Abstract
Obesity is a major risk factor for cardiometabolic diseases. Nevertheless, a substantial proportion of individuals with obesity do not suffer cardiometabolic comorbidities. The mechanisms that uncouple adiposity from its cardiometabolic complications are not fully understood. Here, we identify 62 loci of which the same allele is significantly associated with both higher adiposity and lower cardiometabolic risk. Functional analyses show that the 62 loci are enriched for genes expressed in adipose tissue, and for regulatory variants that influence nearby genes that affect adipocyte differentiation. Genes prioritized in each locus support a key role of fat distribution (FAM13A, IRS1 and PPARG) and adipocyte function (ALDH2, CCDC92, DNAH10, ESR1, FAM13A, MTOR, PIK3R1 and VEGFB). Several additional mechanisms are involved as well, such as insulin–glucose signalling (ADCY5, ARAP1, CREBBP, FAM13A, MTOR, PEPD, RAC1 and SH2B3), energy expenditure and fatty acid oxidation (IGF2BP2), browning of white adipose tissue (CSK, VEGFA, VEGFB and SLC22A3) and inflammation (SH2B3, DAGLB and ADCY9). Some of these genes may represent therapeutic targets to reduce cardiometabolic risk linked to excess adiposity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Summary statistics for the cross-phenotype GWAS analyses are available at http://www.thelooslab.com/#data/ and the GWAS catalogue.
References
Must, A. et al. The disease burden associated with overweight and obesity. JAMA 282, 1523–1529 (1999).
Mokdad, A. H. et al. Prevalence of obesity, diabetes and obesity-related health risk factors, 2001. JAMA 289, 76–79 (2003).
Canoy, D. et al. Body fat distribution and risk of coronary heart disease in men and women in the European Prospective Investigation Into Cancer and Nutrition in Norfolk cohort: a population-based prospective study. Circulation 116, 2933–2943 (2007).
Wildman, R. P. et al. The obese without cardiometabolic risk factor clustering and the normal weight with cardiometabolic risk factor clustering: prevalence and correlates of 2 phenotypes among the US population (NHANES 1999-2004). Arch. Intern. Med. 168, 1617–1624 (2008).
Primeau, V. et al. Characterizing the profile of obese patients who are metabolically healthy. Int. J. Obes. 35, 971–981 (2011).
Bradshaw, P. T., Monda, K. L. & Stevens, J. Metabolic syndrome in healthy obese, overweight, and normal-weight individuals: The Atherosclerosis Risk in Communities Study. Obesity 21, 203–209 (2012).
Ghaben, A. L. & Scherer, P. E. Adipogenesis and metabolic health. Nat. Rev. Mol. Cell Biol. 20, 242–258 (2019).
Smith, G. I., Mittendorfer, B. & Klein, S. Metabolically healthy obesity: facts and fantasies. J. Clin. Invest. 129, 3978–3989 (2019).
Carobbio, S., Pellegrinelli, V. & Vidal-Puig, A. Adipose tissue function and expandability as determinants of lipotoxicity and the metabolic syndrome. Adv. Exp. Med. Biol. 960, 161–196 (2017).
Sun, K., Kusminski, C. M. & Scherer, P. E. Adipose tissue remodeling and obesity. J. Clin. Invest. 121, 2094–2101 (2011).
Kilpelainen, T. O. et al. Genetic variation near IRS1 associates with reduced adiposity and an impaired metabolic profile. Nat. Genet. 43, 753–760 (2011).
Lu, Y. et al. New loci for body fat percentage reveal link between adiposity and cardiometabolic disease risk. Nat. Commun. 7, 10495 (2016).
Yaghootkar, H. et al. Genetic evidence for a link between favorable adiposity and lower risk of type 2 diabetes, hypertension and heart disease. Diabetes 65, 2448–2460 (2016).
Yaghootkar, H. et al. Genetic evidence for a normal-weight ‘metabolically obese’ phenotype linking insulin resistance, hypertension, coronary artery disease and type 2 diabetes. Diabetes 63, 4369–4377 (2014).
Lotta, L. A. et al. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat. Genet. 49, 17–26 (2017).
