Abstract
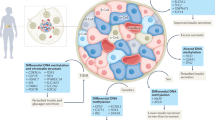
Metabolic memory, the persistent benefits of early glycaemic control on preventing and/or delaying the development of diabetic complications, has been observed in the Diabetes Control and Complications Trial (DCCT) and in the Epidemiology of Diabetes Interventions and Complications (EDIC) follow-up study, but the underlying mechanisms remain unclear. Here, we show the involvement of epigenetic DNA methylation (DNAme) in metabolic memory by examining its associations with preceding glycaemic history, and with subsequent development of complications over an 18-yr period in the blood DNA of 499 randomly selected DCCT participants with type 1 diabetes who are also followed up in EDIC. We demonstrate the associations between DNAme near the closeout of DCCT and mean HbA1c during DCCT (mean-DCCT HbA1c) at 186 cytosine–guanine dinucleotides (CpGs) (FDR < 15%, including 43 at FDR < 5%), many of which were located in genes related to complications. Exploration studies into biological function reveal that these CpGs are enriched in binding sites for the C/EBP transcription factor, as well as enhancer/transcription regions in blood cells and haematopoietic stem cells, and open chromatin states in myeloid cells. Mediation analyses show that, remarkably, several CpGs in combination explain 68–97% of the association of mean-DCCT HbA1c with the risk of complications during EDIC. In summary, DNAme at key CpGs appears to mediate the association between hyperglycaemia and complications in metabolic memory, through modifying enhancer activity at myeloid and other cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The DNA methylation dataset of 499 DCCT participants from this study has been deposited in the Database of Genotype and Phenotype (dbGaP) under the accession number phs002024.v1.p1. Other data that support the findings from this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Nathan, D. M. et al. The effect of intensive treatment of diabetes on the development and progression of long-term complications in insulin-dependent diabetes mellitus. N. Engl. J. Med. 329, 977–986 (1993).
Nathan, D. M. The Diabetes Control and Complications Trial/epidemiology of diabetes interventions and complications study at 30 years: overview. Diabetes Care 37, 9–16 (2014).
Lachin, J. M. et al. Effect of intensive diabetes therapy on the progression of diabetic retinopathy in patients with type 1 diabetes: 18 years of follow-up in the DCCT/EDIC. Diabetes 64, 631–642 (2015).
de Boer, I. H. et al. Intensive diabetes therapy and glomerular filtration rate in type 1 diabetes. N. Engl. J. Med. 365, 2366–2376 (2011).
DCCT/EDIC research group. Effect of intensive diabetes treatment on albuminuria in type 1 diabetes: long-term follow-up of the diabetes control and complications trial and epidemiology of diabetes interventions and complications study. Lancet Diabetes Endocrinol. 2, 793–800 (2014).
Kato, M. & Natarajan, R. Epigenetics and epigenomics in diabetic kidney disease and metabolic memory. Nat. Rev. Nephrol. 15, 327–345 (2019).
Reddy, M. A., Zhang, E. & Natarajan, R. Epigenetic mechanisms in diabetic complications and metabolic memory. Diabetologia 58, 443–455 (2015).
Cooper, M. E. & El-Osta, A. Epigenetics: mechanisms and implications for diabetic complications. Circ. Res. 107, 1403–1413 (2010).
Bird, A. Perceptions of epigenetics. Nature 447, 396–398 (2007).
Russo, V. E. A., Martienssen, R. A. & Riggs, A. D. Epigenetic Mechanisms of Gene Regulation (Cold Spring Harbor Laboratory Press, 1996).
Jirtle, R. L. & Skinner, M. K. Environmental epigenomics and disease susceptibility. Nat. Rev. Genet. 8, 253–262 (2007).
Simmons, R. Epigenetics and maternal nutrition: nature v. nurture. Proc. Nutr. Soc. 70, 73–81 (2011).
