Abstract
Ageing impairs tissue repair. This defect is pronounced in skeletal muscle, whose regeneration by muscle stem cells (MuSCs) is robust in young-adult animals, but inefficient in older organisms. Despite this functional decline, old MuSCs are amenable to rejuvenation through strategies that improve the systemic milieu, such as heterochronic parabiosis. One such strategy, exercise, has long been appreciated for its benefits on healthspan, but its effects on aged stem-cell function in the context of tissue regeneration are incompletely understood. Here, we show that exercise in the form of voluntary wheel running accelerates muscle repair in old mice and improves old MuSC function. Through transcriptional profiling and genetic studies, we discovered that the restoration of old MuSC activation ability hinges on restoration of Cyclin D1, whose expression declines with age in MuSCs. Pharmacologic studies revealed that Cyclin D1 maintains MuSC activation capacity by repressing TGF-β signalling. Taken together, these studies demonstrate that voluntary exercise is a practicable intervention for old MuSC rejuvenation. Furthermore, this work highlights the distinct role of Cyclin D1 in stem-cell quiescence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. RNA-seq data have been deposited in the NCBI Gene Expression Omnibus with the accession code GSE77178. Source data for Figs. 2–4 and Extended Data Figs. 6 and 9 are presented with the paper.
References
Sherwin, C. Voluntary wheel running: a review and novel interpretation. Anim. Behav. 56, 11–27 (1998).
van Praag, H., Shubert, T., Zhao, C. & Gage, F. H. Exercise enhances learning and hippocampal neurogenesis in aged mice. J. Neurosci. 25, 8680–8685 (2005).
Abreu, P., Mendes, S. V. D., Ceccatto, V. M. & Hirabara, S. M. Satellite cell activation induced by aerobic muscle adaptation in response to endurance exercise in humans and rodents. Life Sci. 170, 33–40 (2017).
Kurosaka, M., Naito, H., Ogura, Y., Machida, S. & Katamoto, S. Satellite cell pool enhancement in rat plantaris muscle by endurance training depends on intensity rather than duration. Acta Physiol. (Oxf.) 205, 159–166 (2012).
Dungan, C. M. et al. Elevated myonuclear density during skeletal muscle hypertrophy in response to training is reversed during detraining. Am. J. Physiol. Cell Physiol. 316, C649–C654 (2019).
Egner, I. M., Bruusgaard, J. C. & Gundersen, K. Satellite cell depletion prevents fiber hypertrophy in skeletal muscle. Development 143, 2898–2906 (2016).
Kadi, F., Johansson, F., Johansson, R., Sjöström, M. & Henriksson, J. Effects of one bout of endurance exercise on the expression of myogenin in human quadriceps muscle. Histochem. Cell Biol. 121, 329–334 (2004).
Begue, G. et al. Early activation of rat skeletal muscle IL-6/STAT1/STAT3 dependent gene expression in resistance exercise linked to hypertrophy. PLoS One 8, e57141 (2013).
Fujimaki, S., Hidaka, R., Asashima, M., Takemasa, T. & Kuwabara, T. Wnt protein-mediated satellite cell conversion in adult and aged mice following voluntary wheel running. J. Biol. Chem. 289, 7399–7412 (2014).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Cosgrove, B. D. et al. Rejuvenation of the muscle stem cell population restores strength to injured aged muscles. Nat. Med. 20, 255–264 (2014).
Price, F. D. et al. Inhibition of JAK–STAT signaling stimulates adult satellite cell function. Nat. Med. 20, 1174–1181 (2014).
Nishijo, K. et al. Biomarker system for studying muscle, stem cells, and cancer in vivo. FASEB J. 23, 2681–2690 (2009).
Schultz, E. & Lipton, B. H. Skeletal muscle satellite cells: changes in proliferation potential as a function of age. Mech. Ageing Dev. 20, 377–383 (1982).