Ji, Y. et al. Genome-wide and abdominal MRI data provide evidence that a genetically determined favorable adiposity phenotype is characterized by lower ectopic liver fat and lower risk of type 2 diabetes, heart disease and hypertension. Diabetes 68, 207–219 (2019).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Willer, C. J. et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
Manning, A. K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Warren, H. R. et al. Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat. Genet. 49, 403–415 (2017).
Nikpay, M. et al. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Ginguay, A., Cynober, L., Curis, E. & Nicolis, I. Ornithine aminotransferase, an important glutamate-metabolizing enzyme at the crossroads of multiple metabolic pathways. Biology. 6, 18 (2017).
Rauch, A. et al. Osteogenesis depends on commissioning of a network of stem cell transcription factors that act as repressors of adipogenesis. Nat. Genet. 51, 716–727 (2019).
Pers, T. H., Timshel, P. & Hirschhorn, J. N. SNPsnap: a web-based tool for identification and annotation of matched SNPs. Bioinformatics 31, 418–420 (2015).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Tranchevent, L. C. et al. Candidate gene prioritization with Endeavour. Nucleic Acids Res. 44, W117–W121 (2016).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Itan, Y. et al. The human gene connectome as a map of short cuts for morbid allele discovery. Proc. Natl Acad. Sci. USA 110, 5558–5563 (2013).
Sharma, A. M. & Staels, B. Review: peroxisome proliferator-activated receptor gamma and adipose tissue–understanding obesity-related changes in regulation of lipid and glucose metabolism. J. Clin. Endocrinol. Metab. 92, 386–395 (2007).
Melvin, A., Stears, A. & Savage, D. B. Recent developments in lipodystrophy. Curr. Opin. Lipidol. 30, 284–290 (2019).
Kwok, A. et al. Truncation of Pik3r1 causes severe insulin resistance uncoupled from obesity and dyslipidaemia by increased energy expenditure. Mol. Metab. 40, 101020 (2020).
McCurdy, C. E. et al. Attenuated Pik3r1 expression prevents insulin resistance and adipose tissue macrophage accumulation in diet-induced obese mice. Diabetes 61, 2495–2505 (2012).
Thauvin-Robinet, C. et al. PIK3R1 mutations cause syndromic insulin resistance with lipoatrophy. Am. J. Hum. Genet. 93, 141–149 (2013).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism and disease. Cell 168, 960–976 (2017).
Tamemoto, H. et al. Insulin resistance syndrome in mice deficient in insulin receptor substrate-1. Ann. N. Y. Acad. Sci. 827, 85–93 (1997).
Araki, E. et al. Alternative pathway of insulin signalling in mice with targeted disruption of the IRS-1 gene. Nature 372, 186–190 (1994).
Sun, K. et al. Dichotomous effects of VEGF-A on adipose tissue dysfunction. Proc. Natl Acad. Sci. USA 109, 5874–5879 (2012).
Fathzadeh, M. et al. FAM13A affects body fat distribution and adipocyte function. Nat. Commun. 11, 1465 (2020).
Wardhana, D. A. et al. Family with sequence similarity 13, member A modulates adipocyte insulin signaling and preserves systemic metabolic homeostasis. Proc. Natl Acad. Sci. USA 115, 1529–1534 (2018).
Yamauchi, T. et al. Increased insulin sensitivity despite lipodystrophy in Crebbp heterozygous mice. Nat. Genet. 30, 221–226 (2002).
Qi, L. et al. Adipocyte CREB promotes insulin resistance in obesity. Cell Metab. 9, 277–286 (2009).
Chiu, T. T., Jensen, T. E., Sylow, L., Richter, E. A. & Klip, A. Rac1 signalling towards GLUT4/glucose uptake in skeletal muscle. Cell. Signal. 23, 1546–1554 (2011).
Raun, S. H. et al. Rac1 muscle knockout exacerbates the detrimental effect of high-fat diet on insulin-stimulated muscle glucose uptake independently of Akt. J. Physiol. 596, 2283–2299 (2018).
Hodson, D. J. et al. ADCY5 couples glucose to insulin secretion in human islets. Diabetes 63, 3009–3021 (2014).
Knigge, A. et al. ADCY5 gene expression in adipose tissue is related to obesity in men and mice. PLoS ONE 10, e0120742 (2015).
