Abstract
Inhibiting glycolysis remains an aspirational approach for the treatment of cancer. We have previously identified a subset of cancers harbouring homozygous deletion of the glycolytic enzyme enolase (ENO1) that have exceptional sensitivity to inhibition of its redundant paralogue, ENO2, through a therapeutic strategy known as collateral lethality. Here, we show that a small-molecule enolase inhibitor, POMHEX, can selectively kill ENO1-deleted glioma cells at low-nanomolar concentrations and eradicate intracranial orthotopic ENO1-deleted tumours in mice at doses well-tolerated in non-human primates. Our data provide an in vivo proof of principle of the power of collateral lethality in precision oncology and demonstrate the utility of POMHEX for glycolysis inhibition with potential use across a range of therapeutic settings.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
There is no restriction in any data presented in this work. The data that support the findings of this study are available from the corresponding author upon request. The POMHEX NCI-60 Human Tumor Cell Lines Screen data are attached as Supplementary Note 1. The metabolomic data, and Figures and Extended Data Figures source data have been deposited in Figshare (https://doi.org/10.6084/m9.figshare.12931676.v1). The DICOM source MRI scans are stored on a secure sever at MDA and are available on request. X-ray structures of ENO2 bound with HEX (PDB 5IDZ) have been deposited in the Protein Data Bank. Source data are provided with this paper.
Change history
18 December 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Fonvielle, M., Mariano, S. & Therisod, M. New inhibitors of rabbit muscle triose-phosphate isomerase. Bioorg. Med. Chem. Lett. 15, 2906–2909 (2005).
Anderson, V. E., Weiss, P. M. & Cleland, W. W. Reaction intermediate analogues for enolase. Biochemistry 23, 2779–2786 (1984).
Muller, F. L. et al. Passenger deletions generate therapeutic vulnerabilities in cancer. Nature 488, 337–343 (2012).
Leonard, P. G. et al. SF2312 is a natural phosphonate inhibitor of enolase. Nat. Chem. Biol. 12, 1053–1058 (2016).
Boulard-Heitzmann, P. et al. Decreased red cell enolase activity in a 40-year-old woman with compensated haemolysis. Scand. J. Haematol. 33, 401–404 (1984).
Stefanini, M. Chronic hemolytic anemia associated with erythrocyte enolase deficiency exacerbated by ingestion of nitrofurantoin. Am. J. Clin. Pathol. 58, 408–414 (1972).
Muller, F. et al. Enolase inhibitors and methods of treatment therewith. US Patent WO2016145113A1 (2016).
Krucinska, J. et al. Structural and functional studies of bacterial enolase, a potential target against gram-negative pathogens. Biochemistry 58, 1188–1197 (2019).
Jiang H. et al. Examination of the therapeutic potential of Delta-24-RGD in brain tumor stem cells: role of autophagic cell death. J. Natl. Cancer Inst. 99, 1410–1414 (2007).
Figueroa, J. et al. Exosomes from glioma-associated mesenchymal stem cells increase the tumorigenicity of glioma stem-like cells via transfer of miR-1587. Cancer Res. 77, 5808–5819 (2017).
Yang, Y. et al. UOK 262 cell line, fumarate hydratase deficient (FH–/FH–) hereditary leiomyomatosis renal cell carcinoma: in vitro and in vivo model of an aberrant energy metabolic pathway in human cancer. Cancer Genet. Cytogenet. 196, 45–55 (2010).
Chan, D. A. et al. Targeting GLUT1 and the Warburg effect in renal cell carcinoma by chemical synthetic lethality. Sci. Transl. Med. 3, 94ra70 (2011).
Pradere, U., Garnier-Amblard, E. C., Coats, S. J., Amblard, F. & Schinazi, R. F. Synthesis of nucleoside phosphate and phosphonate prodrugs. Chem. Rev. 114, 9154–9218 (2014).
Foster, K. J. et al. Blood intermediary metabolite and insulin concentrations after an overnight fast: reference ranges for adults, and interrelations. Clin. Chem. 24, 1568–1572 (1978).
Vasconcelos-dos-Santos, A. et al. Biosynthetic machinery involved in aberrant glycosylation: promising targets for developing of drugs against cancer. Front. Oncol. 5, 138 (2015).
Tarrado-Castellarnau, M., de Atauri, P. & Cascante, M. Oncogenic regulation of tumor metabolic reprogramming. Oncotarget 7, 62726–62753 (2016).
Pantelouris, E. M. Absence of thymus in a mouse mutant. Nature 217, 370–371 (1968).
Farquhar, D., Khan, S., Srivastva, D. N. & Saunders, P. P. Synthesis and antitumor evaluation of bis[(pivaloyloxy)methyl] 2′-deoxy-5-fluorouridine 5′-monophosphate (FdUMP): a strategy to introduce nucleotides into cells. J. Med. Chem. 37, 3902–3909 (1994).
