Abstract
In non-small-cell lung cancer (NSCLC), concurrent mutations in the oncogene KRAS and the tumour suppressor STK11 (also known as LKB1) encoding the kinase LKB1 result in aggressive tumours prone to metastasis but with liabilities arising from reprogrammed metabolism. We previously demonstrated perturbed nitrogen metabolism and addiction to an unconventional pathway of pyrimidine synthesis in KRAS/LKB1 co-mutant cancer cells. To gain broader insight into metabolic reprogramming in NSCLC, we analysed tumour metabolomes in a series of genetically engineered mouse models with oncogenic KRAS combined with mutations in LKB1 or p53. Metabolomics and gene expression profiling pointed towards activation of the hexosamine biosynthesis pathway (HBP), another nitrogen-related metabolic pathway, in both mouse and human KRAS/LKB1 co-mutant tumours. KRAS/LKB1 co-mutant cells contain high levels of HBP metabolites, higher flux through the HBP pathway and elevated dependence on the HBP enzyme glutamine-fructose-6-phosphate transaminase [isomerizing] 2 (GFPT2). GFPT2 inhibition selectively reduced KRAS/LKB1 co-mutant tumour cell growth in culture, xenografts and genetically modified mice. Our results define a new metabolic vulnerability in KRAS/LKB1 co-mutant tumours and provide a rationale for targeting GFPT2 in this aggressive NSCLC subtype.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All primary data are included in the supplementary materials accompanying this article. Any additional information required to interpret, replicate or build on the methods or findings reported in the article are available from the corresponding authors upon request. Source data are provided with this paper.
References
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Calles, A. et al. Immunohistochemical loss of LKB1 is a biomarker for more aggressive biology in KRAS-mutant lung adenocarcinoma. Clin. Cancer Res. 21, 2851–2860 (2015).
Skoulidis, F. & Heymach, J. V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 19, 495–509 (2019).
Skoulidis, F. et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov. 8, 822–835 (2018).
Racker, E., Resnick, R. J. & Feldman, R. Glycolysis and methylaminoisobutyrate uptake in rat-1 cells transfected with ras or myc oncogenes. Proc. Natl Acad. Sci. USA 82, 3535–3538 (1985).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Bryant, K. L., Mancias, J. D., Kimmelman, A. C. & Der, C. J. KRAS: feeding pancreatic cancer proliferation. Trends Biochem. Sci. 39, 91–100 (2014).
Padanad, M. S. et al. Fatty acid oxidation mediated by Acyl-CoA synthetase long chain 3 is required for mutant KRAS lung tumorigenesis. Cell Rep. 16, 1614–1628 (2016).
Hardie, D. G., Ross, F. A. & Hawley, S. A. AMPK: a nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell Biol. 13, 251–262 (2012).
Shackelford, D. B. & Shaw, R. J. The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009).
Shackelford, D. B. et al. LKB1 inactivation dictates therapeutic response of non-small cell lung cancer to the metabolism drug phenformin. Cancer Cell 23, 143–158 (2013).
Liu, Y. et al. Metabolic and functional genomic studies identify deoxythymidylate kinase as a target in LKB1-mutant lung cancer. Cancer Discov. 3, 870–879 (2013).
Kim, H. S. et al. Systematic identification of molecular subtype-selective vulnerabilities in non-small-cell lung cancer. Cell 155, 552–566 (2013).
Kim, J. et al. CPS1 maintains pyrimidine pools and DNA synthesis in KRAS/LKB1-mutant lung cancer cells. Nature 546, 168–172 (2017).
Hanover, J. A., Krause, M. W. & Love, D. C. The hexosamine signaling pathway: O-GlcNAc cycling in feast or famine. Biochim. Biophys. Acta 1800, 80–95 (2010).
Hawkins, M., Angelov, I., Liu, R., Barzilai, N. & Rossetti, L. The tissue concentration of UDP-N-acetylglucosamine modulates the stimulatory effect of insulin on skeletal muscle glucose uptake. J. Biol. Chem. 272, 4889–4895 (1997).
Denzel, M. S. et al. Hexosamine pathway metabolites enhance protein quality control and prolong life. Cell 156, 1167–1178 (2014).