Wagner, R. et al. Glucose-raising genetic variants in MADD and ADCY5 impair conversion of proinsulin to insulin. PLoS ONE 6, e23639 (2011).
Rautureau, Y. et al. ADCY9 (adenylate cyclase type 9) inactivation protects from atherosclerosis only in the absence of CETP (cholesteryl ester transfer protein). Circulation 138, 1677–1692 (2018).
Crujeiras, A. B. et al. Genome-wide DNA methylation pattern in visceral adipose tissue differentiates insulin-resistant from insulin-sensitive obese subjects. Transl. Res. 178, 13–24 (2016).
Chen, Z. et al. Functional screening of candidate causal genes for insulin resistance in human preadipocytes and adipocytes. Circ. Res. 126, 330–346 (2020).
Hu, Y. et al. A phenomics-based approach for the detection and interpretation of shared genetic influences on 29 biochemical indices in southern Chinese men. BMC Genomics 20, 983 (2019).
Kulzer, J. R. et al. A common functional regulatory variant at a type 2 diabetes locus upregulates ARAP1 expression in the pancreatic beta cell. Am. J. Hum. Genet. 94, 186–197 (2014).
Fatima, L. A. et al. Estradiol stimulates adipogenesis and Slc2a4/GLUT4 expression via ESR1-mediated activation of CEBPA. Mol. Cell. Endocrinol. 498, 110447 (2019).
Riant, E. et al. Estrogens protect against high-fat diet-induced insulin resistance and glucose intolerance in mice. Endocrinology 150, 2109–2117 (2009).
Small, K. S. et al. Regulatory variants at KLF14 influence type 2 diabetes risk via a female-specific effect on adipocyte size and body composition. Nat. Genet. 50, 572–580 (2018).
Small, K. S. et al. Identification of an imprinted master trans regulator at the KLF14 locus related to multiple metabolic phenotypes. Nat. Genet. 43, 561–564 (2011).
Song, W. et al. Organic cation transporter 3 (Oct3) is a distinct catecholamines clearance route in adipocytes mediating the beiging of white adipose tissue. PLoS Biol. 17, e2006571 (2019).
Ryu, V. & Buettner, C. Fat cells gobbling up norepinephrine? PLoS Biol. 17, e3000138 (2019).
Saxton, S. N. et al. Role of sympathetic nerves and adipocyte catecholamine uptake in the vasorelaxant function of perivascular adipose tissue. Arterioscler. Thromb. Vasc. Biol. 38, 880–891 (2018).
Mori, T., Suzuki-Yamazaki, N. & Takaki, S. Lnk/Sh2b3 regulates adipose inflammation and glucose tolerance through group 1 ILCs. Cell Rep. 24, 1830–1841 (2018).
Robciuc, M. R. et al. VEGFB/VEGFR1-induced expansion of adipose vasculature counteracts obesity and related metabolic complications. Cell Metab. 23, 712–724 (2016).
Lu, X. et al. Resistance to obesity by repression of VEGF gene expression through induction of brown-like adipocyte differentiation. Endocrinology 153, 3123–3132 (2012).
Hsu, K.-L. New lipid metabolic targets for treatment of inflammation. J. Immunol. 198, 81.28 (2017).
Powell, D. R. et al. Diacylglycerol lipase alpha knockout mice demonstrate metabolic and behavioral phenotypes similar to those of cannabinoid receptor 1 knockout mice. Front. Endocrinol. 6, 86 (2015).
Dai, N. et al. IGF2BP2/IMP2-deficient mice resist obesity through enhanced translation of Ucp1 mRNA and other mRNAs encoding mitochondrial proteins. Cell Metab. 21, 609–621 (2015).
Regue, L., Minichiello, L., Avruch, J. & Dai, N. Liver-specific deletion of IGF2 mRNA binding protein-2/IMP2 reduces hepatic fatty acid oxidation and increases hepatic triglyceride accumulation. J. Biol. Chem. 294, 11944–11951 (2019).
Zhang, L. et al. Global Inactivation of the Pla2g6 gene in mice does not cause dyslipidemia under chow or high-fat diet conditions. J. Cancer Prev. 18, 235–248 (2013).