Duysen, E. G. et al. Production of ES1 plasma carboxylesterase knockout mice for toxicity studies. Chem. Res. Toxicol. 24, 1891–1898 (2011).
Rudakova, E. V., Boltneva, N. P. & Makhaeva, G. F. Comparative analysis of esterase activities of human, mouse, and rat blood. Bull. Exp. Biol. Med. 152, 73–75 (2011).
Bahar, F. G., Ohura, K., Ogihara, T. & Imai, T. Species difference of esterase expression and hydrolase activity in plasma. J. Pharm. Sci. 101, 3979–3988 (2012).
Yan, V. C. & Muller, F. L. Advantages of the parent nucleoside GS-441524 over emdesivir for Covid-19 treatment. ACS Med. Chem. Lett. 11, 1361–1366 (2020).
Humle, N. et al. Targeted vascular drug delivery in cerebral cancer. Curr. Pharm. Des. 22, 5487–5504 (2016).
de Groot, J. F. & Yung, W. K. A. Bevacizumab and irinotecan in the treatment of recurrent malignant gliomas. Cancer J. 14, 279–285 (2008).
Nau, R., Sörgel, F. & Eiffert, H. Penetration of drugs through the blood-cerebrospinal fluid/blood–brain barrier for treatment of central nervous system infections. Clin. Microbiol. Rev. 23, 858–883 (2010).
Brunner, M. et al. Penetration of fosfomycin into the parenchyma of human brain: a case study in three patients. Br. J. Clin. Pharm. 54, 548–550 (2002).
Pfeifer, G., Frenkel, C. & Entzian, W. Pharmacokinetic aspects of cerebrospinal fluid penetration of fosfomycin. Int. J. Clin. Pharmacol. Res. 5, 171–174 (1985).
Naesens, L., Balzarini, J., Bischofberger, N. & De Clercq, E. Antiretroviral activity and pharmacokinetics in mice of oral bis(pivaloyloxymethyl)-9-(2-phosphonylmethoxyethyl)adenine, the bis(pivaloyloxymethyl) ester prodrug of 9-(2-phosphonylmethoxyethyl)adenine. Antimicrob. Agents Chemother. 40, 22–28 (1996).
Van Rompay, K. K. et al. Biological effects of short-term or prolonged administration of 9-[2-(phosphonomethoxy)propyl]adenine (tenofovir) to newborn and infant rhesus macaques. Antimicrob. Agents Chemother. 48, 1469–1487 (2004).
Lockiec, F. Ifosfamide: pharmacokinetic properties for central nervous system metastasis prevention. Ann. Oncol. 17, iv33-6 (2006).
Benito, J. et al. Hypoxia-activated prodrug TH-302 targets hypoxic bone marrow niches in pre-clinical leukemia models. Clin. Cancer Res. 22, 1687–1698 (2016).
Beran, M., Andersson, B. S., Wang, Y., McCredie, K. B. & Farquhar, D. The effects of acetaldophosphamide, a novel stable aldophosphamide analogue, on normal human and leukemic progenitor cells in vitro: implications for use in bone marrow purging. Cancer Res. 48, 339–345 (1988).
Ludeman, S. M. The chemistry of the metabolites of cyclophosphamide. Curr. Pharm. Des. 5, 627–643 (1999).
Mazur, L., Opyodo-Chanek, M., Stojak, M. & Wojcieszek, K. Mafosfamide as a new anticancer agent: preclinical investigations and clinical trials. Anticancer Res. 32, 2783–2789 (2012).
Guise, C. P. et al. Bioreductive prodrugs as cancer therapeutics: targeting tumor hypoxia. Chin. J. Cancer 33, 80–86 (2014).
Mazur, L., Opyodo-Chanek, M. & Stojak, M. Glufosfamide as a new oxazaphosphorine anticancer agent. Anticancer Drugs 22, 488–493 (2011).
Tobias, SandraC. & Borch, RichardF. Synthesis and biological studies of novel nucleoside phosphoramidate prodrugs. J. Med. Chem. 44, 4475–4480 (2001).
Wu, W., Sigmond, J., Peters, G. J. & Borch, R. F. Synthesis and biological activity of a gemcitabine phosphoramidate prodrug. J. Med. Chem. 50, 3743–3746 (2007).
Sjövall, J., Bergdahl, S., Movin, G., Ogenstad, S. & Saarimäki, M. Pharmacokinetics of foscarnet and distribution to cerebrospinal fluid after intravenous infusion in patients with human immunodeficiency virus infection. Antimicrob. Agents Chemother. 33, 1023–1031 (1989).
