and activated by nutrient-activated mTORC1, phosphorylates TPI1 Ser 117 and promotes nuclear translocation of TPI1, decreases nuclear dihydroxyacetone phosphate (DHAP) and induces nuclear acetate accumulation because DHAP scavenges acetate via the formation of 1-acetyl-DHAP. CDK2 accumulates in the cytosol during the late G1/S phases. Inactivation or blockade of nuclear translocation of TPI1 abrogates nutrient-dependent and cell-cycle-dependent global histone acetylation, chromatin condensation, gene transcription and DNA replication. These results identify the mechanism of maintaining global histone acetylation by nutrient and cell cycle signals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw data for Figs. 1–7 and Extended Data Figs. 1–9 are available from the corresponding authors on request. Source data are provided with this paper.
Code availability
Code for Prism 6.0 (GraphPad Software) is available from the corresponding authors on request.
References
Rowicka, M., Kudlicki, A., Tu, B. P. & Otwinowski, Z. High-resolution timing of cell cycle-regulated gene expression. Proc. Natl Acad. Sci. USA 104, 16892–16897 (2007).
Ferguson, B. M., Brewer, B. J., Reynolds, A. E. & Fangman, W. L. A yeast origin of replication is activated late in S-phase. Cell 65, 507–515 (1991).
Raghuraman, M. K., Brewer, B. J. & Fangman, W. L. Cell cycle-dependent establishment of a late replication program. Science 276, 806–809 (1997).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Phillips, D. M. The presence of acetyl groups of histones. Biochem. J. 87, 258–263 (1963).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Eberharter, A. & Becker, P. B. Histone acetylation: a switch between repressive and permissive chromatin–second in review series on chromatin dynamics. EMBO Rep. 3, 224–229 (2002).
Kruhlak, M. J. et al. Regulation of global acetylation in mitosis through loss of histone acetyltransferases and deacetylases from chromatin. J. Biol. Chem. 276, 38307–38319 (2001).
Krebs, J. E., Kuo, M. H., Allis, C. D. & Peterson, C. L. Cell cycle-regulated histone acetylation required for expression of the yeast HO gene. Gene Dev. 13, 1412–1421 (1999).
Kurat, C. F. et al. Cell cycle-regulated oscillator coordinates core histone gene transcription through histone acetylation. Proc. Natl Acad. Sci. USA 111, 14124–14129 (2014).
Yang, X. J. & Seto, E. HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention. Oncogene 26, 5310–5318 (2007).
Bolduc, J. F., Hany, L., Barat, C., Ouellet, M. & Tremblay, M. J. Epigenetic metabolite acetate inhibits class I/II histone deacetylases, promotes histone acetylation and increases HIV-1 integration in CD4+ T cells. J. Virol. 91, e01943-16 (2017).
Wellen, K. E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Li, X. J., Qian, X. & Lu, Z. M. Local histone acetylation by ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Autophagy 13, 1790–1791 (2017).
Bulusu, V. et al. Acetate recapturing by nuclear acetyl-CoA synthetase 2 prevents loss of histone acetylation during oxygen and serum limitation. Cell Rep. 18, 647–658 (2017).
Li, X. J. et al. Nucleus-translocated ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Mol. Cell 66, 684–697 (2017).
Boon, R., Silveira, G. G. & Mostoslavsky, R. Nuclear metabolism and the regulation of the epigenome. Nat. Metab. 2, 1190–1203 (2020).
Sutendra, G. et al. A nuclear pyruvate dehydrogenase complex is important for the generation of acetyl-CoA and histone acetylation. Cell 158, 84–97 (2014).
de Boer, V. C. J. & Houten, S. M. A mitochondrial expatriate: nuclear pyruvate dehydrogenase. Cell 158, 9–10 (2014).
Friis, R. M. et al. A glycolytic burst drives glucose induction of global histone acetylation by picNuA4 and SAGA. Nucleic Acids Res. 37, 3969–3980 (2009).
Yu, Z. et al. PKM2 Thr454 phosphorylation increases its nuclear translocation and promotes xenograft tumor growth in A549 human lung cancer cells. Biochem. Biophys. Res. Commun. 473, 953–958 (2016).
