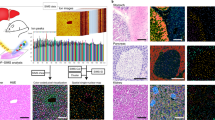
Abstract
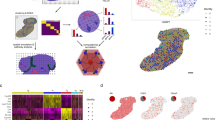
The mammalian liver is composed of repeating hexagonal units termed lobules. Spatially resolved single-cell transcriptomics has revealed that about half of hepatocyte genes are differentially expressed across the lobule, yet technical limitations have impeded reconstructing similar global spatial maps of other hepatocyte features. Here, we show how zonated surface markers can be used to sort hepatocytes from defined lobule zones with high spatial resolution. We apply transcriptomics, microRNA (miRNA) array measurements and mass spectrometry proteomics to reconstruct spatial atlases of multiple zonated features. We demonstrate that protein zonation largely overlaps with messenger RNA zonation, with the periportal HNF4α as an exception. We identify zonation of miRNAs, such as miR-122, and inverse zonation of miRNAs and their hepatocyte target genes, highlighting potential regulation of gene expression levels through zonated mRNA degradation. Among the targets, we find the pericentral Wingless-related integration site (Wnt) receptors Fzd7 and Fzd8 and the periportal Wnt inhibitors Tcf7l1 and Ctnnbip1. Our approach facilitates reconstructing spatial atlases of multiple cellular features in the liver and other structured tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
The code used to generate the processed data and figures is available upon request.
Data availability
The RNA-seq data that support the findings of this study have been deposited with the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under bioproject accession code PRJNA556572, and the SRA identifiers SAMN12360372–SAMN12360382. Supplementary Table 2 summarizes the unique molecular identifier counts per million for each sample. Supplementary Table 11 summarizes the zUMI barcodes used for each sample and the corresponding zUMI settings.
The liquid chromatography–tandem mass spectrometry proteomic data were uploaded to the ProteomeXchange via the PRIDE archive, with project identifier PXD014512. Processed data can be found in Supplementary Table 3.
The miRNA microarray data have been deposited with the NCBI Gene Expression Omnibus under accession code GSE134827. Supplementary Table 6 summarizes the miRNA data.
References
Hoehme, S. et al. Prediction and validation of cell alignment along microvessels as order principle to restore tissue architecture in liver regeneration. Proc. Natl Acad. Sci. USA 107, 10371–10376 (2010).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2 + cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
Colnot, S. & Perret, C. in Molecular Pathology of Liver Diseases (ed. Monga, P. S.) 7–16 (Springer, 2011).
Ben-Moshe, S. & Itzkovitz, S. Spatial heterogeneity in the mammalian liver. Nat. Rev. Gastroenterol. Hepatol. 16, 395–410 (2019).
Gebhardt, R. Metabolic zonation of the liver: regulation and implications for liver function. Pharmacol. Ther. 53, 275–354 (1992).
Jungermann, K. & Keitzmann, T. Zonation of parenchymal and nonparenchymal metabolism in liver. Annu. Rev. Nutr. 16, 179–203 (1996).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Lindros, K. O. & Penttilä, K. E. Digitonin-collagenase perfusion for efficient separation of periportal or perivenous hepatocytes. Biochem. J. 228, 757–760 (1985).
Quistorff, B., Grunnet, N. & Cornell, N. W. Digitonin perfusion of rat liver. A new approach in the study of intra-acinar and intracellular compartmentation in the liver. Biochem. J. 226, 289–297 (1985).
Budnik, B., Levy, E., Harmange, G. & Slavov, N. SCoPE-MS: mass spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation. Genome Biol. 19, 161 (2018).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Doi, Y. et al. Development of complementary expression patterns of E- and N-cadherin in the mouse liver. Hepatol. Res. 37, 230–237 (2007).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Azimifar, S. B., Nagaraj, N., Cox, J. & Mann, M. Cell-type-resolved quantitative proteomics of murine liver. Cell Metab. 20, 1076–1087 (2014).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Odom, D. T. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004).
Torre, C., Perret, C. & Colnot, S. Molecular determinants of liver zonation. Prog. Mol. Biol. Transl. Sci. 97, 127–150 (2010).
Stanulović, V. S. et al. Hepatic HNF4α deficiency induces periportal expression of glutamine synthetase and other pericentral enzymes. Hepatology 45, 433–444 (2007).
Colletti, M. et al. Convergence of Wnt signaling on the HNF4α-driven transcription in controlling liver zonation. Gastroenterology 137, 660–672 (2009).
Brosch, M. et al. Epigenomic map of human liver reveals principles of zonated morphogenic and metabolic control. Nat. Commun. 9, 4150 (2018).
Holloway, M. G., Miles, G. D., Dombkowski, A. A. & Waxman, D. J. Liver-specific hepatocyte nuclear factor-4α deficiency: greater impact on gene expression in male than in female mouse liver. Mol. Endocrinol. 22, 1274–1286 (2008).
Guo, H., Ingolia, N. T., Weissman, J. S. & Bartel, D. P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Lagos-Quintana, M. et al. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739 (2002).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009).