Olson, A. K., Bouchard, B., Zhu, W. Z., Chatham, J. C. & Des Rosiers, C. First characterization of glucose flux through the hexosamine biosynthesis pathway (HBP) in ex vivo mouse heart. J. Biol. Chem. 295, 2018–2033 (2020).
Hardivillé, S. & Hart, G. W. Nutrient regulation of signaling, transcription, and cell physiology by O-GlcNAcylation. Cell Metab. 20, 208–213 (2014).
Dennis, J. W., Nabi, I. R. & Demetriou, M. Metabolism, cell surface organization, and disease. Cell 139, 1229–1241 (2009).
Kornfeld, R. & Kornfeld, S. Assembly of asparagine-linked oligosaccharides. Annu. Rev. Biochem. 54, 631–664 (1985).
Lau, K. S. et al. Complex N-glycan number and degree of branching cooperate to regulate cell proliferation and differentiation. Cell 129, 123–134 (2007).
Peixoto, A., Relvas-Santos, M., Azevedo, R., Santos, L. L. & Ferreira, J. A. Protein glycosylation and tumor microenvironment alterations driving cancer hallmarks. Front. Oncol. 9, 380 (2019).
Oki, T., Yamazaki, K., Kuromitsu, J., Okada, M. & Tanaka, I. cDNA cloning and mapping of a novel subtype of glutamine:fructose-6-phosphate amidotransferase (GFAT2) in human and mouse. Genomics 57, 227–234 (1999).
Wan, L. et al. ENL links histone acetylation to oncogenic gene expression in acute myeloid leukaemia. Nature 543, 265–269 (2017).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Huang, F. et al. Inosine monophosphate dehydrogenase dependence in a subset of small cell lung cancers. Cell Metab. 28, 369–382.e5 (2018).
Mollaoglu, G. et al. MYC drives progression of small cell lung cancer to a variant neuroendocrine subtype with vulnerability to aurora kinase inhibition. Cancer Cell 31, 270–285 (2017).
Bolscher, J. G. et al. Ras (proto)oncogene induces N-linked carbohydrate modification: temporal relationship with induction of invasive potential. EMBO J. 7, 3361–3368 (1988).
Wojciechowicz, D. C., Park, P. Y. & Paty, P. B. β1-6 branching of N-linked carbohydrate is associated with K-ras mutation in human colon carcinoma cell lines. Biochem. Biophys. Res. Commun. 212, 758–766 (1995).
Wellen, K. E. et al. The hexosamine biosynthetic pathway couples growth factor-induced glutamine uptake to glucose metabolism. Genes Dev. 24, 2784–2799 (2010).
Yang, C. et al. High expression of GFAT1 predicts poor prognosis in patients with pancreatic cancer. Sci. Rep. 6, 39044 (2016).
Taparra, K. et al. O-GlcNAcylation is required for mutant KRAS-induced lung tumorigenesis. J. Clin. Invest. 128, 4924–4937 (2018).
Prat, A. et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 12, R68 (2010).
Zibrova, D. et al. GFAT1 phosphorylation by AMPK promotes VEGF-induced angiogenesis. Biochem. J. 474, 983–1001 (2017).
Hershfield, M. S. & Seegmiller, J. E. Regulation of de novo purine biosynthesis in human lymphoblasts. J. Biol. Chem. 251, 7348–7354 (1976).
Longnecker, D. S. & Curphey, T. J. Adenocarcinoma of the pancreas in azaserine-treated rats. Cancer Res. 35, 2249–2258 (1975).
Ricciardiello, F. et al. Inhibition of the hexosamine biosynthetic pathway by targeting PGM3 causes breast cancer growth arrest and apoptosis. Cell Death Dis. 9, 377 (2018).
Olivier-Van Stichelen, S. et al. The hexosamine biosynthetic pathway and O-GlcNAcylation drive the expression of β-catenin and cell proliferation. Am. J. Physiol. Endocrinol. Metab. 302, E417–E424 (2012).
Slawson, C., Copeland, R. J. & Hart, G. W. O-GlcNAc signaling: a metabolic link between diabetes and cancer? Trends Biochem. Sci. 35, 547–555 (2010).
Häuselmann, I. & Borsig, L. Altered tumor-cell glycosylation promotes metastasis. Front. Oncol. 4, 28 (2014).