Conboy, I. M., Conboy, M. J., Smythe, G. M. & Rando, T. A. Notch-mediated restoration of regenerative potential to aged muscle. Science 302, 1575–1577 (2003).
Rao, S. S. & Kohtz, D. S. Positive and negative regulation of D-type cyclin expression in skeletal myoblasts by basic fibroblast growth factor and transforming growth factor beta. A role for cyclin D1 in control of myoblast differentiation. J. Biol. Chem. 270, 4093–4100 (1995).
Skapek, S. X., Rhee, J., Spicer, D. B. & Lassar, A. B. Inhibition of myogenic differentiation in proliferating myoblasts by cyclin D1-dependent kinase. Science 267, 1022–1024 (1995).
Panda, A. C. et al. Novel RNA-binding activity of MYF5 enhances Ccnd1/Cyclin D1 mRNA translation during myogenesis. Nucleic Acids Res. 44, 2393–2408 (2016).
Ju, X. et al. Identification of a cyclin D1 network in prostate cancer that antagonizes epithelial-mesenchymal restraint. Cancer Res. 74, 508–519 (2014).
Pauklin, S., Madrigal, P., Bertero, A. & Vallier, L. Initiation of stem cell differentiation involves cell cycle-dependent regulation of developmental genes by Cyclin D. Genes Dev. 30, 421–433 (2016).
Zou, P. et al. p57(Kip2) and p27(Kip1) cooperate to maintain hematopoietic stem cell quiescence through interactions with Hsc70. Cell Stem Cell. 9, 247–261 (2011).
Chaves-Ferreira, M. et al. The cyclin D1 carboxyl regulatory domain controls the division and differentiation of hematopoietic cells. Biol. Direct. 11, 21 (2016).
Jeselsohn, R. et al. Cyclin D1 kinase activity is required for the self-renewal of mammary stem and progenitor cells that are targets of MMTV-ErbB2 tumorigenesis. Cancer Cell. 17, 65–76 (2010).
Ma, J. et al. Proliferation and differentiation of neural stem cells are selectively regulated by knockout of cyclin D1. J. Mol. Neurosci. 42, 35–43 (2010).
Bizen, N. et al. A growth-promoting signaling component cyclin D1 in neural stem cells has antiastrogliogenic function to execute self-renewal. Stem Cells 32, 1602–1615 (2014).
Pauklin, S. & Vallier, L. The cell-cycle state of stem cells determines cell fate propensity. Cell 155, 135–147 (2013).
Quelle, D. E. et al. Overexpression of mouse D-type cyclins accelerates G1 phase in rodent fibroblasts. Genes Dev. 7, 1559–1571 (1993).
Resnitzky, D., Gossen, M., Bujard, H. & Reed, S. I. Acceleration of the G1/S phase transition by expression of cyclins D1 and E with an inducible system. Mol. Cell. Biol. 14, 1669–1679 (1994).
Kim, H. A. et al. A developmentally regulated switch directs regenerative growth of Schwann cells through cyclin D1. Neuron 26, 405–416 (2000).
Choi, Y. J. et al. The requirement for cyclin D function in tumor maintenance. Cancer Cell. 22, 438–451 (2012).
Casimiro, M. C. et al. ChIP sequencing of cyclin D1 reveals a transcriptional role in chromosomal instability in mice. J. Clin. Invest. 122, 833–843 (2012).
Casimiro, M. C. et al. Kinase-independent role of cyclin D1 in chromosomal instability and mammary tumorigenesis. Oncotarget. 6, 8525–8538 (2015).
Hinds, P. W., Dowdy, S. F., Eaton, E. N., Arnold, A. & Weinberg, R. A. Function of a human cyclin gene as an oncogene. Proc. Natl Acad. Sci. USA 91, 709–713 (1994).
Du, Z., Tong, X. & Ye, X. Cyclin D1 promotes cell cycle progression through enhancing NDR1/2 kinase activity independent of cyclin-dependent kinase 4. J. Biol. Chem. 288, 26678–26687 (2013).