Jiang, Y. et al. PKM2 regulates chromosome segregation and mitosis progression of tumor cells. Mol. Cell 53, 75–87 (2014).
Lee, J. W., Kim, H. K., Han, Y. M. & Kim, J. H. Pyruvate kinase isozyme type M2 (PKM2) interacts and cooperates with Oct-4 in regulating transcription. Int. J. Biochem. Cell Biol. 40, 1043–1054 (2008).
Mouveaux, T. et al. Nuclear glycolytic enzyme enolase of Toxoplasma gondii functions as a transcriptional regulator. PLoS ONE 9, e105820 (2014).
Yalcin, A. et al. Nuclear targeting of 6-phosphofructo-2-kinase (PFKFB3) increases proliferation via cyclin-dependent kinases. J. Biol. Chem. 284, 24223–24232 (2009).
Zhao, S. et al. Regulation of cellular metabolism by protein lysine acetylation. Science 327, 1000–1004 (2010).
Xu, W. S., Parmigiani, R. B. & Marks, P. A. Histone deacetylase inhibitors: molecular mechanisms of action. Oncogene 26, 5541–5552 (2007).
Nagaraj, R. et al. Nuclear localization of mitochondrial TCA cycle enzymes as a critical step in mammalian zygotic genome activation. Cell 168, 210–223 (2017).
Zaret, K. Micrococcal nuclease analysis of chromatin structure. Curr. Protoc. Mol. Biol. 21, 21.1 (2005).
Chang, M. L. et al. Human triosephosphate isomerase deficiency resulting from mutation of Phe-240. Am. J. Hum. Genet. 52, 1260–1269 (1993).
Castillo, J. et al. Genomic and proteomic dissection and characterization of the human sperm chromatin. Mol. Hum. Reprod. 20, 1041–1053 (2014).
de Mateo, S., Castillo, J., Estanyol, J. M., Ballesca, J. L. & Oliva, R. Proteomic characterization of the human sperm nucleus. Proteomics 11, 2714–2726 (2011).
Woo, R. A. & Poon, R. Y. Cyclin-dependent kinases and S phase control in mammalian cells. Cell Cycle 2, 316–324 (2003).
Stevenson-Lindert, L. M., Fowler, P. & Lew, J. Substrate specificity of CDK2–cyclin A. What is optimal? J. Biol. Chem. 278, 50956–50960 (2003).
Sielecki, T. M., Boylan, J. F., Benfield, P. A. & Trainor, G. L. Cyclin-dependent kinase inhibitors: useful targets in cell cycle regulation. J. Med. Chem. 43, 1–18 (2000).
Holy, J., Lamont, G. & Perkins, E. Disruption of nucleocytoplasmic trafficking of cyclin D1 and topoisomerase II by sanguinarine. BMC Cell Biol. 7, 13 (2006).
Brown, N. R. et al. Effects of phosphorylation of threonine 160 on cyclin-dependent kinase 2 structure and activity. J. Biol. Chem. 274, 8746–8756 (1999).
Slepokura, K. & Lis, T. Dihydroxyacetone phosphate, DHAP, in the crystalline state: monomeric and dimeric forms. Carbohydr. Res. 345, 512–529 (2010).
Cai, L., Sutter, B. M., Li, B. & Tu, B. P. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol. Cell 42, 426–437 (2011).
Orosz, F. et al. Enhanced association of mutant triosephosphate isomerase to red cell membranes and to brain microtubules. Proc. Natl Acad. Sci. USA 97, 1026–1031 (2000).
Orozco, J.M. et al. Dihydroxyacetone phosphate signals glucose availability to mTORC1. Nat Metab 2, 893–901 (2020).
Mamczur, P., Dus, D. & Dzugaj, A. Colocalization of aldolase and FBPase in cytoplasm and nucleus of cardiomyocytes. Cell Biol. Int. 31, 1122–1130 (2007).
Mamczur, P. & Dzugaj, A. Aldolase A is present in smooth muscle cell nuclei. Acta Biochim. Pol. 55, 799–805 (2008).
Saez, D. E. & Slebe, J. C. Subcellular localization of aldolase B. J. Cell. Biochem. 78, 62–72 (2000).