Susztak, K. Understanding the epigenetic syntax for the genetic alphabet in the kidney. J. Am. Soc. Nephrol. 25, 10–17 (2014).
Rosen, E. D. et al. Epigenetics and epigenomics: implications for diabetes and obesity. Diabetes. 67, 1923–1931 (2018).
Sandholm, N. et al. The genetic landscape of renal complications in type 1 diabetes. J. Am. Soc. Nephrol. 28, 557–574 (2017).
Pollack, S. et al. Multiethnic genome-wide association study of diabetic retinopathy using liability threshold modeling of duration of diabetes and glycemic control. Diabetes. 68, 441–456 (2019).
Hosseini, S. M. et al. The association of previously reported polymorphisms for microvascular complications in a meta-analysis of diabetic retinopathy. Hum Genet. 134, 247–257 (2015).
Ko, Y. A. et al. Cytosine methylation changes in enhancer regions of core pro-fibrotic genes characterize kidney fibrosis development. Genome Biol. 14, R108 (2013).
Wing, M. R. et al. DNA methylation profile associated with rapid decline in kidney function: findings from the CRIC study. Nephrol. Dial. Transplant. 29, 864–872 (2014).
Chu, A. Y. et al. Epigenome-wide association studies identify DNA methylation associated with kidney function. Nat. Commun. 8, 1286 (2017).
Qiu, C. et al. Cytosine methylation predicts renal function decline in American Indians. Kidney Int. 93, 1417–1431 (2018).
Miao, F. et al. Evaluating the role of epigenetic histone modifications in the metabolic memory of type 1 diabetes. Diabetes 63, 1748–1762 (2014).
Chen, G. et al. Aberrant DNA methylation of mTOR pathway genes promotes inflammatory activation of immune cells in diabetic kidney disease. Kidney Int. https://doi.org/10.1016/j.kint.2019.02.020 (2019).
Gluck, C. et al. Kidney cytosine methylation changes improve renal function decline estimation in patients with diabetic kidney disease. Nat. Commun. 10, 2461 (2019).
Salem, R. M. et al. Genome-wide association study of diabetic kidney disease highlights biology involved in glomerular basement membrane collagen. J. Am. Soc. Nephrol. 30, 2000–2016 (2019).
Chen, Z. et al. Epigenomic profiling reveals an association between persistence of DNA methylation and metabolic memory in the DCCT/EDIC type 1 diabetes cohort. Proc. Natl Acad. Sci. USA 113, E3002–E3011 (2016).
Shalev, A. Minireview: thioredoxin-interacting protein: regulation and function in the pancreatic beta-cell. Mol. Endocrinol. 28, 1211–1220 (2014).
De Marinis, Y. et al. Epigenetic regulation of the thioredoxin-interacting protein (TXNIP) gene by hyperglycemia in kidney. Kidney Int. 89, 342–353 (2016).
Kumar, A. & Mittal, R. Mapping Txnip: key connexions in progression of diabetic nephropathy. Pharmacol. Rep. 70, 614–622 (2018).
Singh, L. P. Thioredoxin interacting protein (TXNIP) and pathogenesis of diabetic retinopathy. J. Clin. Exp. Ophthalmol. https://doi.org/10.4172/2155-9570.1000287 (2013).
Rich, S. S. et al. A genome-wide association scan for acute insulin response to glucose in Hispanic-Americans: the insulin resistance atherosclerosis family study (IRAS FS). Diabetologia 52, 1326–1333 (2009).
Neve, B. et al. Role of transcription factor KLF11 and its diabetes-associated gene variants in pancreatic beta cell function. Proc. Natl Acad. Sci. USA 102, 4807–4812 (2005).
Fernandez-Zapico, M. E. et al. MODY7 gene, KLF11, is a novel p300-dependent regulator of Pdx-1 (MODY4) transcription in pancreatic islet beta cells. J. Biol. Chem. 284, 36482–36490 (2009).