Li, Z. et al. Alternate cyclin D1 mRNA splicing modulates p27KIP1 binding and cell migration. J. Biol. Chem. 283, 7007–7015 (2008).
Hydbring, P., Malumbres, M. & Sicinski, P. Non-canonical functions of cell cycle cyclins and cyclin-dependent kinases. Nat. Rev. Mol. Cell Biol. 17, 280–292 (2016).
Pestell, R. G. New roles of cyclin D1. Am. J. Pathol. 183, 3–9 (2013).
Rubio, M. F. et al. Cyclin D1 is a NF-κB corepressor. Biochim. Biophys. Acta 1823, 1119–1131 (2012).
Liu, Q. et al. Cyclin D1 and C/EBPβ LAP1 operate in a common pathway to promote mammary epithelial cell differentiation. Mol. Cell. Biol. 34, 3168–3179 (2014).
Matsuura, I. et al. Cyclin-dependent kinases regulate the antiproliferative function of Smads. Nature 430, 226–231 (2004).
Carlson, M. E., Hsu, M. & Conboy, I. M. Imbalance between pSmad3 and Notch induces CDK inhibitors in old muscle stem cells. Nature 454, 528–532 (2008).
Yousef, H. et al. Systemic attenuation of the TGF-β pathway by a single drug simultaneously rejuvenates hippocampal neurogenesis and myogenesis in the same old mammal. Oncotarget 6, 11959–11978 (2015).
Sawyer, J. S. et al. Synthesis and activity of new aryl- and heteroaryl-substituted pyrazole inhibitors of the transforming growth factor-beta type I receptor kinase domain. J. Med. Chem. 46, 3953–3956 (2003).
Luo, K. Signaling cross talk between TGF-β/Smad and other signaling pathways. Cold Spring Harb. Perspect. Biol. 9, a022137 (2017).
Liu, L. et al. Chromatin modifications as determinants of muscle stem cell quiescence and chronological aging. Cell Rep. 4, 189–204 (2013).
Liu, L., Cheung, T. H., Charville, G. W. & Rando, T. A. Isolation of skeletal muscle stem cells by fluorescence-activated cell sorting. Nat. Protoc. 10, 1612–1624 (2015).
Butler, K. V. et al. Rational design and simple chemistry yield a superior, neuroprotective HDAC6 inhibitor, tubastatin A. J. Am. Chem. Soc. 132, 10842–10846 (2010).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol 1, 4 (2001).
Luo, D. et al. Deltex2 represses MyoD expression and inhibits myogenic differentiation by acting as a negative regulator of Jmjd1c. Proc. Natl Acad. Sci. USA 114, E3071–E3080 (2017).
Burzyn, D. et al. A special population of regulatory T cells potentiates muscle repair. Cell 155, 1282–1295 (2013).
Cordelières, F. P. et al. Automated cell tracking and analysis in phase-contrast videos (iTrack4U): development of Java software based on combined mean-shift processes. PLoS One 8, e81266 (2013).
Nussbaum-Krammer, C. I., Neto, M. F., Brielmann, R. M., Pedersen, J. S. & Morimoto, R. I. Investigating the spreading and toxicity of prion-like proteins using the metazoan model organism C. elegans. J. Vis. Exp. https://doi.org/10.3791/52321 (2015).
Gilbert, P. M. et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329, 1078–1081 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Risso, D., Ngai, J., Speed, T. P. & Dudoit, S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 32, 896–902 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Wingender, E. Compilation of transcription regulating proteins. Nucleic Acids Res. 16, 1879–1902 (1988).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Daily, K., Patel, V. R., Rigor, P., Xie, X. & Baldi, P. MotifMap: integrative genome-wide maps of regulatory motif sites for model species. BMC Bioinformatics 12, 495 (2011).
Lachmann, A. et al. ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26, 2438–2444 (2010).