Mamczur, P., Rakus, D., Gizak, A., Dus, D. & Dzugaj, A. The effect of calcium ions on subcellular localization of aldolase–FBPase complex in skeletal muscle. FEBS Lett. 579, 1607–1612 (2005).
Hajra, A. K. Dihydroxyacetone phosphate acyltransferase. Biochim. Biophys. Acta 1348, 27–34 (1997).
Yamashita, A. et al. Glycerophosphate/acylglycerophosphate acyltransferases. Biology 3, 801–830 (2014).
Hadzhiev, Y. et al. A cell cycle-coordinated polymerase II transcription compartment encompasses gene expression before global genome activation. Nat. Commun. 10, 691 (2019).
Neganova, I. et al. An important role for CDK2 in G1 to S checkpoint activation and DNA damage response in human embryonic stem cells. Stem Cells 29, 651–659 (2011).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Guertin, D. A. & Sabatini, D. M. Defining the role of mTOR in cancer. Cancer Cell 12, 9–22 (2007).
Malumbres, M. & Barbacid, M. Cell cycle, CDKs and cancer: a changing paradigm. Nat. Rev. Cancer 9, 153–166 (2009).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Shechter, D., Dormann, H. L., Allis, C. D. & Hake, S. B. Extraction, purification and analysis of histones. Nat. Protoc. 2, 1445–1457 (2007).
Dai, H., Xiao, C., Liu, H., Hao, F. & Tang, H. Combined NMR and LC–DAD-MS analysis reveals comprehensive metabonomic variations for three phenotypic cultivars of Salvia Miltiorrhiza Bunge. J. Proteome Res. 9, 1565–1578 (2010).
Monier, K., Armas, J. C. G., Etteldorf, S., Ghazal, P. & Sullivan, K. F. Annexation of the interchromosomal space during viral infection. Nat. Cell Biol. 2, 661–665 (2000).
Maul, G. G. & Deaven, L. Quantitative-determination of nuclear-pore complexes in cycling cells with differing DNA content. J. Cell Biol. 73, 748–760 (1977).
Sancak, Y. et al. PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol. Cell 25, 903–915 (2007).
Lambeir, A. M., Opperdoes, F. R. & Wierenga, R. K. Kinetic properties of triose-phosphate isomerase from Trypanosoma brucei brucei—a comparison with the rabbit muscle and yeast enzymes. Eur. J. Biochem. 168, 69–74 (1987).
Noursadeghi, M. et al. Quantitative imaging assay for NF-κB nuclear translocation in primary human macrophages. J Immunol. Methods. 329, 194–200 (2008).
Sancho, M., Diani, E., Beato, M. & Jordan, A. Depletion of human histone H1 variants uncovers specific roles in gene expression and cell growth. PLoS Genet. 4, e1000227 (2008).
Acknowledgements
This work was supported by grants from the State Key Development Programs of China (2018YFA0801300, 2018YFA0800300, 2018YFC1004700, 2019YFA0801900 and 2020YFA0803600), the National Science Foundation of China (nos. 31821002, 31930062 and 91857000), Shanghai Rising-Star Program (18QA1400300), Medical and Health Commission of Shanghai (2018YQ36), Innovation-oriented Science and Technology Grant from Key Laboratory of Reproduction Regulation of NHC (CX2017-0X) and a grant from the Key Laboratory of Reproduction Regulation of NPFPC.
Author information
Authors and Affiliations
Contributions
S.-M.Z. and W.X. conceived the project. S.-M.Z. and W.X. supervised experiments, analysed the data and wrote the manuscript. J.-J.Z., T.-T.F., Y.-Z.M., M.W., Y.L., P.-C.L., Y.-Y.Y. and J.-Y.Z. carried out the cellular, biochemical and molecular biologic analysis; J.-L.H. and M.Z. synthesized 1-acetyl-DHAP; T.-T.F., J.-J.Z., L.Z., Y.-P.A., G.-Q.Y., C.Z. and J.Y. performed the LC–MS, NMR and ESI/MS analysis. All the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks David Sabatini and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: George Caputa.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nuclear acetate regulates global histone acetylation (related to Fig. 1).