Park, I. Y. et al. Dual chromatin and cytoskeletal remodeling by SETD2. Cell 166, 950–962 (2016).
Kulkarni, H. et al. Novel epigenetic determinants of type 2 diabetes in Mexican-American families. Hum. Mol. Genet. 24, 5330–5344 (2015).
Yamanouchi, M. et al. Improved clinical trial enrollment criterion to identify patients with diabetes at risk of end-stage renal disease. Kidney Int. 92, 258–266 (2017).
Toperoff, G. et al. Genome-wide survey reveals predisposing diabetes type 2-related DNA methylation variations in human peripheral blood. Hum. Mol. Genet. 21, 371–383 (2012).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Avellino, R. & Delwel, R. Expression and regulation of C/EBPα in normal myelopoiesis and in malignant transformation. Blood 129, 2083–2091 (2017).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 e1319 (2016).
Rickels, R. & Shilatifard, A. Enhancer logic and mechanics in development and disease. Trends Cell Biol. 28, 608–630 (2018).
Reynolds, L. M. et al. Age-related variations in the methylome associated with gene expression in human monocytes and T cells. Nat. Commun. 5, 5366 (2014).
Roshandel, D. et al. Meta-genome-wide association studies identify a locus on chromosome 1 and multiple variants in the MHC region for serum C-peptide in type 1 diabetes. Diabetologia 61, 1098–1111 (2018).
Paterson, A. D. et al. A genome-wide association study identifies a novel major locus for glycemic control in type 1 diabetes, as measured by both A1C and glucose. Diabetes 59, 539–549 (2010).
Hainsworth, D. P. et al. Risk factors for retinopathy in type 1 diabetes: The DCCT/EDIC study. Diabetes Care 42, 875–882 (2019).
Perkins, B. A. et al. Risk factors for kidney disease in type 1 diabetes. Diabetes Care 42, 883–890 (2019).
Kriebel, J. et al. Association between DNA methylation in whole blood and measures of glucose metabolism: KORA F4 study. PLoS ONE 11, e0152314 (2016).
Ronn, T. et al. Impact of age, BMI and HbA1c levels on the genome-wide DNA methylation and mRNA expression patterns in human adipose tissue and identification of epigenetic biomarkers in blood. Hum. Mol. Genet. 24, 3792–3813 (2015).
Hidalgo, B. et al. Epigenome-wide association study of fasting measures of glucose, insulin, and HOMA-IR in the genetics of lipid lowering drugs and diet network study. Diabetes 63, 801–807 (2014).
Walaszczyk, E. et al. DNA methylation markers associated with type 2 diabetes, fasting glucose and HbA1c levels: a systematic review and replication in a case-control sample of the Lifelines study. Diabetologia 61, 354–368 (2018).
Chambers, J. C. et al. Epigenome-wide association of DNA methylation markers in peripheral blood from Indian Asians and Europeans with incident type 2 diabetes: a nested case-control study. Lancet Diabetes Endocrinol. 3, 526–534 (2015).
Soriano-Tarraga, C. et al. Epigenome-wide association study identifies TXNIP gene associated with type 2 diabetes mellitus and sustained hyperglycemia. Hum. Mol. Genet. 25, 609–619 (2016).
Cardona, A. et al. Epigenome-wide association study of incident type 2 diabetes in a british population: EPIC-Norfolk study. Diabetes 68, 2315–2326 (2019).
Ye, J. et al. Identification of loci where DNA methylation potentially mediates genetic risk of type 1 diabetes. J. Autoimmun. 93, 66–75 (2018).
Vigorelli, V. et al. Abnormal DNA methylation induced by hyperglycemia reduces CXCR 4 gene expression in CD 34+ stem cells. J. Am. Heart Assoc. 8, e010012 (2019).