Wager, T. T., Hou, X., Verhoest, P. R. & Villalobos, A. Central nervous system multiparameter optimization desirability: application in drug discovery. ACS Chem. Neurosci. 7, 3 (2016).
Borch, R. F. et al. Synthesis and evaluation of nitroheterocyclic phosphoramidates as hypoxia-selective alkylating agents. J. Med. Chem. 43, 2258–2265 (2000).
Yan, V. C. et al. Bioreducible phosphonoamidate pro-drug inhibitor of enolase: proof of concept study. ACS Med. Chem. Lett. 11, 1484–1489 (2020).
Valk, P. E., Mathis, C. A., Prados, M. D., Gilbert, J. C. & Budinger, T. F. Hypoxia in human gliomas: demonstration by PET with fluorine-18-fluoromisonidazole. J. Nucl. Med. 33, 2133–2137 (1992).
Yan, V. C., Pham, C.-D. & Muller, F. L. Expedient method for direct mono-amidation of phosphonic and phosphoric acids. Preprint at ChemRxiv https://doi.org/10.26434/CHEMRXIV.12073131.V1 (2020).
Yan, V. C., Pham, C.-D., Arthur, K. & Muller, F. L. Aliphatic amines are viable pro-drug moieties in phosphonoamidate drugs. Preprint at bioRxiv https://doi.org/10.1101/2020.04.05.026583 (2020).
Powers, J. F. et al. A unique model for SDH-deficient GIST: an endocrine-related cancer. Endocr. Relat. Cancer 25, 943–954 (2018).
Maitituoheti, M. et al. Enhancer reprogramming confers dependence on glycolysis and IGF signaling in KMT2D mutant melanoma. Cell Rep. 33, 108293 (2020).
Alam, H. et al. Super-enhancer impairment is a link between MLL4-inactivated lung tumors and their vulnerability to glycolysis pathway inhibition. Preprint at bioRxiv https://doi.org/10.1101/507202 (2018).
German, M. S. Glucose sensing in pancreatic islet beta cells: the key role of glucokinase and the glycolytic intermediates. Proc. Natl Acad. Sci. USA 90, 1781–1785 (1993).
Gaffney, DominiqueO. et al. Non-enzymatic lysine lactoylation of glycolytic enzymes. Cell Chem. Biol. 27, 206–213 (2020).
Muller, F., Muller, F., Aquilanti, E. & DePinho, R. In vitro enzymatic activity assay for ENOLASE in mammalian cells in culture. Preprint at Protocol Exchange https://doi.org/10.1038/protex.2012.040 (2012).
Gillies, R. J., Didier, N. & Denton, M. Determination of cell number in monolayer cultures. Anal. Biochem. 159, 109–113 (1986).
Kueng, W., Silber, E. & Eppenberger, U. Quantification of cells cultured on 96-well plates. Anal. Biochem. 182, 16–19 (1989).
Bady, P. et al. DNA fingerprinting of glioma cell lines and considerations on similarity measurements. Neuro. Oncol. 14, 701–711 (2012).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008).
Nistér, M. et al. Evidence for progressional changes in the human malignant glioma line U-343 MGa: analysis of karyotype and expression of genes encoding the subunit chains of platelet-derived growth factor. Cancer Res. 47, 4953–4960 (1987).
Lal, S. et al. An implantable guide-screw system for brain tumor studies in small animals. J. Neurosurg. 92, 326–333 (2000).
Yuan, M., Breitkopf, S. B., Yang, X. & Asara, J. M. A positive/negative ion–switching, targeted mass spectrometry–based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Geoff, T. et al. Biological crystallography iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. Sect. D. Biol. Crystallogr. 67, 271–281 (2011).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. Sect. D. Biol. Crystallogr. 69, 1204–1214 (2013).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. Sect. D. Biol. Crystallogr. 68, 352–367 (2012).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 213–221 (2010).
Lacy, S. A., M. J. Hitchcock, Lee, W. A., Telliert, P. & Cundy, K. C. Effect of oral probenecid coadministration on the chronic toxicity and pharmacokinetics of intravenous cidofovir in cynomolgus monkeys effect of oral probenecid coadministration on the chronic toxicity and pharmacokinetics of intravenous cidofovir in cyno-molgus monkeys. Toxicol. Sci. 44, 97–106 (1998).
Van Rompay, K. K. A., Hamilton, M., Kearney, B. & Bischofberger, N. Pharmacokinetics of tenofovir in breast milk of lactating rhesus macaques. Antimicrob. Agents Chemother. 49, 2093–2094 (2005).