Carrat, G. R. et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am. J. Hum. Genet 100, 238–256 (2017).
Stolerman, E. S. et al. TCF7L2 variants are associated with increased proinsulin/insulin ratios but not obesity traits in the Framingham Heart Study. Diabetologia 52, 614–620 (2009).
Lee, P. L., Tang, Y., Li, H. & Guertin, D. A. Raptor/mTORC1 loss in adipocytes causes progressive lipodystrophy and fatty liver disease. Mol. Metab. 5, 422–432 (2016).
Stijnen, P., Ramos-Molina, B., O’Rahilly, S. & Creemers, J. W. PCSK1 mutations and human endocrinopathies: from obesity to gastrointestinal disorders. Endocr. Rev. 37, 347–371 (2016).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. Am. J. Hum. Genet. 96, 21–36 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
The Genomes Project, C. A global reference for human genetic variation. Nature 526, 68 (2015).
R core Team. R: a Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2014); http://www.R-project.org/
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Liu, B., Gloudemans, M. & Montgomery, S. LocusCompare: a Tool to Visualize Pairs of Association (2018); http://locuscompare.com/
Choi, S. W. & O’Reilly, P. F. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience 8, giz082 (2019).
Simonsen, J. L. et al. Telomerase expression extends the proliferative life-span and maintains the osteogenic potential of human bone marrow stromal cells. Nat. Biotechnol. 20, 592–596 (2002).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Li, Q. H., Brown, J. B., Huang, H. Y. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Carithers, L. J. et al. A novel approach to high-quality postmortem tissue procurement: The GTEx Project. Biopreserv. Biobank 13, 311–319 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Zhu, L. J. et al. ChIPpeakAnno: a Bioconductor package to annotate ChIP–seq and ChIP–chip data. BMC Bioinformatics 11, 237 (2010).
Acknowledgements
R.J.F.L. is supported by the National Institutes of health (NIH; R01DK11011, R01DK107786 and R01DK124097). T.O.K. is supported by the Independent Research Fund Denmark (6110-00183) and the Novo Nordisk Foundation (NNF17OC0026848, NNF18CC0034900 and NNF20OC0063707). M.d.H. is a fellow of the Swedish Heart-Lung Foundation (20170872 and 20200781) and a Kjell and Märta Beijer Foundation researcher. M.d.H. is further supported by the Swedish Heart-Lung Foundation (20140543, 20170678, 20180706 and 20200602) and the Swedish Research Council (2015-03657 and 2019-01417). A.V.-P. is supported by NIH R01DK107786, an ERC Senior Investigator award (669879) and an MRC MDU Programme grant (PO 4050281695). Yuval Itan is supported by the NIH (R01DK123530). The work of A.R. and S.M. was supported by grants from the Independent Research Fund Denmark (Sapere Aude Advanced grant no. 12-125524), the Danish National Research Foundation (grant no. 141) to the Center for Functional Genomics and Tissue Plasticity and the Novo Nordisk Foundation. A.R. was further supported by a postdoctoral grant from the Danish Diabetes Academy supported by the Novo Nordisk Foundation. N.C. is supported by a grant from the Canadian Institutes of Health Research (fellowship MFE-158192).
Author information
Authors and Affiliations
Contributions
R.J.F.L. and T.O.K. conceived the project, designed the experiments, supervised analyses and edited the manuscript. L.O.H. performed the final computational analyses for gene discovery, follow-up analyses and wrote the first draft of the manuscript and edited subsequent versions. U.M.S. and N.Y. performed the initial gene-discovery analyses. A.R. and S.M. led the human adipocyte analyses. C.S.B. and Y.I. performed IPA enrichment analysis and HGC biological distance calculations. E.M. performed the ToppGene and Endeavour screens, integrated results across all gene prioritization approaches and drafted the section on gene prioritization. M.d.H. supervised the integration of gene prioritization results, liaising with E.M. M.P. performed GRS analyses in the UKBB population. N.C. and Z.W. performed validation analyses in the UKBB population. S.C., A.V.-P., S.M., M.d.H., T.O.K. and R.J.F.L. examined the genes prioritized for their biological contribution. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Jose Florez, Xueling Sim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Pooja Jha; Isabella Samuelson.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
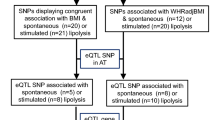
Extended Data Fig. 1
Overview of the different stages, approaches and results.