Taniguchi, N. & Kizuka, Y. Glycans and cancer: role of N-glycans in cancer biomarker, progression and metastasis, and therapeutics. Adv. Cancer Res. 126, 11–51 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Mullen, A. R. et al. Oxidation of alpha-ketoglutarate is required for reductive carboxylation in cancer cells with mitochondrial defects. Cell Rep. 7, 1679–1690 (2014).
Tomayko, M. M. & Reynolds, C. P. Determination of subcutaneous tumor size in athymic (nude) mice. Cancer Chemother. Pharmacol. 24, 148–154 (1989).
Pham, N. D. et al. Effects of altered sialic acid biosynthesis on N-linked glycan branching and cell surface interactions. J. Biol. Chem. 292, 9637–9651 (2017).
Collisson, E. A. et al. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Győrffy, B., Surowiak, P., Budczies, J. & Lánczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE 8, e82241 (2013).
Acknowledgements
R.J.D. is supported by the Howard Hughes Medical Institute, a Robert L. Moody Sr. Faculty Scholar endowment and by grants from the National Cancer Institute (NCI) (no. R35CA22044901) and Cancer Prevention and Research Institute of Texas (no. RP160089). Jiyeon Kim is supported by the NCI (grant no. 1K22CA226676-01A1), American Lung Association (grant no. LCD-614827) and V Foundation (grant no. V2019-022). J.D.M. is supported by the NCI (SPORE grant no. P50CA070907) and Cancer Prevention and Research Institute of Texas (grant no. RP160652). James Kim is supported by NCI grant no. 1R01CA196851, SPORE grant no. P50CA70907 and the American Cancer Society (grant no. RSG-16-090-01-TBG). S.K. was supported by the National Heart, Lung, and Blood Institute (grant no. 5T32HL098040).
Author information
Authors and Affiliations
Contributions
Jiyeon Kim and R.J.D. conceived the research project, designed the experiments and wrote the paper. Jiyeon Kim, H.M.L., B.K., C.Y., E.L.L., N.M., S.R., K.L., M.H., W.G., B.F., A.K.K., L.C., K.N., S.K., U.M., L.G., X.S. and James Kim performed the research and/or contributed to the analysis and discussions. F.C. developed the LC–MS/MS methods and performed the analysis. J.D.M. provided the cell lines and contributed to the analysis and discussions. H.W. and K.U-K. provided the mouse tumour metabolomics and RNA-seq data.
Corresponding authors
Ethics declarations
Competing interests
R.J.D. is an advisor for Agios Pharmaceuticals. J.D.M. receives cell line licensing royalties from the National Institutes of Health and UTSW. The other authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Christoph Schmitt. Nature Metabolism thanks Sulagna Banerjee, Thales Papagiannakopoulos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Hexosamine-related metabolic pathways are associated with Kras/Lkb1 mouse tumours.
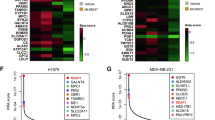
a, GSEA between Kras/Lkb1 tumours and adjacent lungs returned ‘Amino sugar and nucleotide metabolism’, ‘Fructose and mannose metabolism’, and ‘Oxidative Phosphorylation’ as the top ranked metabolic gene ontology terms from KEGG database. Enrichment statistics include nominal p value and nominal enrichment score (NES). b, The leading-edge genes of ‘Amino sugar and nucleotide sugar metabolism’ (left) and ‘Fructose and mannose metabolism’ (right) are in Extended Data Fig. 1a. The leading-edge genes of ‘Oxidative Phosphorylation’ (not displayed here) are in Supplementary Table 3. c, The leading-edge genes of ‘Amino sugar and nucleotide sugar metabolism’, ‘Fructose and mannose metabolism’ and ‘O-Glycan biosynthesis’ in Fig. 1a.
Extended Data Fig. 2 Mouse Kras/Lkb1 tumours do not display CPS1 induction.
a, RNAseq result for CPS1 in mouse lung and mouse tumours of the indicated genotypes. Data are the average and SD of tissues from three independent mice except Kras mice (n = 2). b, CPS1 expression in the tissues used in Extended Data Fig. 2a. q-rt-PCR Data are the average and SD of three fragments from three independent mice except Kras mice (three fragments from two mice). c, CPS1 protein expression in the tissues used in Extended Data Fig. 2b. Statistical significance was assessed using one-way ANOVA followed by Tukey’s post hoc test was used and there was no statistical significance among the groups.