Van Rompay, K. K. A. et al. Biological effects of short-term or prolonged administration of 9-[2-(phosphonomethoxy)propyl]adenine (tenofovir) to newborn and infant rhesus macaques. Antimicrob. Agents Chemother. 48, 2346 (2004).
Naesens, L. et al. Antiretroviral efficacy and pharmacokinetics of oral bis(isopropyloxycarbonyloxymethyl)-9-(2-phosphonylmethoxypropyl)adenine in mice. Antimicrob. Agents Chemother. 42, 1568–1573 (1998).
Duwal, S., Schütte, C. & von Kleist, M. Pharmacokinetics and pharmacodynamics of the reverse transcriptase inhibitor tenofovir and prophylactic efficacy against HIV-1 infection. PLoS ONE 7, e40382 (2012).
Deeks, S. G. et al. Safety, pharmacokinetics, and antiretroviral activity of intravenous 9-[2-(R)-(phosphonomethoxy)propyl]adenine, a novel anti-human immunodeficiency virus (HIV) therapy, in HIV-infected adults. Antimicrob. Agents Chemother. 42, 2380–2384 (1998).
Cundy, K. C. et al. Pharmacokinetics and bioavailability of the anti-human immunodeficiency virus nucleotide analog 9-[(R)-2-(phosphonomethoxy)propyl]adenine (PMPA) in dogs. Antimicrob. Agents Chemother. 42, 687–690 (1998).
Zykov, I. N. et al. Pharmacokinetics and pharmacodynamics of fosfomycin and its activity against extended-spectrum-lactamase-, plasmid-mediated AmpC-, and carbapenemase-producing Escherichia coli in a murine urinary tract infection model. Antimicrob. Agents Chemother. 62, e02560-17 (2018).
Pérez, D. S., Tapia, M. O. & Soraci, A. L. Fosfomycin: uses and potentialities in veterinary medicine. Open Vet. J. 4, 26–43 (2014).
Kirby, W. M. M. Pharmacokinetics of fosfomycin. Chemotherapy 23, 141–151 (1977).
Murakawa, T., Sakamoto, H., Fukada, S., Konishi, T. & Nishida, M. Pharmacokinetics of fosmidomycin, a new phosphonic acid antibiotic. Antimicrob. Agents Chemother. 21, 224–230 (1982).
Acknowledgements
We thank S. Millward, S. Gammon, F. Lang, D. Pwinica-Worms, J. De Groot, P. Cserjesi, C. Halbrook, C. Lyssiotis and J. Huse for critical comments and suggestions. We thank H. Butterfield, L. Jacobs, A. Portal, P. Shresta, M. Washington and P. Suriyamongkol for technical assistance. We thank J. Holton and G. Meigs for their assistance with X-ray diffraction data collection. We thank K. Kalurachchi and J. McMurray for assistance with NMR. The UT M.D. Anderson Cancer Center NMR Core Facility is supported in part by the NCI Cancer Center Support Grant (CA16672). We thank M. Yuan and S. Breitkopf for help with the mass-spectrometry experiments. We thank Z. Jia for technical support with NHP experiments at CRL. We thank J. Long for help with statistical analysis and for critical reading. We thank K. Michel, C. Kingsley, V. Tran and the rest of the SAIF team for expert assistance with MRI imaging, and G. Rao, V. Henry, J. Gumin and F. Lang for GSC’s and intracranial cell implantations. The Pharmaceutical Chemistry Facility and the Small Animal Imaging Facility at The UT M.D. Anderson Cancer Center were supported by the NIH/NCI Cancer Center Support Grant under award number P30CA016672. Financial support was provided by NIH CDP SPORE P50CA127001-07 (F.L.M.), NIH SPORE 2P50CA127001-11A1 (F.L.M.), ACS RSG-15-145-01-CDD (F.L.M.), NCCN YIA170032 (F.L.M. Young Investigator Award), The Andrew Sabin Family Foundation (F.L.M. Fellow award), The Dr. Marnie Rose Foundation (F.L.M.), The Uncle Kory Foundation (F.L.M.) and The University of Texas MD Anderson Cancer Center Institutional Research Grant (F.L.M.), CPRIT RP140612 (R.A.D.) and NIH R01 CA225955 (R.A.D). S.K. was supported in part by MD Anderson Cancer Center CPRIT Research Training Program Grant RP170067. This work is dedicated to the memory of Dr. John Stuart McMurray, an expert in phosphonate prodrug synthesis, without whom this would not have been possible.