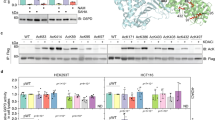
a, HeLa cells were synchronized to G1/S boundary with DTB. The released cells were collected at different time and the percentage of cells at different cell cycle phases were determined by flow cytometry (upper) and quantified (middle and bottom)), the gating strategy of flow cytometry was shown in Extended Data Fig. 6b.b, Verification of site-specific histone acetylation antibodies. The acetylation levels of site-specific acetylation levels HeLa cells and TSA-treated HeLa cells were probed by correspondent antibodies. c, Stability of Acetyl-CoA in nuclear and post nuclear fractions. 0.1 mM acetyl-CoA was added to PBS that contained 10% nuclear or post nuclear lysate, remaining acetyl-CoA levels in the mix was determined at time indicated points. n = 3 biologically independent samples (Nuclear: P = 0.0983, 0.0693, 0.0708, 0.0552; Post-Nuclear: P = 0.5527, 0.0621, 0.1039, 0.0707). d, The effect of ACLY knockdown and its effects on histone acetylation were detected. e, The synchronization of HeLa cells to G2/M boundary was confirmed by flow cytometry. f, The effect of ST045849 to specifically block nPDC translocation were detected in HeLa cells.
Extended Data Fig. 2 TPI1 regulates global histone acetylation (Related to Fig. 2).
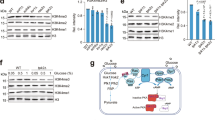
a, TPI1 overexpression upregulated histone acetylation. The global acetylation levels were compared between HeLa cells and HeLa cells overexpressing TPI1. The levels of endogenous (lower bands) and overexpressed (upper bands) TPI1 were indicated. b-c, TPI1 regulates global histone acetylation. The effects of TPI1 overexpression (b) and TPI1 knockdown with small hairpin RNA (c) were detected in HepG2, HEK293T and MCF7 cells. d, TPI1 and CDK2 knockdown inhibited cell proliferation. The growth of HeLa cells and HeLa cells with either TPI1 or CDK2 knocked down was compared. n = 3 biologically independent samples (P = 0.0025, 0.0003). e, TPI1 knockdown altered nuclear DHAP and acetate levels. Levels of acetyl-CoA, DHAP and acetate were compared in nuclear and post-nuclear fractions of HeLa cells and TPI1 knockdown HeLa cells. n = 3 biologically independent samples (P = 0.0061, 0.0006, 0.7426, 0.0201, 0.0158, 0.2641).
Extended Data Fig. 3 TPI1 regulates global histone acetylation via altering DHAP (Related to Fig. 2).
a, TPI1F278L is defective in glycolytic activity. The specific activities of TPI1 and TPI1F278L were analyzed, relative (to TPI1) specific activity of both enzymes were presented. n = 3 biologically independent experiments (P < 0.0001). b, TPI1F278L localized to nucleus and cytoplasm. The cellular compartmentation of TPI1F278L was detected by immunofluorescence. Scale bar: 10um. c, TPI1 overexpression decreased HDACs activities. The HDACs activities were compared between nucleus of HeLa cells and of HeLa cells overexpressing TPI1. n = 3 biologically independent samples (P = 0.0009). d, HDACs revered TPI1 effects on histone acetylation. The pan- and site specific histone acetylation levels were compared among HeLa cells, TPI1-expressing HeLa cells and HeLa cells that co-expressing TPI1 and each of HDAC1, HDAC2 or HDAC3. e, DHAP treatment increased nuclear DHAP levels. 0.5 mM DHAP was supplemented in the DMEM that was used to culture HeLa cells. The nuclear DHAP levels were measured for both untreated and treated cells. n = 3 biologically independent samples (P < 0.0001). f, DHAP decrease global histone acetylation. Histone acetylation levels were detected in HEK293T cells cultured without and with DHAP as indicated concentrations in the culture media. g, GAP treatment increased nuclear GAP levels. 0.5 mM GAP was supplemented in the DMEM that was used to culture HEK293T cells. The nuclear GAP levels were measured for both untreated and treated cells. n = 3 biologically independent samples (P < 0.0001).