Tsukada, J., Yoshida, Y., Kominato, Y. & Auron, P. E. The CCAAT/enhancer (C/EBP) family of basic-leucine zipper (bZIP) transcription factors is a multifaceted highly-regulated system for gene regulation. Cytokine 54, 6–19 (2011).
Nagareddy, P. R. et al. Hyperglycemia promotes myelopoiesis and impairs the resolution of atherosclerosis. Cell Metab. 17, 695–708 (2013).
Mitroulis, I. et al. Modulation of myelopoiesis progenitors is an integral component of trained immunity. Cell. 172, 147–161 e112 (2018).
Woroniecka, K. I. et al. Transcriptome analysis of human diabetic kidney disease. Diabetes 60, 2354–2369 (2011).
Kojima, H., Kim, J. & Chan, L. Emerging roles of hematopoietic cells in the pathobiology of diabetic complications. Trends Endocrinol. Metab. 25, 178–187 (2014).
The Diabetes Control and Complications Trial (DCCT). Design and methodologic considerations for the feasibility phase. The DCCT Research Group. Diabetes 35, 530–545 (1986).
Fortin, J. P., Triche, T. J. Jr. & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 13, 86 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Niewczas, M. A. et al. A signature of circulating inflammatory proteins and development of end-stage renal disease in diabetes. Nat. Med. 25, 805–813 (2019).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Acknowledgements
A complete list of participants in the DCCT DCCT/EDIC Study Group is presented at the end of the Supplementary Information. We are deeply grateful to A. Krolewski (Joslin Diabetes Center, Harvard Medical School, Boston, MA) for generously providing DNA samples and related clinical information from his Joslin Kidney Study cohort of people with T1D. We are also grateful to the DCCT/EDIC Central Biochemistry Laboratory at the University of Minnesota for assistance in providing the archived DNA samples. This study was supported by grants from the National Institutes of Health (NIH): DP3 DK106917-01 and R01 DK065073 (to R.N.), and the Wanek family project at the City of Hope (to R.N., A.D.R. and J.D.T.). Research reported in this publication included work performed in the following Campus Cores: Integrative Genomics, and DNA/RNA synthesis (supported by the National Cancer Institute of the NIH under award number P30CA33572), and the Genomics Core at the University of Southern California (D. Weisenberger, Director). The DCCT/EDIC has been supported by cooperative agreement grants (1982-1993, 2012-2017, 2017-2022) and contracts (1982-2012) with the Division of Diabetes Endocrinology and Metabolic Diseases of the National Institute of Diabetes and Digestive and Kidney Disease (current grant numbers U01 DK094176 and U01 DK094157), and through support by the National Eye Institute, the National Institute of Neurologic Disorders and Stroke, the General Clinical Research Centers Program (1993-2007) and Clinical Translational Science Center Program (2006-present). Industry contributors have had no role in the DCCT/EDIC study but have provided free or discounted supplies or equipment to support participants’ adherence to the study: Abbott Diabetes Care, Animas, Bayer Diabetes Care, Becton Dickinson, Eli Lilly, Extend Nutrition, Insulet Corporation, Lifescan, Medtronic Diabetes, Nipro Home Diagnostics, Nova Diabetes Care, Omron, Perrigo Diabetes Care,Roche Diabetes Care and Sanofi-Aventis. The trial is registered at clinicaltrials.gov, NCT00360815 and NCT00360893.
Author information
Authors and Affiliations
Consortia
Contributions
Z.C. and R.N. designed the study and wrote the manuscript; Z.C., F.M., A.D.P., J.M.L., B.H.B. and R.N. designed the project, analysed data and edited the manuscript; Z.C., F.M., L.Z., X.W. and D.R. performed experiments and research and analysed data; X.A.L., J.S.K., A.D.R. and J.D.T. assisted in the analyses, provided valuable advice and reviewed and edited the manuscript; M.C. provided data for validations and reviewed and edited the manuscript; the DCCT/EDIC Research Study Group helped in the project design and reviewed the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sensitivity analyses of additional clinical variables on the association significance between mean-DCCT HbA1c and DNAme at the HbA1c-associated CpGs.