Author information
Authors and Affiliations
Contributions
F.L.M. and D.M. conceived HEX and POMHEX, which were synthesized by Z.P. and W.B. Z.K., F.P., P.M., B.C., C.-D.P, V.C.Y. and F.L.M. repeated chemical syntheses; V.C.Y., B.C. and F.L.M. wrote synthetic procedures with characterizations. N.S. and F.L.M. performed in vitro enzymatic activity inhibitor experiments. P.G.L. performed isolation of ENO2 recombinant protein and X-ray crystallography with HEX. R.Z. and W.P. performed Seahorse experiments. N.H., N.S., S.K., S.C.P., X.W., T.T. and K.A performed cell-culture enolase-inhibitor sensitivity testing. J.J.A. performed western blots. Y.B. performed in NMR experiments. Y.-H.L. and F.L.M. performed 13C-NMR tracing and biochemical profiling experiments. J.M.A. performed mass spectrometry and small-molecule metabolite analysis. Q.X., Q.W., Y.J., Y.S. and J.R.M. performed pharmacological studies. X.W., S.K. and Y-H.L. performed mouse drug treatment and Y-H.L. performed small-animal MRI tumour imaging. J.R.M., Y.S., D.K.G., V.C.Y. and F.L.M. generated figures, and performed data analysis and statistics. F.L.M. and R.A.D. oversaw the project. V.C.Y. and F.L.M. wrote the manuscript
Corresponding author
Ethics declarations
Competing interests
F.L.M. and R.A.D. are inventors on a patent covering the concept of targeting ENO1-deleted tumours with inhibitors of ENO2 (US patent 9,452,182 B2). F.L.M., R.A.D., D.M., W.B., Y.-H.L, N.S., Z.P., B.C. and F.P. are inventors on a patent application for the use of enolase inhibitors for the treatment of ENO1-deleted tumours (US patent 10,363,261 B2). V.C.Y, C-D.P and FLM are inventors on a patent application describing the synthesis and utility of novel prodrug inhibitors of enolase US 62/797,315 (filed 27 January 2019) R.A.D. is founder, director and/or advisor of Tvardi Therapeutics, Asylia Therapeutics and Stellanova Therapeutics, and the focus of these companies is not related to the content of this manuscript.
Additional information
Peer review information Primary Handling Editor: George Caputa.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Rapid induction of anemia by non-ENO2-specific Enolase inhibitors.
a, Non pro-drugged phosphonate Enolase inhibitors were administered to mice by tail vein IV injection at the times indicated (white arrows) to determine effects on hematocrit (fraction of RBC / total plasma volume). Each trace represents an individual mouse treated and sampled repeatedly over 10 days, with two animals per drug or vehicle (green). Phosphoacetohydroxamate (PhAH; purple) caused a rapid and significant drop in hematocrit (and visible jaundice), which is then restored to initial levels after discontinuing treatment. PhAH is a pan-Enolase inhibitor with slightly greater potency against ENO1 over ENO2. As ENO1 is the sole isoform expressed in RBCs, anemia caused by PhAH is most likely due to on-target inhibitory activity against ENO1. b, Representative capillary tube after centrifugation from the hematocrit experiment described in a. The ratio of RBC fraction to total blood volume is decreased in response to PhAH treatment; pale yellow plasma in the PhAH sample indicates hemolysis. c,d, POMSF is a POM-pro-drug version of SF2312 that was generated for in vivo testing. Mice were IV tail vein injected with either vehicle (DMSO), POMSF, or POMHEX dissolved in PBS at the indicated doses. Capillary tubes were used to measure percent hematocrit. c, Hematocrit was determined as a function of time in response to Enolase inhibitors over a 10-day treatment course. Mice were administered either DMSO (green), POMSF (blue), or POMHEX (red) for 5 days before discontinuing treatment. Compared to the DMSO control and POMHEX, administration of POMSF results in decreased percent hematocrit, which is then restored to initial levels after discontinuing treatment. Significant differences by non-paired t-test with Bonferroni correction are indicated in d. e, Representative capillary tubes from mice treated with DMSO vehicle (left), POMSF (middle), and POMHEX (right). Note the decrease hematocrit (red bracket) as fraction of total blood volume (black bracket) from POMSF treatment; pale orange plasma indicates hemolysis.
Extended Data Fig. 2 HEX is a substrate-competitive inhibitor with preference for ENO2.
a, b, Enolase activity (1/v, y-axis) was measured spectrophotometrically in vitro using an NADH-coupled assay1,2, with ENO1 and ENO2 activity plotted as a function of substrate concentration (x-axis, 1/S: 2-phosphoglycerate in mM; HEX in nM). Data were plotted as Lineweaver-Burke with x-axis showing inverse of substrate versus inverse activity (1/V). Each dot represents on independent biochemical rate determination. Slopes (Km-apparent/Vmax) were re-plotted as a function of inhibitor concentration in Fig. 1c. c. Enolase activity in lysates from ENO1 and ENO2 overexpressing D423 cells, measured as a function of HEX concentration; 0.5 mM 2-PG substrate concentration.