Extended Data Fig. 4 CDK2 phosphorylates TPI1 to promote its nucleus translocation (Related to Fig. 3).
a, TPI1 localize to nucleus and cytosol. Immunofluorescence staining of HeLa and HEK293T cells with DAPI and anti-TPI1 antibodies. Scale bar: 10 μm. b-c, CDK2 interacted with TPI1. The interaction between CDK2 and was determined by that tandem affinity purification analysis using TPI1 as bait identified CDK2 as binding partner (b) and co-immunoprecipitation analysis revealed that co-expressed TPI1-HA and CDK2-Flag interacted with each other (c). d, Dot blot and western blot was employed to verify reactivity and specificity of anti- S117-P antibody. e, GW8510 prevented CDK2 phosphorylate serine 117 of TPI1 in vitro. The in vitro protein kinase of CDK2 toward S117 of TPI1 was carried out under with and without GW8510 in the reaction mix. S117-P was detected with site-specific anti- S117-P antibody. f, Transient CDK2 inhibition didn’t inhibit cell proliferation. The growth of HeLa cells cultured with and without GW8510 was monitored during 12 hours. n = 3 biologically independent samples (P = 0.1623).
Extended Data Fig. 5 TPI1 S117 phosphorylation promote TPI1 nucleus translocation (Related to Fig. 3).
a, Confirmation of homozygous knock-in of TPI1 and CDK2 mutants. Sequencing results of S117A and S117D of TPI1 mutants knock-in and T160A and T160D of CDK2 mutants knockin were shown. b, TPI1 and CDK2 mutations altered cell cycle progression. FACS results of HeLa cells and homozygous knockin S117A and S117D of TPI1 and T160A and T160D of CDK2 were shown, the gating strategy of flow cytometry was shown in Extended Data Fig. 6b. c, TPI1 S117 mutations affected cell growth. The growth of HeLa, TPI1S117A and TPI1S117D knockin HeLa cells were compared. n = 3 biologically independent samples (P = 0.0089, 0.0412). d, S117D and S117A retained catalytic activities. Specific activities were determined for TPI1 (set as 100% arbitrarily), S117D and S117A that were expressed and purified from HEK293T. n = 4 biologically independent experiments (P =0.5734, 0.0798).
Extended Data Fig. 6 Synchronizing and releasing of CDK2 knockdown cells (related to Fig. 4).
a, CDK2 knockdown HeLa cells were synchronized to G1/S boundary with DTB. The released cells were collected at different time and the percentage of cells at different cell cycle phases were determined by flow cytometry (upper) and quantified (middle and bottom), the gating strategy of flow cytometry was shown in Extended Data Fig. 6b. b, Gating strategy of flow cytometry for normal HeLa cells(upper) and DTB Block Released 0hr HeLa cells(bottom).
Extended Data Fig. 7 Nutrient regulates DHAP and mTORC1 (related to Fig. 5).
a, Nutrient failed to alter histone acetylation in TPI1 S117 mutants knockin cells. The responses of histone acetylation levels to nutrient were detected in HeLa and S117A and S117D knockin HeLa cells. b-c, Nutrient failed to alter nuclear acetate and DHAP levels and histone acetylation levels in TPI1 S117 mutants knockin cells. The nuclear acetate and DHAP levels (b) and histone acetylation levels (c) were detected in HeLa and S117A and S117D knockin HeLa cells that were cultured under absence and presence of glucose and glutamine in the culture media. n = 3 biologically independent samples (P = 0.0015, 0.0009, 0.3713, 0.5402, 0.0735, 0.1577). d-e, Activating mTORC1 abrogated nutrient to regulate nuclear acetate and DHAP levels and histone acetylation levels. The nuclear acetate and DHAP levels (d) and histone acetylation levels (e) were detected in wildtype and NPRL3 knockout HeLa cells cultured in the absence and presence of glucose or glutamine. n = 3 biologically independent samples (P = 5.72e-6, 0.0007, 3.95e-5, 0.0003, 0.5726, 0.5161, 0.1373, 0.5944). f, Rapamycin failed to alter histone acetylation in TPI1 S117 mutants knockin cells. The responses of histone acetylation levels to rapamycin were detected in HeLa and S117A and S117D knockin HeLa cells. g, Raptor interacted with CDK2. Raptor and CDK2 were co-expressed in HeLa cells, co-immunoprecipitation was carried out to detect interaction between them. h. CDK2 T160 mutations affect cell growth. The growth of HeLa, CDK2T160A and CDK2T160D knockin HeLa cells were compared. n = 3 biologically independent samples.