For each HbA1c-associated CpG, one additional clinical variable (indicated on top of each plot) was added as covariate in the multiple linear regression model used for the identification of HbA1c-associated CpGs using all the samples (two-sided tests based on t-statistic, n=499). The resulting association significance in –log10P (y-axis) was compared with that obtained from the model without the specific variable (x-axis) and shown as one dot in the scatter plots. The blue line represents all the dots with same significance values. Abbreviations: SBP- systolic blood pressure; DBP-diastolic blood pressure; BMI-body mass index; CHL-cholesterol; TRG-triglyceride; LDL-low density lipoprotein; HDL-high density lipoprotein; PDR- proliferative diabetic retinopathy; SNPDR-severe nonproliferative diabetic retinopathy; CSME- clinically significant macular edema; AER30-AER > 30 mg/24 h; CARV-cardiovascular disease.
Extended Data Fig. 2 Sensitivity analyses of additional clinical variables on the percentage of association coefficients changes between mean-DCCT HbA1c and DNAme at the HbA1c-associated CpGs.
For each HbA1c-associated CpG, one additional clinical variable (indicated on top of each plot) was added as covariate in the multiple linear regression model with two-sided tests based on t-statistic used for identification of HbA1c-associated CpGs using all the samples (n=499). For each clinical variable, the percentage change of the resulting association coefficients versus the coefficients estimated using the original model without the specific variable was calculated. For each additional variable, the distribution of the calculated changes was plotted using density plots. Abbreviations: COEF-coefficient; SBP- systolic blood pressure; DBP-diastolic blood pressure; BMI-body mass index; CHL-cholesterol; TRG-triglyceride; LDL-low density lipoprotein; HDL-high density lipoprotein; PDR- proliferative diabetic retinopathy; SNPDR-severe nonproliferative diabetic retinopathy; CSME- clinically significant macular edema; AER30-AER > 30 mg/24 h; CARV-cardiovascular disease.
Extended Data Fig. 3 Validation of HbA1c-associated CpGs identified in DCCT499 in a meta-analysis of DCCT499 and Joslin195.
Fixed meta-analysis was performed using Bacon R package based on coefficients and SER of CpGs obtained in both datasets (n=499 in DCCT499 and n=195 in Joslin195). Scatter plot is shown to compare the association estimates generated from meta-analysis and original models across 185 HbA1c-associated CpGs, after excluding one HbA1c-associated CpG not covered reliably in Joslin195 dataset due to detection P > 0.05 in at least one sample. Each dot represents one CpG with colour indicating the significance levels obtained from meta-analysis. Red dots represent Bonferroni-adjusted P < 0.05 (cutoff P = 0.05/185 = 2.7 × 10−4), orange dots represent nominal P < 0.05, black dots represent remaining CpGs. Blue line represents all the dots with same association estimates obtained from meta-analysis and original model.
Extended Data Fig. 4 Genomic locations and IPA analyses of HbA1c-associated genes.