Extended Data Fig. 2 Tissue and cell type enrichment analyses for genes in cluster 1.
Tissues [A], cells [B] and physiological systems [C] significantly enriched for genes in 15 loci of cluster 1. Built-in iterative statistical computation using scoring, bias adjustment, and FDR estimation, based on the membership z-score. The p-value, called ‘nominal p-value’ is not corrected for multiple testing, whereas the FDR (shown in Tables), is corrected for multiple testing.
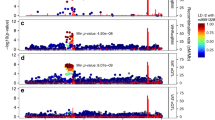
Extended Data Fig. 3 DNase-seq, H3K27ac ChIP-seq and RNA-seq data of hMSC-TERT4 cells during adipocyte differentiation.
(a) DNase-seq, H3K27ac ChIP-seq and RNA-seq data of hMSC-TERT4 cells during adipocyte differentiation. (a) DNase-seq and H3K27ac ChIP-seq read density (combined tracks from two independent experiments) in the NT5C2 locus and bar plot with expression level of NT5C2 from RNA-seq (n = 3 biologically independent samples). Bar plot with average and error bars representing standard deviation. DEseq2 derived Benjamin-Hochberg adjusted P-values below 0.01 when comparing to MSC are indicated. Lead SNP rs10883832 is indicated by red arrow. Black lines represent eQTLs that are in high LD with rs10883832. Black arrow shows rs11191548. DNase I hypersensitive sites are shaded. (b) PPARG locus as in (a). Lead SNP rs2881654 is indicated by red arrow. Black lines represent SNPs that are in high LD with rs2881654. (c) PIK3R1 locus as in (a). Lead SNP rs4976033 is indicated by red arrow.
Extended Data Fig. 4 Gene prioritization tools and genes prioritized.
61 unique candidate genes were prioritized by at least one of eight criteria used to integrate results from individual bioinformatics approaches.
Extended Data Fig. 5 A Sunburst plot showing the criteria by which each candidate gene was prioritized.
The root (inner circle) displays the chromosome on which the prioritized genes (middle circle) are located; the leaves (outer circle) display the criteria (color-coded) fulfilled by the gene. The plot shows that half of the candidate genes have been prioritized by multiple criteria. E-TG: Endeavour and ToppGene; IPA: ingenuity pathway analysis; HGC: human gene connectome. Criteria: 1 = DEPICT P ≤ 0.1; 2 = (eQTL or chromatin-chromatin interaction in relevant tissue) AND top 10% E-TG; 3 = eQTL in relevant tissue AND IPA ≥ 5 pathways, functions, networks; 4 = top 2.5% E-TG; 5 = top 10% E-TG AND IPA ≥ 5 pathways, functions, networks; 6 = Enhancer enrichment analysis; 7 = IPA ≥ 5 pathways, functions, networks; 8 HGC = average biological distance to PPARG and IRS1 < 10.
Extended Data Fig. 6
Number of genes prioritized by one or more of the eight criteria.
Supplementary information
Rights and permissions
About this article
Cite this article
Huang, L.O., Rauch, A., Mazzaferro, E. et al. Genome-wide discovery of genetic loci that uncouple excess adiposity from its comorbidities. Nat Metab 3, 228–243 (2021). https://doi.org/10.1038/s42255-021-00346-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00346-2
This article is cited by
-
CCDC92 promotes podocyte injury by regulating PA28α/ABCA1/cholesterol efflux axis in type 2 diabetic mice
Acta Pharmacologica Sinica (2024)
-
FTO is a major genetic link between breast cancer, obesity, and diabetes
Breast Cancer Research and Treatment (2024)
-
Diverse WGBS profiles of longissimus dorsi muscle in Hainan black goats and hybrid goats
BMC Genomic Data (2023)
-
A phenome-wide comparative analysis of genetic discordance between obesity and type 2 diabetes
Nature Metabolism (2023)
-
Polygenic risk scores analyses of psychiatric and metabolic traits with antipsychotic-induced weight gain in schizophrenia: an exploratory study
The Pharmacogenomics Journal (2023)