Extended Data Fig. 8 DHAP scavenges nuclear acetate to form 1-acetyl-DHAP (Related to Fig. 6).
a, A non-enzymatic mechanism of the chemical reaction of DHAP with acetate. DHAP can form a dimer, and acetate may attack the DHAP dimer and form 1-acetyl-DHAP. b, Detection of formation of 1-acetyl-DHAP with MS under the negative mode. c, Time-dependent formation of 1-acetyl-DHAP. The amount of 1-acetyl-DHAP formed in reaction mix contained 3 μM acetate and 100 μM DHAP were with or without cell lysate was determined at different times after the initiation of the reaction. The product level of acetate and DHAP mix reacted for 0.25 min was set as 100% arbitrarily. n = 3 biologically independent samples (P = 0.0244, 0.3482, 0.4409, 0.5173). d, The nuclear levels of acetate (left) and DHAP (right) were determined at indicated times after 0.25 mM acetate and 1mM DHAP was supplemented, respectively, to the culture media. n = 3 biologically independent samples. e, Media 1-acetyl-DHAP supplemental increased nuclear 1-acetyl-DHAP levels. n = 3 biologically independent samples (P = 0.0008). f, Acetyltransferases activities were not affected by DHAP, acetate and 1-acetyl-DHAP. The acetyltransferases activities in nuclear extract of HeLa cells were determined by their ability to acetylate a synthetic peptide under absence or presence of these compounds. n = 3 biologically independent samples (P = 0.0848, 0.1021, 0.1297).
Extended Data Fig. 9 mTORC1-CDK2-TPI1 axis regulates histone (Related to Fig. 7).
a, Rapamycin prevented CDK2 overexpression-promoted TPI1 nuclear translocation and histone hyperacetylation. Global histone acetylation and TPI1 levels were detected in lysate, nuclear and histone fractions of HeLa cells and CDK2-overexpressing HeLa cells that were cultured in DMEM media under presence and absence of rapamycin. b, CDK2 inactivation abrogated glucose to induce global histone hyperacetylation. Global histone acetylation levels were detected in lysate and histone fractions of HeLa cells that were cultured in DMEM base media supplemented with different levels of glucose under presence and absence of GW8510. c, Mutations on serine 117 of TPI1 abrogated nutrient to alter TIP1 localization. The cytosolic and nucleus localization of wildtype, S117A and S117D KI HeLa cells that were cultured in media with or without glucose and glutamine were detected with fluorescence staining (left) and the relative nuclear fluorescence intensity to whole cell fluorescence intensity was quantified (right). Scale bar: 10 µm. n = 3 biologically independent samples (P = 0.0002, 0.3528, 0.7319). d, Gene transcriptions that were upregulated by TPI1 overexpression. The enriched pathways which gene transcriptions were upregulated by TPI1 overexpression were listed.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Fig. 7
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 9
Unprocessed western blots and/or gels.
Rights and permissions
About this article
Cite this article
Zhang, JJ., Fan, TT., Mao, YZ. et al. Nuclear dihydroxyacetone phosphate signals nutrient sufficiency and cell cycle phase to global histone acetylation. Nat Metab 3, 859–875 (2021). https://doi.org/10.1038/s42255-021-00405-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00405-8
This article is cited by
-
Newly discovered roles of triosephosphate isomerase including functions within the nucleus
Molecular Medicine (2023)
-
Triosephosphate isomerase 1 may be a risk predictor in laryngeal squamous cell carcinoma: a multi-centered study integrating bulk RNA, single-cell RNA, and protein immunohistochemistry
European Journal of Medical Research (2023)
-
Modulation of cellular processes by histone and non-histone protein acetylation
Nature Reviews Molecular Cell Biology (2022)
-
Reviewing cancer’s biology: an eclectic approach
Journal of the Egyptian National Cancer Institute (2021)
-
TiPpIng the balance in histone acetylation
Nature Metabolism (2021)