Extended Data Fig. 3 Lkb1, but not p53, mutation significantly alters metabolism of Kras mutant mouse lung tumour.
a, Key metabolites differentiating between Kras (K) and Kras/Lkb1 (KL) tumours (left), Kras/Lkb1 (KL) and Kras/p53 (KP) tumours (middle) and Kras (K) and Kras/p53 (KP) tumours (right) (VIP > ~ 1.5). Metabolites shown in all three groups are in light blue whereas ones only shown in the first and second group are in red and ones only shown in the first and third group are in green. Relative metabolite abundance is indicated in the bar, with red representing metabolite accumulation. b, Venn diagrams of Variable Importance in the Projection metabolites (VIP > 1.0) between [K and KL tumour tissues] and [KP and KL tumour tissues] (left) and between [K and KP tumour tissues] and [K and KL tumour tissues]. c, Abundance of M6P and N-glycolylneuraminic acid (Neu5GC) from the metabolomics in Extended Data Fig. 3a. Individual data points are shown with mean values and SD for ten (KP) or ten (KL) mouse tumours. M6P, mannose-6-phosphate; Neu5Gc, N-glycolylneuraminic acid. Statistical significance was assessed using Wilcoxon signed rank test. *p < 0.05; **p < 0.01.
Extended Data Fig. 4 KRAS/LKB1 (KL) cells show lower basal levels of UPR-ER stress than KRAS (K) cells.
a, Relative intensity of O-GlcNAcylation to loading control in H2122 cells; O-GlcNAcylation signal was derived from the bracketed region of the blot in Fig. 3d. b, Doubling time of LKB1-proficient and -deficient Calu-1 cells (n = 9). c, Left, Abundance of BiP protein in K and KL cell lines. Actin is used as a loading control. Right, Relative intensity of BiP to loading control in K and KL cells. Statistical significance was assessed using two sample unequal variance Student’s t-tests. *p < 0.05. Cell counting for doubling time calculation was performed more than three times and Western blot was repeated twice.
Extended Data Fig. 5 KL cells are sensitive to inhibition of the HBP.
a, Effect of OSMI-1 treatment on global O-GlcNAcylation of K and KL cells. b, Abundance of OGT protein in K and KL cell lines transfected with a control siRNA or siRNA directed against OGT. Actin is used as a loading control. c, Effect of OGT silencing on global O-GlcNAcylation of K and KL cells. d, Relative viability of EV and LKB1-WT expressing H2122 cells following a 72 hr exposure to OSMI-1 (25 µM). e, Relative viability of EV and LKB1-WT expressing H2122 cells to OGT silencing for 96 hr. f, Relative viability of shGFP and shLKB1-expressing Calu-1 (Left) and H1373 (Right) cells to OGT silencing for 96 hr. g,h, Abundance of OGT protein in EV and LKB1 expressing KL cells (H460 and H2122) (g) and shGFP and shLKB1 expressing K cells (Calu-1 and H1373) (h) transfected with a control siRNA or siRNA directed against OGT. Actin is used as a loading control except H1373 (Vinculin is used as a loading control). Statistical significance was assessed using two-tailed Student’s t-tests. ***p < 0.001; ****p < 0.0001. Cell viability assays were repeated twice, and all Western blots were repeated three times or more.
Extended Data Fig. 6 KL cells are sensitive to inhibition of GFPT activity.
a, Effect of azaserine treatment (1 µM) on K and KL cells’ viability (14 cell lines, n = 6). Data are the average and SD of six independent cultures. b, Effect of azaserine treatment on cell death in K and KL cells (n = 3). c, Relative viability of shGFP and shLKB1-expressing H1373 following a 72 hr exposure to azaserine (1 µM). d, Effect of azaserine treatment on global O-GlcNAcylation of K and KL cells. Actin is used as a loading control. Statistical significance was assessed using Student’s t-tests (a) and one-way ANOVA followed by Tukey’s multiple comparisons test (c). In c, *, p < 0.05 comparing to shGFP without azaserine treatment; $, p < 0.05 comparing to shGFP with azaserine treatment; #, p < 0.05 comparing to shLKB1 without azaserine treatment. ****, p < 0.0001. Cell viability assay and Western blots were repeated at least twice. FACS analysis was performed once.