Rouillard, A. D. et al. The harmonizome: a collection of processed datasets gathered to serve and mine knowledge about genes and proteins. Database (Oxford) 2016, baw100 (2016).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Raj, A., van den Bogaard, P., Rifkin, S. A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Mueller, F. et al. FISH-quant: automatic counting of transcripts in 3D FISH images. Nat. Methods 10, 277–278 (2013).
Acknowledgements
We thank J. T. Rodgers, L. Liu and Z. De Miguel for intellectual support, and M. Wagner and I. Akimenko for technical assistance. This work was supported by funding from the Stanford University School of Medicine Medical Scientist Training Program (T32 GM007365) and CIRM Scholar Training Program (TG2 01159) to J.O.B., funding from FAPESP (BEPE 2015/26767-1) to L.A.P., funding from the NIH (TR01 AG047820) to T.W-C. and T.A.R., and funding from the Glenn Foundation for Medical Research, the NIH (P01 AG036695, R37 AG023806, and R01 AR062185) and the Department of Veterans Affairs (Merit Review) to T.A.R.
Author information
Authors and Affiliations
Contributions
J.O.B, M.A., M.I., T.W-C., and T.A.R. designed experiments. J.O.B., M.A., M.I., M.Q., A.d.M., I.M.E., L.A.P., H.D.I., A.G., C.R-M., P.B, D.I.B., and M.J.B. conducted and analysed experiments. J.O.B., M.A. and T.A.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effects of voluntary wheel running on muscle.
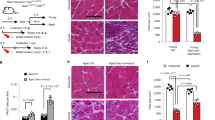
a, Non-strenuous voluntary exercise by wheel running in mice. Young or old mice are provided access to a freely rotating wheel or to a locked wheel as a control. Three weeks later, muscles are either assayed with MuSCs in their quiescent state, without injury or MuSC isolation, or assayed for MuSC exit from quiescence, induced by experimental injury or MuSC isolation into culture. b, Throughout the final week of locked wheel (-Ex) or free wheel (+Ex) access, thymidine analog (EdU or BrdU) was administered continuously in the drinking water. MuSCs were FACS-isolated and immediately fixed for EdU staining. For comparison, also shown are the results from young mice receiving muscle injury at the time of onset of labeling (n=12 for Y(-Ex), 6 for Y(+Ex), 9 for O(-Ex), 9 for O(+Ex), and 2 for Inj mice). c, FACS-isolated MuSCs were assayed for cell size based on forward scatter in flow cytometry. For comparison, also shown are results from young muscles injured three days prior to analysis. Data were normalized to the mean injured level in each experiment (n=5 for Y(-Ex), 5 for Y(+Ex), 6 for O(-Ex), 6 for O(+Ex), and 3 for Inj mice). d, FACS-isolated MuSCs were assayed for RNA content based on Pyronin Y intensity in flow cytometry. For comparison, also shown are results from young muscles injured three days prior to analysis. Data were normalized to the mean injured level in each experiment (n=5 for Y(-Ex), 5 for Y(+Ex), 6 for O(-Ex), 6 for O(+Ex), and 3 for Inj mice). e, FACS-isolated MuSCs were assayed for MyoD expression based on immunocytochemistry. For comparison, also shown are results from young muscles partially injured three days prior to analysis (n=3 for Y(-Ex), 3 for Y(+Ex), 3 for O(-Ex), 3 for O(+Ex), and 1 for Inj mice). f-h, TA muscles were sectioned and assayed for MyoD-expressing cells (f), Ki67-expressing cells (g), and Pax7-expressing cells (h) by immunohistochemistry. For comparison, also shown are results from young muscles injured seven days prior to analysis (n=3 mice per group). i, TA muscle cross-sections were stained with H&E. Representative images (quantified in (j) are shown). j, The mean CSA of myofibers was quantified (n=8 mice per group). k, For each mouse, left and right TA muscles were isolated and their weights averaged (n=6 for Y(-Ex), 6 for Y(+Ex), 5 for O(-Ex), and 5 for O(+Ex) mice). l-n, TA muscles were sectioned and assayed for macrophages expressing F4/80 (l), regenerating myofibers expressing eMHC (m), and regenerating myofibers with central nuclei (n) by immunohistochemistry. For comparison, also shown are results from muscles injured seven days prior to analysis (n=3 mice per group). Scale bar in l, 100 μm. Data are summarized with mean + s.e.m. NS, not significant; *P<0.05; two-tailed Welch’s t-test in b-h, j-n.