Extended Data Fig. 3 The Gli56 ENO1-homozygously deleted glioma cell line is sensitive to POMHEX and is pyruvate auxotrophic.
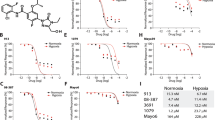
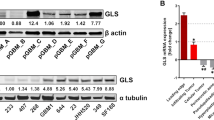
Sensitivity to POMHEX was determined in additional ENO1-homozygous deleted cell line, Gli56 (dark red) and its isogenic ENO1-rescued control3,4. a, Absence of ENO1 protein confirmed by western blot in the Gli56 cell line; note levels of ENO2 are lower than in D423 (ENO1-/-), which is confirmed by mRNA. b, c, RNA-seq mRNA expression of ENO1 and ENO2 in cell lines differing in ENO1-deletion status, confirming lower ENO2-expression in Gli56 compared to D423 cell. Each point represent one biological replicate, means are shown, n number is indicated for each cell line. d, Gli56 (dark red) is selectively sensitive to POMHEX compared to its isogenic ENO1-rescued control3 (blue). Another Gli56 rescued cell line (purple), in which ENO1 and a neighboring deleted gene, NMNAT1, are re-expressed (generated for a separate collateral lethality project), also shows resistance to POMHEX. Cell density was visualized by crystal violet after 14 days of growth; fresh media (DMEM, contains 1.25 mM pyruvate) containing Enolase inhibitor was provided every 4 days (N = 4 biological replicates +/- S.EM.). In contrast to the D423 cell line, Gli56 cannot be grown in pyruvate-free DMEM media, likely due to lower level of residual ENO2 expression. e, f, Gli56 ENO1-deleted, but not Gli56 ENO1-rescued cells, can grow in pyruvate-free DMEM. g, h, D423 ENO1-deleted cells can grow in pyruvate-free DMEM media, but slower than D423 ENO1-rescued or LN319 (ENO1-WT) cells. N = 8 biological replicates, mean +/- S.E.M. P value by 2-way ANOVA are indicated.
Extended Data Fig. 4 Relative sensitivities of glioma cell lines to the pro-drug POMHEX versus the active Enolase inhibitor HEX.
Sensitivity of a panel of glioma cell lines with with varying ENO1 statuses to the Enolase inhibitors POMHEX and HEX. IC50 values were calculated based on terminal cell density measured by crystal violet. D502 and U343 are ENO1-heterozygous deleted cell lines (~50% total Enolase3,4). Consistent with our previous reports for pan-Enolase inhibitors3,4, ENO1-homozygous deletion confers the greatest sensitivity to HEX, while ENO1-heterozygous deletion status shows intermediate sensitivity. While sensitivity to HEX is dependent on ENO1-deletion status, the sensitivity to POMHEX like depends not only on ENO1-deletion status but also expression of pro-drug bioactivation enzymes that transform POMHEX into active HEX enolase inhibitor (carboxylesterases, CES and phosphodiesterases, PDE). On average, the potency of POMHEX is ~75-fold greater than HEX though with substantial variation across cell lines. D502 is considerably more sensitive to POMHEX than U343 (IC50 82 vs 559 nM), yet U343 is more sensitive to HEX than D502 (IC50 19,723 nM vs 28,756 nM). This can be explained by higher levels of expression of pro-drug activating enzymes in the D502 glioma cell line result in greater sensitivity to POMHEX as compared to U343. Identification of the specific genes responsible, and their expression could be used for patient stratification, expanding the utility of Enolase inhibitors beyond those with ENO1-homozygous deletions.
Extended Data Fig. 5 ENO1-deletion status predicts sensitivity to Enolase inhibitors in glioma sphere- forming cells (GSC).
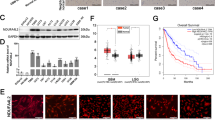
GSCs and omics data were kindly shared by the Lang lab9,10. a, Copy number variation at the 1p36 locus identifies GSC296 (red) as ENO1-homozygous deleted and GSC231, 268, 275, 289 as heterozygous deleted (pink). Dark blue regions represent bi-allelic (homozygous) deletion, while light blue regions correspond to mono-allelic loss. b, Confirmed lack of ENO1 expression by western blot in GSC296 (red), with corresponding levels of ENO2 shown. The western blot analysis for GSC296 was performed at least 3 times with similar outcomes. c, RNA-seq mRNA expression of ENO1 (bottom) and ENO2 (top) in ENO1-homozygous deleted and heterozygous deleted GSCs, confirming lack of ENO1 expression. Each bar represents one biological replicate. d,e, Sensitivity of GSCs with varying ENO1-deletion status to POMHEX. Dissociated GSCs were seeded in 96-well round bottom-low attachment plates in duplicate and allowed to form spheroids. Spheroids were treated with a serial dilution of POMHEX for 1 week; media was changed every 3 days with fresh drug. Viability was visualized with 5 nM TMRE (red). TMRE was quantified by ImageJ and expressed as a function of vehicle controls. Each dot represents one biological replicate. GSC296 (ENO1-/-) was distinctly sensitive to POMHEX. ENO1-heterozygous deleted GSCs showed intermediate sensitivity. ENO1-WT GSCs with the lowest residual ENO2 expression showed the greatest sensitivity amongst ENO1-WT GSCs. Experiments with GSC296 were reproduced independently once.