Extended Data Fig. 7 KL cells are sensitive to inhibition of GFPT2.
a, Abundance of GFPT1 protein in cell lines transfected with a control esiRNA or esiRNA directed against GFPT1. Actin is used as a loading control. b, Abundance of GFPT2 protein in cell lines transfected with a control esiRNA or esiRNA directed against GFPT2. Actin is used as a loading control. c, Kaplan−Meier plot associating GFPT1 mRNA expression with survival. Dataset is from KM Plotter (http://kmplot.com/analysis/index.php?p=service&cancer=lung). d, Kaplan−Meier plot associating GFPT2 mRNA expression with survival. e, Abundance of GFPT1 in a panel of K and KL cells. Actin is used as a loading control. f, Abundance of GFPT2 in a panel of K and KL cells. Actin is used as a loading control. g, Effect of Dox-induced GFPT2 deletion on growth in a monolayer culture. Data are the average and SD of 6 replicates. h, Abundance of GFPT2 in Dox-inducible GFPT2 KO H157 (left) and H460 (right) cells with or without Dox induction. Actin is used as a loading control. i, Representative images of colonies grown in soft agar in Fig. 5g. j, Effects of a GFPT2 KO on cell proliferation (H460 cells, n = 5). k, Effect of GlcNAc on anchorage-independent growth of GFPT2 KO cells. l, Global O-GlcNAcylation of GFPT2 WT, KO and KO treated with GlcNAc. m, Abundance of GlcNAc-6-P and ManNAc in Dox-inducible GFPT2 KO H157 cells (n = 3). n, Effect of Dox-inducible GFPT2 KO H157 cells on cell surface L-PHA lectin binding. Statistical significance in g, j, m, and n was assessed using two-tailed Student’s t-tests. In j, to calculate significance on repeated measurements over time, a two-way ANOVA with Tukey’s post hoc test was used. In k, statistical significance was assessed using one-way ANOVA with Tukey post hoc test. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Targeted metabolomics was performed once. Soft agar assay, monolayer cell growth, GFPT1 and 2, and O-GlcNAcylation western blotting were assayed twice. All other experiments were repeated three times or more.
Extended Data Fig. 8 KL cells are sensitive to inhibition of GFPT2 (cont.).
a,b, Abundance of GlcNAc-6-P, ManNAc and GlcNAc-6-P in Dox-inducible negative control (NC) KO H460 (a) and H157 (b) cells (n = 3). c, Abundance of GFPT2 protein in EV and LKB1-expressing H460 transfected with a control esiRNA or esiRNA directed against GFPT2. Actin is used as a loading control. d, Left, Relative viability of EV and LKB1-expressing H2122 cells after GFPT2 silencing for 96 hr. Right, Abundance of GFPT2 protein in EV and LKB1-expressing H2122 cells transfected with a control esiRNA or esiRNA directed against GFPT2. Actin is used as a loading control. e, Relative viability of EV and LKB1-expressing H460 (Left) and H2122 (Right) cells after GFPT1 silencing for 96 hr. f, Abundance of GFPT1 protein in EV and LKB1-expressing H2122 cells transfected with a control esiRNA or esiRNA directed against GFPT1. Actin is used as a loading control. g, Abundance of GFPT2 protein in EV and constitutively active AMPK (CA AMPK)-expressing H460 cells transfected with a control esiRNA or esiRNA directed against GFPT2. Actin is used as a loading control.