a, Genomic locations of HbA1c-associated CpGs and non-associated CpGs relative to Refseq genes depicted using pie charts. TSS, transcription start site; TSS1500, distal promoter region from 1,500 bp upstream of TSS to 200 bp upstream of TSS; TSS200, proximal promoter from 200 bp upstream of TSS to TSS; 5’UTR, 5’ untranslated region; 3’UTR, 3’ untranslated region; intergenic, genomic regions excluding TSS1500, TSS200 and gene body. b, Enriched canonical pathways (Benjamini-Hochberg adjusted P < 0.05) identified among annotated genes of HbA1c-associated CpGs. Right-tailed Fisher’s exact test was used in IPA analysis. Total of 143 unique annotated genes were used as input for IPA, and 141 genes after excluding 2 that did not match in the IPA database were finally included in the analysis. Pathway names are listed on the left with corresponding significance level in –log10 form presented in the middle as bar plot. The pathways labeled in red font are known to be associated with diabetes and its complications. HbA1c-associated genes identified in each pathway are shown on the right. Each row represents one pathway and each column represents one HbA1c-associated gene found in the enriched pathway. Red indicates the corresponding specific gene (with name shown on the top of the column) that is involved in the pathway specified on the left of the panel. The majority of them contain several NFAT pathway genes, including TGFBR2, GNAI2, CACNB1, PLCB4, MEF2D, TGFB2, ADCY7, PRKD1 and CACNA1A. c, Top network related to cellular growth, proliferation and embryonic development. The 24 HbA1c-associated genes labeled in red font are those identified to depict interactions with several proteins related to diabetes, its complications and insulin sensitivity including nuclear factor kappaB (NF-kB), protein kinaseB (PKB), phosphoinositide 3-kinase (PI3K), p38 mitogen-activated protein kinase (P38MAPK) and protein phosphatase2 catalytic (PPP2C) subunits. HbA1c-assoc CpGs: HbA1c-associated CpGs.
Extended Data Fig. 5 Enrichment of 15 chromatin states at 186 HbA1c-associated CpGs compared to all the non-HbA1c-associated CpGs.
Fifteen genome-wide chromatin states were defined in 5 major blood cell types and hematopoietic stem cell (HSC) by the NIH roadmap Epigenomics Program (https://egg2.wustl.edu/roadmap/web_portal/chr_state_learning.html#core_15state for details). For each cell type, the percentage of CpGs located in the genomic regions depicting each chromatin state among HbA1c-associated CpGs (black bars) versus all CpGs covered by the EPIC array (named as ‘all CpGs’, white bars) are shown side by side using bar plots. The enrichment significances of HbA1c-associated CpGs versus the other CpGs were tested by right-tailed Fisher’s exact tests on 186 HbA1c-associated CpGs vs. all the 815,246 non HbA1c-associated CpGs reliably covered by the EPIC array in each specific chromatin state. ****P < 5 × 10−8; ***P < 5 × 10−5; **P < 5 × 10−3 and *P < 5 × 10−2. States with statistically significant enrichment in at least one cell type are highlighted with colored background/shading: green for transcription-related (TxFlnk, Tx and TxWk) and yellow for enhancer-related (EnhG or Enh). The other states are shown in alternating grey and white shades for better visualization. Abbreviations: HSC-hematopoietic stem cell; TSS-transcription start site; TssA-Active TSS; TssAFlnk-Flanking Active TSS; TxFlnk-Transcribed state at gene 5’ and 3’; Tx-Strong transcription; TxWk-Weak transcription; EnhG-Genic enhancers; Enh-Enhancers; ZNF/Rpts- ZNF genes & repeats; Het- Heterochromatin; TssBiv- Bivalent/Poised TSS; BivFlnk- Flanking Bivalent TSS/Enh; EnhBiv- Bivalent Enhancer; ReprPC- Repressed PolyComb; ReprPCWk- Weak Repressed PolyComb; Quies-Quiescent/Low. HbA1c-assoc CpGs: HbA1c-associated CpGs.
Extended Data Fig. 6 Enrichment of chromatin (epigenetic) states at HbA1c-associated CpG sites in blood cells.