Extended Data Fig. 6 Normal and near-normal cell lines are minimally sensitive to POMHEX.
All indicated cells were grown in RPMI and treated with a serial dilution of POMHEX for 5 days. Cell density was then quantified by crystal violet after 5 days of growth and expressed relative to control (n = 2 biological replicates). a, A panel of hepatocellular carcinoma cell lines (SNU-423, SNU-398, SK-HEP-1; grey) and highly differentiated, hepatoblastoma line (HepG2 C3A; blue) were treated with POMHEX; D423 (red) was tested as a point of reference. The IC50 of POMHEX for most HCC cell lines is about 750 nM (grey), which is slightly lower than that for ENO1-WT glioma cell lines. However, the highly differentiated cell line, HepG2 C3A, was essentially insensitive with an IC50 >10,000 nM. This resistance likely derives from the dependence of hepatocytes on OxPhos, with minimal requirements for glycolysis-derived ATP (excess ATP allows for glycolysis reversal, gluconeogenesis) and concurs with our data in NHP, which indicate no hepatotoxicity. b, HEK293 kidney cells (grey) are immortalized, non-transformed kidney cells. Sensitivity to POMHEX was comparable to that observed in ENO1-WT cancer cell lines. c, Non-immortalized human astrocytes (grey) showed IC50 of >2,500 nM to POMHEX, which is also comparable to ENO1-WT glioma cell lines and far higher than that for ENO1-deleted D423 cells.
Extended Data Fig. 7 Utility of Enolase inhibitors beyond cancers with ENO1-deletions.
Hyperactivation of HIF by loss of VHL tumor suppressor gene has been reported to sensitize cells to glycolysis inhibition12. a, Proliferation of renal clear cell carcinoma cell lines that lack functional VHL (786-O and RCC4, red) and isogenic rescued cell lines re-expressing VHL (blue) was followed by Incucyte live cell imager (x-axis, time; y-cell confluence) in response to POMHEX treatment. Each box represents one biological replicates. VHL-deleted parental lines showed 4- to 8-fold greater sensitivity than isogenic rescued VHL lines. b, Cell density quantified by crystal violet for the same cell lines corroborates Incucyte live imaging data (N = 4 biological replicates, with +/- S.E.M.). IC50 for POMHEX in the RCC4 line is ~250 nM and 1,000 nM in the isogenic rescued line. TCA-cycle deficiency and defective oxidative phosphorylation by loss of function of Fumarase (FH) increase reliance on glycolysis and sensitize to glucose deprivation11. c, Likewise, the FH-null renal carcinoma cell line, UOK262, (red) shows dramatically higher sensitivity to the Enolase inhibitor POMHEX than isogenic FH-rescued cell line control (blue). Cell density in response to POMHEX treatment was quantified by crystal violet (N = 4 biological replicates, with +/- S.E.M.).
Extended Data Fig. 8 The synthetic cell permeable pyruvate pro-metabolite, methyl 2-oxopropanoate, significantly attenuates toxicity of POMHEX.
ENO1-deleted (D423, red), ENO1-isogenically rescued (D423 ENO1, blue), and ENO1-WT (LN319, grey) cells were treated with POMHEX at the doses indicated (x-axis) in media (DMEM) free of pyruvate with either (a) 2.5 mM methyl 2-oxopropanoate “methyl pyruvate,” (b) 5 mM lactate, or (c) 5 mM acetate (d). Cell density after 5 days exposure was determined by crystal violet staining and expressed relative to non-drug contain controls (n = 4 biological replicates as indicated, +/ S.E.M.). The IC50 for pyruvate-free media is indicated by a dashed line, for comparison. Exogenous methyl pyruvate attenuates sensitivity to Enolase inhibitors especially in ENO1-homozygously deleted cells (IC50 shifted from ~20 nM to ~150 nM), but supplementation with lactate or acetate had minimal effects. (e) Methyl pyruvate is a cell-permeable, synthetic pro-metabolite of pyruvate that can passively diffuse into the cell without requiring monocarboxylate transporter (SLC16) activity. It can then be hydrolyzed by intracellular carboxylesterases to release pyruvate. While lactate and acetate could serve a similar purpose, the present data (c, d) suggest that the rate of acetyl-CoA production by these carbon sources is insufficient to compensate for loss of ATP and pyruvate by POMHEX-mediated inhibition of glycolysis.