Extended Data Fig. 2 Exercise improves multiple aspects of old MuSC regenerative ability.
a–d, Exercise and muscle injury were performed as in Fig. 1a. After either four days (a), five days (b) or twenty-eight days (c-d), muscles were isolated and stained to examine regeneration. a-b, Muscles were sectioned and assayed for eMHC+ myofibers (a, n=7 for Y(-Ex), 5 for Y(+Ex), 7 for O(-Ex), and 4 for O(+Ex) mice. b, n=4 for Y(-Ex), 7 for O(-Ex), and 8 for O(+Ex) mice. Y(+Ex) at five days was not done (N.D.)). c-d, Twenty-eight days post-injury (dpi), the mean cross-sectional areas (CSA) of myofibers (c) and the number of Pax7-expressing cells (d) were quantified (n=3 mice per group). e, Gating strategy for FACS isolation of MuSCs, following a published protocol45,46. Purity of isolated MuSCs is >98% as assessed by routine staining for Pax7 of cells fixed one hour after plating. f, FACS-isolated MuSCs were cultured for eighteen hours and then analyzed for RNA content by flow cytometry based on Pyronin Y staining (n=6 mice per group). g, FACS-isolated MuSCs were tracked by time-lapse microscopy to determine time to first division (n=7 for O(-Ex), 7 for O(+Ex), and 5 for Y(-Ex) mice). h, FACS-isolated MuSCs were tracked by time-lapse microscopy to determine the distance migrated by each cell between serial images (n=7 for O(-Ex), 7 for O(+Ex), and 5 for Y(-Ex) mice). i, FACS-isolated MuSCs were cultured for one day and then stained with 7AAD to determine viability by flow cytometry. Shown is the gating strategy for analysis and the quantification of the fraction of dead cells (n=3 mice per condition). Scale bar in e, 50 μm. Data are summarized with mean + s.e.m. *P<0.05; one-tailed Welch’s t-test in a-d, f-i.
Extended Data Fig. 3 The exercise-induced improvement in old MuSC activation gradually subsides after exercise cessation.
a, Mice were given no access or free access to a running wheel, followed by wheel removal for zero, one, or two weeks. The onset of exercise was staggered so that MuSC isolation was performed at the same time for all groups. b, FACS-isolated MuSCs were cultured continuously in EdU to assess S-phase progression (n=8 for O(-Ex), 7 for O(+Ex)(0 wk), 8 for O(+Ex)(1 wk), and 3 for O(+Ex)(2 wk) mice). Data are summarized with mean + s.e.m. *P<0.05; two-tailed Welch’s t-test in b.
Extended Data Fig. 4 The exercise-induced improvement in old MuSC activation is transferable through serum.
a, Old recipient mice that had never exercised received three consecutive daily tail-vein injections with serum collected from old non-exercising or exercising mice. MuSCs were isolated from recipient mice one day after the last injection. b, FACS-isolated MuSCs were cultured continuously in EdU to assess S-phase progression (n=8 recipient mice for O(-Ex), comprising 4, 3, and 1 recipients for three different serum pools, and n=6 recipient mice for O(+Ex), comprising 3, 2, and 1 recipients for three different serum pools). Data are summarized with mean + s.e.m. *P<0.05; two-tailed Mann-Whitney U-test in b.