Sekine, S., Ogawa, R., Mcmanus, M. T., Kanai, Y. & Hebrok, M. Dicer is required for proper liver zonation. J. Pathol. 219, 365–372 (2009).
Arvey, A., Larsson, E., Sander, C., Leslie, C. S. & Marks, D. S. Target mRNA abundance dilutes microRNA and siRNA activity. Mol. Syst. Biol. 6, 363 (2010).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73 (2014).
Bissels, U. et al. Absolute quantification of microRNAs by using a universal reference. RNA 15, 2375–2384 (2009).
Tsai, W.-C. et al. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J. Clin. Invest. 122, 2884–2897 (2012).
Li, W. F., Dai, H., Ou, Q., Zuo, G.-Q. & Liu, C. A. Overexpression of microRNA-30a-5p inhibits liver cancer cell proliferation and induces apoptosis by targeting MTDH/PTEN/AKT pathway. Tumour Biol. 37, 5885–5895 (2016).
Kornfeld, J.-W. et al. Obesity-induced overexpression of miR-802 impairs glucose metabolism through silencing of Hnf1b. Nature 494, 111–115 (2013).
Trajkovski, M. et al. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 474, 649–653 (2011).
Agarwal, V., Bell, G. W., Nam, J.-W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, e05005 (2015).
Benhamouche, S. et al. Apc tumor suppressor gene is the “zonation-keeper” of mouse liver. Dev. Cell 10, 759–770 (2006).
Birchmeier, W. Orchestrating Wnt signalling for metabolic liver zonation. Nat. Cell Biol. 18, 463–465 (2016).
Burke, Z. D. & Tosh, D. The Wnt/β‐catenin pathway: master regulator of liver zonation? Bioessays 28, 1072–1077 (2006).
Gebhardt, R. & Hovhannisyan, A. Organ patterning in the adult stage: the role of Wnt/β‐catenin signaling in liver zonation and beyond. Dev. Dyn. 239, 45–55 (2010).
Thompson, M. D. & Monga, S. P. S. WNT/β-catenin signaling in liver health and disease. Hepatology 45, 1298–1305 (2007).
Yang, J. et al. β-catenin signaling in murine liver zonation and regeneration: a Wnt-Wnt situation! Hepatology 60, 964–976 (2014).
Planas-Paz, L. et al. The RSPO–LGR4/5–ZNRF3/RNF43 module controls liver zonation and size. Nat. Cell Biol. 18, 467–479 (2016).
Preziosi, M., Okabe, H., Poddar, M., Singh, S. & Monga, S. P. Endothelial Wnts regulate β‐catenin signaling in murine liver zonation and regeneration: a sequel to the Wnt–Wnt situation. Hepatol. Commun. 2, 845–860 (2018).
Rocha, A. S. et al. The angiocrine factor Rspondin3 is a key determinant of liver zonation. Cell Rep. 13, 1757–1764 (2015).
Shy, B. R. et al. Regulation of Tcf7l1 DNA binding and protein stability as principal mechanisms of Wnt/β-catenin signaling. Cell Rep. 4, 1–9 (2013).
Tago, K. et al. Inhibition of Wnt signaling by ICAT, a novel β-catenin-interacting protein. Genes Dev. 14, 1741–1749 (2000).
Wang, K. et al. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl Acad. Sci. USA 106, 4402–4407 (2009).
Zucman-Rossi, J., Villanueva, A., Nault, J.-C. & Llovet, J. M. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology 149, 1226–1239.e4 (2015).
Chu, A. et al. Large-scale profiling of microRNAs for The Cancer Genome Atlas. Nucleic Acids Res. 44, e3 (2016).
Moor, A. E. et al. Spatial reconstruction of single enterocytes uncovers broad zonation along the intestinal villus axis. Cell 175, 1156–1167.e15 (2018).
Park, J. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Guder, W. G. & Ross, B. D. Enzyme distribution along the nephron. Kidney Int. 26, 101–111 (1984).
Xu, H. et al. Liver‐enriched transcription factors regulate microRNA‐122 that targets CUTL1 during liver development. Hepatology 52, 1431–1442 (2010).
Berndt, N., Horger, M. S., Bulik, S. & Holzhütter, H.-G. A multiscale modelling approach to assess the impact of metabolic zonation and microperfusion on the hepatic carbohydrate metabolism. PLoS Comput. Biol. 14, e1006005 (2018).
Godoy, P. et al. Recent advances in 2D and 3D in vitro systems using primary hepatocytes, alternative hepatocyte sources and non-parenchymal liver cells and their use in investigating mechanisms of hepatotoxicity, cell signaling and ADME. Arch. Toxicol. 87, 1315–1530 (2013).
Holzhütter, H.-G., Drasdo, D., Preusser, T., Lippert, J. & Henney, A. M. The virtual liver: a multidisciplinary, multilevel challenge for systems biology. Wiley Interdiscip. Rev. Syst. Biol. Med. 4, 221–235 (2012).
Jopling, C. L., Yi, M., Lancaster, A. M., Lemon, S. M. & Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific microRNA. Science 309, 1577–1581 (2005).