Extended Data Fig. 9 ShRNA knockdown of ENO2 recapitulates metabolic disruptions caused by small-molecule ENO2 inhibitors.
a, Knockdown of ENO2 by doxycycline-inducible shRNA in D423 ENO1-deleted and ENO1-rescued cells with the same constructs and cell lines used previously3. Induction of shENO2 results in ~70% decrease in ENO2-protein levels (ratios of band densities ENO2 to TPI, loading control, are indicated). The western blot analysis was performed once with each line representing a single biological replicate. Polar metabolites were profiled at specific times after induction of shENO2 with doxycycline (times indicated in x-axis, hours) in both conditioned media (extracellular) and cell pellet (intracellular), with cells passaged after 120 h. b, Schematic showing the Enolase reaction in the context of central carbon metabolism and associated pathways, with metabolites altered selectively in D423 ENO1-deleted cells in response to ENO2-knockdown. c-e, Accumulation of metabolites upstream of Enolase in response to ENO2-knockdown in ENO1-deleted but not ENO1-rescued cells recapitulates observation with Enolase inhibitors (Fig. 4). Glycerate (c, d) was the most significantly altered metabolite in polar metabolomic profile of conditioned media in ENO1-deleted but not rescued cells. Intracellular glycerate levels (c) recapitulate this trend and mirror the levels of 3-PG (e), from which it is hydrolyzed. f-j, TCA-cycle metabolites were selectively decreased in response to ENO2-knockdown, recapitulating effects of small molecule Enolase inhibition (Fig. 4). Similar to observations with small molecule Enolase inhibition, hexosamine biosynthesis pathway (HBP) metabolites increase in response to ENO2 knockdown. k-l, Each bar represents one biological replicate. P values from T-test of the log of the normalized values of each biological replicate are indicated.
Extended Data Fig. 10 Phosphonate-containing drugs show similar, species-specific pharmacokinetics.
The PK properties of phosphonate containing drugs have been well-studied in diverse model species as well as human patients and exhibit remarkable similarity, despite extensive chemical diversity. The very similar PK properties of these diverse drugs, derives from the fact their physiochemistry and pharmacokinetics are predominantly dictated by the negatively charged phosphonate group, conferring water solubility and limited protein binding. This water solubility means that drugs distribute with water and are cleared by a predominantly renal mechanism. Renal excretion is passive glomerular filtration and active tubular secretion, which can be modulated by drugs like Probenecid64. Rodent species rapidly eliminate phosphonate drugs, with >95% of injected drug excreted unchanged within 1 hr. In non-human primates, half-lives are typically 10-times longer, and in human patients even more so. This copious body of PK data provides a guide towards predictions of how HEX might behave in human patients. The comparison with fosmidomycin is especially informative as it is nearly identical, except for 1 C-C bond, to fosmidomycin. The elimination half-life of HEX in rodents and NHP falls very much in line with other phosphonate drugs and allows reasonable confidence in PK prediction in other models (canines) and in human patients. N/A = data not found in the literature.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Notes 1 and 2
Source data
Source Data Fig. 3
Uncropped Western Blot
Source Data Extended Data Fig. 3
Uncropped Western Blot
Source Data Extended Data Fig. 5
Uncropped Western Blot
Source Data Extented Data Fig. 9
Uncropped Western Blot
Rights and permissions
About this article
Cite this article
Lin, YH., Satani, N., Hammoudi, N. et al. An enolase inhibitor for the targeted treatment of ENO1-deleted cancers. Nat Metab 2, 1413–1426 (2020). https://doi.org/10.1038/s42255-020-00313-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00313-3
This article is cited by
-
Integration of transcriptomics and proteomics to elucidate inhibitory effect and mechanism of rosmarinic acid from Perilla frutescens (L.) Britt. in treating Trichophyton mentagrophytes
Chinese Medicine (2023)
-
Mediation of PKM2-dependent glycolytic and non-glycolytic pathways by ENO2 in head and neck cancer development
Journal of Experimental & Clinical Cancer Research (2023)
-
ENO2-derived phosphoenolpyruvate functions as an endogenous inhibitor of HDAC1 and confers resistance to antiangiogenic therapy
Nature Metabolism (2023)
-
Non-metabolic role of alpha-enolase in virus replication
Molecular Biology Reports (2023)
-
Targeting cancer metabolism in the era of precision oncology
Nature Reviews Drug Discovery (2022)