Extended Data Fig. 5 Transcriptional effects of aging and exercise in quiescent MuSCs.
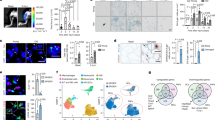
a, RT-qPCR in MuSCs from mice independent of those used in the RNA-Seq experiment. Ct values were normalized first to the mean of Gapdh, Hprt, and Actb1 and then to the mean Y(-Ex) level in each experiment, with Y(-Ex) shown as a dotted line at relative expression 1.0 for comparison (n=13 for O(-Ex), 14 for O(+Ex), and 13 for Y(-Ex) mice). b, GSEA results for the Hallmark gene sets in comparisons of RNA-Seq profiles for O(-Ex) vs. Y(-Ex), O(+Ex) vs. O(-Ex), and Y(+Ex) vs. Y(-Ex) MuSCs. Gene sets are in descending order based on the O(+Ex) vs. O(-Ex) NES. c, Enrichment plots for the INFLAMMATORY RESPONSE gene set. d, Single-cell RT-qPCR for Ccnd1 in freshly isolated MuSCs. For comparison, also shown are results for young MuSCs isolated three days after injury. The pairs on each chip were O(-Ex) vs. O(+Ex) and Y(-Ex) vs. Injured (n=24 cells from one mouse in each group). Data are summarized with mean and s.e.m. in a, box-and-whisker plots (bottom whisker, min; box bottom, 25th percentile; box middle, median; box top, 75th percentile; top whisker, max; “+”, mean) in d. NES, normalized enrichment score in b, c; ES, running enrichment score; S2N, GSEA Signal2Noise ranking metric in c. *P<0.05; two-tailed Welch’s t-test in a, d.
Extended Data Fig. 6 Characterization of Cyclin D1 reduction and expression in MuSCs.
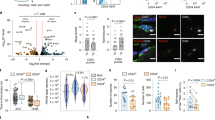
a, Gating strategy for FACS isolation of YFP+ MuSCs after tamoxifen administration to transgenic mice. Shown are MuSC yields in terms of the percentage of size- and doublet-gated cells that are YFP+DAPI-; NS, WT vs. HET and WT vs. KO (n values represent individual mice). Purity of isolated MuSCs is >94% or >98% as assessed by routine staining and quantification of YFP or Pax7, respectively, of cells fixed one hour after plating. b, TA muscles were isolated from twelve-month-old mice that had received tamoxifen injections at three months of age. Muscle sections were stained for Pax7 to identify MuSCs, YFP to identify recombined cells, and laminin to delimit muscle fibers and MuSCs from the interstitium. No MuSCs or YFP+ cells were identified in the interstitium, and no YFP+ cells were Pax7-. The MuSC pool was quantified by counting Pax7+ cells in sections (n=3 mice per group). c, FACS-isolated MuSCs were cultured continuously in the presence of EdU to assess S-phase progression (n=4 for HET(-Ex), 6 for HET(+Ex), 5 for KO(-Ex), 5 for KO(+Ex), and 6 for WT(-Ex) mice). d, To confirm maintenance of ex vivo quiescence by TubA, MuSCs were kept in culture for three days either in quiescence (with TubA) or during activation (with DMSO vehicle) in the continuous presence of EdU and then fixed for analysis. MuSCs were then released for two days in the presence of EdU by removing TubA. MuSCs were then fixed for analysis of exit from quiescence (n=3 mice per condition). e, MuSCs were infected as in Fig. 3j for three days and then harvested for Western blot. Each lane represents a pool of three to six mice split into the two infection conditions. f, MuSCs infected as in Fig. 3j were harvested for RT-qPCR analysis (n=3 mice per group). Scale bar in a, 50 μm, in b, 10 μm. Data are summarized with mean + s.e.m. NS, not significant; *P<0.05; two-tailed Welch’s t-test in a-c, one-tailed Welch’s t-test in d, one-tailed ratio-paired t-test in f.