Extended Data Fig. 9 GFPT2 suppression inhibits KL tumour growth.
a, Global O-GlcNAcylation in A549 (left) and H460 (right) xenografts in presence and absence of azaserine. Actin was used as a loading control. b, Global O-GlcNAcylation in Calu-1 (left) and Calu-6 (right) xenografts in presence and absence of azaserine. Actin was used as a loading control. c and d, Abundance of GFPT2 protein in Dox-inducible GFPT2 KO H460 (c) and H157 (d) xenografts with or without Dox induction. Actin was used as a loading control. e and f, Growth of Dox-inducible NC KO H460 (e) and H157 (f) xenografts in presence and absence of Dox. Mean tumour volume and SEM are shown for each group (n = 5 per group). g, Growth of Dox-inducible GFPT2 KO Calu-1 xenografts in presence and absence of Dox. Mean tumour volume and SEM are shown for each group (n = 5 for GFPT2-Dox, n = 4 for GFPT2 + Dox). h,i, Abundance of GFPT2 protein in Dox-inducible NC KO H460 (h) and H157 (i) xenografts with or without Dox induction. Actin was used as a loading control. j, Abundance of GFPT2 protein in Dox-inducible GFPT2 KO Calu-1 xenografts with or without Dox induction. Actin was used as a loading control. k, Growth of H460 WT or GFPT2 KO (two different clones) xenografts. Mean tumour volume and SEM are shown for each group (n = 5 per group).l, Abundance of GFPT2 protein in GFPT2 WT and KO (two independent clones) H460 xenografts with or without Dox induction. Note that only three mice bearing KO #2 cells developed tumours. Actin was used as a loading control. Statistical significance in f and g was assessed using two-way ANOVA followed by Sidak’s multiple comparisons test. In e and k, statistical significance was assessed using two-way ANOVA with Tukey’s multiple comparisons test. Mouse experiments were performed once. Western blots were repeated twice.
Extended Data Fig. 10 Azaserine treatment reduces KL tumour burden and proliferation.
a, Tumor area in K tumours with or without azaserine treatment was quantified with ImageJ and % of tumour burden out of total lung was analyzed. Scale bar, 3 mm. b, Tumor area from Extended Data Fig. 10a was quantified with ImageJ and % of tumour burden out of total lung was analyzed. c, Representative Ki67 staining images of the same mouse tissues used in Extended Data Fig. 10a. Scale bar, 100 µm. d, Ki67 positive cells and total cells within the area were quantified using ClickMaster2000. Three images per tissue were used for quantification. Statistical significance was assessed using two-tailed Student’s t-tests.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3
Supplementary Tables
Supplementary Tables 1–6
Supplementary Data
Mouse tumour volume raw data
Source data
Source Data Fig. 2
Uncropped WB data related to Fig. 2
Source Data Fig. 3
Uncropped WB data related to Fig. 3
Source Data Fig. 5
Uncropped western blot data related to Fig. 5
Source Data Extended Data Fig. 2
Uncropped western blot data related to Extended Data Fig. 2
Source Data Extended Data Fig. 4
Uncropped western blot data related to Extended Data Fig. 4
Source Data Extended Data Fig. 5
Uncropped western blot data related to Extended Data Fig. 5
Source Data Extended Data Fig. 6
Uncropped western blot data related to Extended Data Fig. 6
Source Data Extended Data Fig. 7
Uncropped western blot data related to Extended Data Fig. 7
Source Data Extended Data Fig. 8
Uncropped western blot data related to Extended Data Fig. 8
Source Data Extended Data Fig. 9
Uncropped western blot data related to Extended Data Fig. 9
Rights and permissions
About this article
Cite this article
Kim, J., Lee, H.M., Cai, F. et al. The hexosamine biosynthesis pathway is a targetable liability in KRAS/LKB1 mutant lung cancer. Nat Metab 2, 1401–1412 (2020). https://doi.org/10.1038/s42255-020-00316-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00316-0
This article is cited by
-
The GFPT2-O-GlcNAcylation-YBX1 axis promotes IL-18 secretion to regulate the tumor immune microenvironment in pancreatic cancer
Cell Death & Disease (2024)
-
Discovery of metal-binding proteins by thermal proteome profiling
Nature Chemical Biology (2024)
-
Current knowledge and potential intervention of hexosamine biosynthesis pathway in lung cancer
World Journal of Surgical Oncology (2023)
-
Nutlin-3a induces KRAS mutant/p53 wild type lung cancer specific methuosis-like cell death that is dependent on GFPT2
Journal of Experimental & Clinical Cancer Research (2023)
-
What is new in cancer-associated fibroblast biomarkers?
Cell Communication and Signaling (2023)