a, Enrichment of chromatin states in each cell-type at the 186 HbA1c-associated CpGs. For each cell-type, the percentage of CpGs located in genomic regions enriched with 18 chromatin states (labeled under the plots) at HbA1c-associated CpGs (dark bars) versus all CpGs covered by the EPIC array (white bars) in 4 major blood cell-types are shown side-by-side in the bar plots. Right-tailed Fisher’s exact tests were conducted to identify the significance of enrichment of each state in 186 HbA1c-associated CpGs relative to all the 815,246 non HbA1c-associated CpGs reliably covered by EPIC array. States with significant enrichment in at least one cell-type are highlighted with colored background/shading: green for transcription-related (Tx and TxWk) and yellow for enhancer-related states (EnhG1, EnhG2, EnhA1, EnhA2 or EnhWk). P values listed on top of each significantly enriched state.****P < 5 × 10−8; ***P < 5 × 10−5; **P < 5 × 10−3 and *P < 5 × 10−2. The other states are shown in alternating grey and white shades for better visualization. b, Enrichment of enhancer- or transcription-related states across 4 different cell-types in HbA1c-associated CpGs versus all CpGs. The states for each plot are shown in the heading. P-values were determined using same tests and sample sizes as in panel a. c, Comparison of heatmaps of 15 chromatin states in 6 blood cell-types (top) with heatmaps of 18 states at 4 blood cell-types (bottom) at HbA1c-associated CpGs. These states are defined by the NIH Roadmap Epigenomics Program. Unsupervised hierarchical analysis was performed on data from 15 states and visualized in the top panel using colours depicted in the boxed legend for 15 states (bottom left). 18-chromatin states at HbA1c-associated CpGs, presented in the same order as 15 states, are shown in the lower panel with corresponding colour legends in the bottom right box. Each row represents one cell-type and each column, one CpG. HbA1c-assoc CpGs: HbA1c-associated CpGs.
Extended Data Fig. 7 Ribbon plots for the association between DNAme at candidate loci (HbA1c-associated CpGs) and the expression of genes located within 500 kb (FDR < 0.05) that were observed in both monocytes and CD4+ cells.
The associations of DNAme at HbA1c-associated CpGs with the expression of gene (s) in monocytes or CD4+ T cells were analyzed on each pair of HbA1c-associated CpGs and corresponding nearby genes within 500 kb. Multiple linear regression models using two-sided tests based on t-statistic (n=1202 for monocytes and n=214 for CD4+ T cells) were applied to published datasets containing both DNAme and gene expression profiles from the same samples to identify HbA1c-associated CpGs whose DNAme is associated with the expression of nearby gene (s) (expression-associated genes) with FDR < 0.05 in monocytes and CD4+ cells. The associations for each CpG in both monocytes (left) and CD4+ cells (right) are shown side-by-side within each panel. The order of CpGs is based on the significance level of the CpG with its most significantly expression-associated gene in both cell types. The height of the ribbon represents the significance level in –log10(P). Blue indicates negative association while red indicates positive association. Grey represents associations with nominal P < 0.05 but FDR >0.05. Most of the 9 common CpGs are in enhancers, suggesting a more ubiquitous regulatory role for DNAme at enhancers across different blood cell types.
Extended Data Fig. 8 Ribbon plots for the association between DNAme at candidate loci (HbA1c-associated CpGs) and expression of genes located within 500 bp (FDR < 0.05) that were observed only in monocytes (not in CD4+ cells).
The associations of DNAme at HbA1c-associated CpGs with the expression of gene (s) in monocytes were analyzed on each pair of HbA1c-associated CpGs and corresponding nearby genes within 500 kb. Multiple linear regression models using two-sided tests based on t-statistic were applied to published datasets containing both DNAme and gene expression profiles from same monocyte samples (n=1202) to identify HbA1c-associated CpGs whose DNAme is associated with the expression of nearby gene (s) (expression-associated genes) with FDR < 0.05. The order of CpGs is based on the significance level of the CpG with its most significantly expression-associated gene. The height of the ribbon represents the significance level in –log10(P). Blue indicates negative association while red indicates positive association. Grey represents the associations with nominal P < 0.05 but FDR >0.05.
Extended Data Fig. 9 Ribbon plots for the association between DNAme at candidate loci (HbA1c-associated CpGs) and expression of genes located within 500 bp (FDR < 0.05) that were observed only in CD4+ cells (not in monocytes).