Roderburg, C. et al. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology 53, 209–218 (2011).
Mitchell, P. S. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl Acad. Sci. USA 105, 10513–10518 (2008).
Farid, W. R. et al. Hepatocyte‐derived microRNAs as serum biomarkers of hepatic injury and rejection after liver transplantation. Liver Transpl. 18, 290–297 (2012).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Hirayama, A. et al. Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res. 69, 4918–4925 (2009).
Llufrio, E. M., Wang, L., Naser, F. J. & Patti, G. J. Sorting cells alters their redox state and cellular metabolome. Redox Biol. 16, 381–387 (2018).
Manco, R. et al. Reactive cholangiocytes differentiate into proliferative hepatocytes with efficient DNA repair in mice with chronic liver injury. J. Hepatol. 70, 1180–1191 (2019).
Mederacke, I., Dapito, D. H., Affò, S., Uchinami, H. & Schwabe, R. F. High-yield and high-purity isolation of hepatic stellate cells from normal and fibrotic mouse livers. Nat. Protoc. 10, 305–315 (2015).
Tanami, S. et al. Dynamic zonation of liver polyploidy. Cell Tissue Res. 368, 405–410 (2017).
Bagnoli, J. W. et al. Sensitive and powerful single-cell RNA sequencing using mcSCRB-seq. Nat. Commun. 9, 2937 (2018).
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. zUMIs: a fast and flexible pipeline to process RNA sequencing data with UMIs. Gigascience 7, giy059 (2018).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Li, B. et al. Adult mouse liver contains two distinct populations of cholangiocytes. Stem Cell Rep. 9, 478–489 (2017).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Milo, R. et al. Network motifs: simple building blocks of complex networks. Science 298, 824–827 (2002).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Bernhardt, J., Funke, S., Hecker, M. & Siebourg, J. Visualizing Gene Expression Data Via Voronoi Treemaps (IEEE, accessed 23 August 2019); https://ieeexplore.ieee.org/document/5362329
Liebermeister, W. et al. Visual account of protein investment in cellular functions. Proc. Natl Acad. Sci. USA 111, 8488–8493 (2014).
Otto, A. et al. Systems-wide temporal proteomic profiling in glucose-starved Bacillus subtilis. Nat. Commun. 1, 137 (2010).
Acknowledgements
We thank E. Hagai and the Flow Cytometry Unit (Weizmann Institute of Science) for FACS technical support; T. Ziv and the Smoler Proteomics Center (Technion) for performing the liquid chromatography–tandem mass spectrometry and analysing the results; and D. Pilzer and the Genomic Technologies Unit (Weizmann Institute of Science) for performing the miRNA microarray measurements. We thank all members of the laboratory for valuable comments. S.I. is supported by the Henry Chanoch Krenter Institute for Biomedical Imaging and Genomics, The Leir Charitable Foundations, the Richard Jakubskind Laboratory of Systems Biology, the Cymerman-Jakubskind Prize, The Lord Sieff of Brimpton Memorial Fund, the I-CORE programme of the Planning and Budgeting Committee (grant no. 1902/12), the Israel Science Foundation (grant no. 1486/16), the Broad Institute-Israel Science Foundation (grant no. 2615/18), the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant no. 768956), the Bert L. and N. Kuggie Vallee Foundation, the Howard Hughes Medical Institute (HHMI) International Research Scholar Award (grant no. 55008734), The Wolfson Family Charitable Trust (grant no. 21376) and the Edmond de Rothschild Foundations.
Author information
Authors and Affiliations
Contributions
K.B.H. and S.I. conceived the study. S.B.M. and S.I. designed the experiments. S.B.M. prepared all the samples. S.B.M. and Y.S. analysed the data. A.E.M. contributed to the data analysis. R.M. and T.V. assisted with the immunofluorescence experiments. K.B.H. contributed to establishing the methodology. S.I. supervised the study. S.B.M., Y.S. and S.I. wrote the manuscript. All authors reviewed the manuscript and provided input.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Elena Bellafante.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ben-Moshe, S., Shapira, Y., Moor, A.E. et al. Spatial sorting enables comprehensive characterization of liver zonation. Nat Metab 1, 899–911 (2019). https://doi.org/10.1038/s42255-019-0109-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0109-9
This article is cited by
-
Single-cell spatial multi-omics and deep learning dissect enhancer-driven gene regulatory networks in liver zonation
Nature Cell Biology (2024)
-
A spatial map of hepatic mitochondria uncovers functional heterogeneity shaped by nutrient-sensing signaling
Nature Communications (2024)
-
Quantifying fat zonation in liver lobules: an integrated multiscale in silico model combining disturbed microperfusion and fat metabolism via a continuum biomechanical bi-scale, tri-phasic approach
Biomechanics and Modeling in Mechanobiology (2024)
-
A spatiotemporal atlas of mouse liver homeostasis and regeneration
Nature Genetics (2024)
-
Liver in infections: a single-cell and spatial transcriptomics perspective
Journal of Biomedical Science (2023)