Extended Data Fig. 7 Gene sets altered by Cyclin D1 reduction and by aging in MuSCs.
a, GSEA results for the Hallmark gene sets in comparisons of RNA-Seq profiles for Y(HET) vs. Y(WT), Y(KO) vs. Y(WT), and O(WT) vs. Y(WT) MuSCs. Gene sets are in ascending order based on the mean NES. b, Enrichment plots for gene sets representing cell cycle genes (E2F TARGETS) and inflammation genes (INFLAMMATORY RESPONSE). NES, normalized enrichment score in a, b; ES, running enrichment score; S2N, GSEA Signal2Noise ranking metric in b.
Extended Data Fig. 8 TGFβ-Smad3 activity is anti-correlated with Ccnd1 in MuSCs.
a, For each gene in the RNA-Seq datasets, a weighted correlation coefficient against Ccnd1 was calculated across all samples. Shown are examples of negative, zero, and positive correlations, in which expression is plotted in log scale and point size conveys sample weight. b, GSEA results for the Hallmark gene sets using the Ccnd1 correlation coefficient of each gene across all samples. Gene sets are in ascending order based on NES. c, GSEA results for TFT gene sets obtained from the Harmonizome database that are experimentally determined (TRANSFAC and ChEA) and computationally predicted (MSigDB and MotifMap). Shown are the top twelve anticorrelated gene sets based on NES for each gene set collection (total gene sets screened: 72 for TRANSFAC, 74 for ChEA, 140 for MSigDB, and 34 for MotifMap). Smad3 is highlighted in each collection. d, Enrichment plots for the Hallmark gene set TGF BETA in each of the previously mentioned RNA-Seq comparisons. NES, normalized enrichment score in b-d; ES, running enrichment score; S2N, GSEA Signal2Noise ranking metric in d.
Extended Data Fig. 9 TGFβ-Smad3 activity in MuSCs with aging, Cyclin D1 deficiency, and pharmacologic modulation.
a-c, Western blots on freshly isolated MuSCs to assess for activating C-terminal phosphorylation of Smad3. Each lane’s phospho-Smad3 level was normalized first to Histone 3 and then to the grand mean of each blot; blots quantified in each figure contained equals numbers of each replicate type. a, MuSCs were from Y(Veh), O(Veh), and O(LY) mice. Shown is a representative blot and quantification of two blots. Each lane represents MuSCs from one mouse (n=6 lanes per group). b, MuSCs were from WT(Veh), HET(Veh), and HET(LY) mice. Shown is a representative blot and quantification of two blots. Each lane represents MuSCs from one mouse (n=6 lanes per group). c, MuSCs were from WT(Veh), KO(Veh), and KO(LY) mice. Shown is a representative blot and quantification of two blots. Each lane represents MuSCs from one mouse (n=6 lanes per group). *P<0.05; one-tailed Mann-Whitney U-test in a-c.
Supplementary information
Source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Unprocessed western blots
Source Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 9
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Brett, J.O., Arjona, M., Ikeda, M. et al. Exercise rejuvenates quiescent skeletal muscle stem cells in old mice through restoration of Cyclin D1. Nat Metab 2, 307–317 (2020). https://doi.org/10.1038/s42255-020-0190-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-0190-0
This article is cited by
-
Depletion of SAM leading to loss of heterochromatin drives muscle stem cell ageing
Nature Metabolism (2024)
-
Exercise Improves Heart Function after Myocardial Infarction: The Merits of AMPK
Cardiovascular Drugs and Therapy (2024)
-
Exercise Promotes Tissue Regeneration: Mechanisms Involved and Therapeutic Scope
Sports Medicine - Open (2023)
-
ATF3 induction prevents precocious activation of skeletal muscle stem cell by regulating H2B expression
Nature Communications (2023)
-
Restoring bone marrow niche function rejuvenates aged hematopoietic stem cells by reactivating the DNA Damage Response
Nature Communications (2023)