The associations of DNAme at HbA1c-associated CpGs with the expression of gene(s) in CD4+ T cells were analyzed on each pair of HbA1c-associated CpGs and corresponding nearby genes within 500 kb. Multiple linear regression models using two-sided tests based on t-statistic were applied to published datasets containing both DNAme and gene expression profiles from same CD4+ T cell samples (n=214) to identify HbA1c-associated CpGs whose DNAme is associated with the expression of nearby gene(s) (expression-associated genes) with FDR < 0.05. The order of the CpGs is based on significance level of the CpG with its most significantly associated gene (expression data). The height of the ribbon represents the significance level in –log10(P). Blue indicates negative association while red indicates positive association. Grey represents the associations with nominal P < 0.05 but FDR >0.05.
Extended Data Fig. 10 Pipeline depicting the step-wise sequence used for selecting CpGs for mediation analyses of DNAme in between mean-DCCT HbA1c and complication development during EDIC.
At each selection step, the number of CpGs (n) is shown at the specified cut-off criteria. Arrows with same color reflect the same analytical step using the same model for each complication. The 4 steps as described in the Methods are specified on the right side of the figure. “n” represents number of CpGs identified at each step. In step 1, Model 1 was applied to each complication to analyze the association of mean-DCCT HbA1c with risk of complication development during EDIC (up to EDIC year 18). In step 2, Model 2 was applied to each complication to identify the HbA1c-associated CpGs whose DNAme is associated with risk of complication development. In step 3, Model 3 was applied to each complication using similar model as Model 1 and Model 2, in which both DNAme and mean-DCCT HbA1c were now included. To identify the best combinations of CpGs to explain the association between HbA1c and complication development in step 4, only the top 10 CpGs (based on the association significance between risk of complication and DNAme identified using model 2 in step2) were considered in order to capture the major mediation effect, while maintaining reasonable computational complexity (which increases dramatically as the number of CpGs included increases using Brute-force approach). HbA1c-assoc CpGs: HbA1c-associated CpGs.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 and Tables 1–6, 10–15, 17–19, 21–23, 25–28 and 31
Source data
Source Data Fig. 1
Source Data for Fig. 1
Source Data Fig. 3
Source Data for Fig. 3
Source Data Fig. 4
Source Data for Fig. 4
Source Data Fig. 5
Source Data for 5
Source Data Fig. 6
Source Data for Fig. 6
Source Data Extended Data Fig. 3
Source Data for Extended Data 3
Source Data Extended Data Fig. 4
Source Data for Extended Data 4
Source Data Extended Data Fig. 5
Source Data for Extended Data 5
Source Data Extended Data Fig. 6
Source Data for Extended Data 6
Source Data Extended Data Fig. 7
Source Data for Extended Data 7
Source Data Extended Data Fig. 8
Source Data for Extended Data 8
Source Data Extended Data Fig. 9
Source Data for Extended Data 9
Rights and permissions
About this article
Cite this article
Chen, Z., Miao, F., Braffett, B.H. et al. DNA methylation mediates development of HbA1c-associated complications in type 1 diabetes. Nat Metab 2, 744–762 (2020). https://doi.org/10.1038/s42255-020-0231-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-0231-8
This article is cited by
-
Metabolic memory: mechanisms and diseases
Signal Transduction and Targeted Therapy (2024)
-
Unraveling the epigenetic code: human kidney DNA methylation and chromatin dynamics in renal disease development
Nature Communications (2024)
-
DNA methylation and 28-year cardiovascular disease risk in type 1 diabetes: the Epidemiology of Diabetes Complications (EDC) cohort study
Clinical Epigenetics (2023)
-
Integrative genomic analyses in adipocytes implicate DNA methylation in human obesity and diabetes
Nature Communications (2023)
-
DNA methylation in diabetic retinopathy: pathogenetic role and potential therapeutic targets
Cell & Bioscience (2022)