Abstract
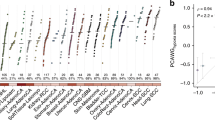
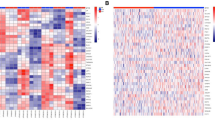
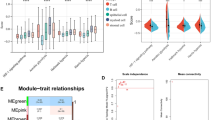
Tumour hypoxia is a major contributor to resistance to anticancer therapies. Given that the results of hypoxia-targeted therapy trials have been disappointing, a more personalized approach may be needed. Here, we characterize multi-omic molecular features associated with tumour hypoxia and identify molecular alterations that correlate with both drug-resistant and drug-sensitive responses to anticancer drugs. Based on a well-established hypoxia gene expression signature, we classify about 10,000 tumour samples into hypoxia score-high and score-low groups across different cancer types from The Cancer Genome Atlas (TCGA) and demonstrate their prognostic associations. Then, we identify various types of molecular features associated with hypoxia status that correlate with drug resistance but, in some cases, also with drug sensitivity, contrasting the conventional view that hypoxia confers drug resistance. We further show that 110 out of 121 (90.9%) clinically actionable genes can be affected by hypoxia status and experimentally validate the predicted effects of hypoxia on the response to several drugs in cultured cells. Our study provides a comprehensive molecular-level understanding of tumour hypoxia and may have practical implications for clinical cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
Codes were implemented in R and have been deposited in GitHub: https://github.com/youqiongye/HAMFA.
References
Bertout, J. A., Patel, S. A. & Simon, M. C. The impact of O2 availability on human cancer. Nat. Rev. Cancer 8, 967–975 (2008).
Walsh, J. C. et al. The clinical importance of assessing tumor hypoxia: relationship of tumor hypoxia to prognosis and therapeutic opportunities. Antioxid. Redox Signal. 21, 1516–1554 (2014).
Wigerup, C., Påhlman, S. & Bexell, D. Therapeutic targeting of hypoxia and hypoxia-inducible factors in cancer. Pharmacol. Ther. 164, 152–169 (2016).
Harris, B. H. L., Barberis, A., West, C. M. L. & Buffa, F. M. Gene expression signatures as biomarkers of tumour hypoxia. Clin. Oncol. (R. Coll. Radiol.) 27, 547–560 (2015).
Fox, N. S., Starmans, M. H. W., Haider, S., Lambin, P. & Boutros, P. C. Ensemble analyses improve signatures of tumour hypoxia and reveal inter-platform differences. BMC Bioinformatics 15, 170 (2014).
Buffa, F. M., Harris, A. L., West, C. M. & Miller, C. J. Large meta-analysis of multiple cancers reveals a common, compact and highly prognostic hypoxia metagene. Br. J. Cancer 102, 428–435 (2010).
Thienpont, B. et al. Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 537, 63–68 (2016).
Black, J. C. et al. Hypoxia drives transient site-specific copy gain and drug-resistant gene expression. Genes Dev. 29, 1018–1031 (2015).
Amelio, I. & Melino, G. The p53 family and the hypoxia-inducible factors (HIFs): determinants of cancer progression. Trends Biochem. Sci. 40, 425–434 (2015).
Marhold, M. et al. HIF1α regulates mTOR signaling and viability of prostate cancer stem cells. Mol. Cancer Res. 13, 556–564 (2015).
Krock, B. L., Skuli, N. & Simon, M. C. Hypoxia-induced angiogenesis: good and evil. Genes Cancer 2, 1117–1133 (2011).
Masoud, G. N. & Li, W. HIF-1α pathway: role, regulation and intervention for cancer therapy. Acta Pharm. Sin. B 5, 378–389 (2015).
Dang, K. & Myers, K. A. The role of hypoxia-induced miR-210 in cancer progression. Int. J. Mol. Sci. 16, 6353–6372 (2015).
Koumenis, C. et al. Regulation of protein synthesis by hypoxia via activation of the endoplasmic reticulum kinase PERK and phosphorylation of the translation initiation factor eIF2α. Mol. Cell. Biol. 22, 7405–7416 (2002).
Wilson, W. R. & Hay, M. P. Targeting hypoxia in cancer therapy. Nat. Rev. Cancer 11, 393–410 (2011).
Lara, P. C. et al. Severe hypoxia induces chemo-resistance in clinical cervical tumors through MVP over-expression. Radiat. Oncol. 4, 29 (2009).
Samanta, D., Gilkes, D. M., Chaturvedi, P., Xiang, L. & Semenza, G. L. Hypoxia-inducible factors are required for chemotherapy resistance of breast cancer stem cells. Proc. Natl Acad. Sci. USA 111, E5429–E5438 (2014).
Gatenby, R. A. et al. Oxygen distribution in squamous cell carcinoma metastases and its relationship to outcome of radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 14, 831–838 (1988).
Minakata, K. et al. Hypoxia induces gefitinib resistance in non-small-cell lung cancer with both mutant and wild-type epidermal growth factor receptors. Cancer Sci. 103, 1946–1954 (2012).
Scharping, N. E., Menk, A. V., Whetstone, R. D., Zeng, X. & Delgoffe, G. M. Efficacy of PD-1 blockade is potentiated by metformin-induced reduction of tumor hypoxia. Cancer Immunol. Res. 5, 9–16 (2017).
Reddy, K. R., Guan, Y., Qin, G., Zhou, Z. & Jing, N. Combined treatment targeting HIF-1α and Stat3 is a potent strategy for prostate cancer therapy. Prostate 71, 1796–1809 (2011).
Seeber, L. M. S., Zweemer, R. P., Verheijen, R. H. M. & van Diest, P. J. Hypoxia-inducible factor-1 as a therapeutic target in endometrial cancer management. Obstet. Gynecol. Int. 2010, 580971 (2010).
Liu, S. K. et al. A novel poly(ADP-ribose) polymerase inhibitor, ABT-888, radiosensitizes malignant human cell lines under hypoxia. Radiother. Oncol. 88, 258–268 (2008).
Jiang, Y. et al. Hypoxia potentiates the radiation-sensitizing effect of olaparib in human non-small cell lung cancer xenografts by contextual synthetic lethality. Int. J. Radiat. Oncol. Biol. Phys. 95, 772–781 (2016).
Patel, P. H. et al. Hypoxia-inducible factor (HIF) 1α and 2α levels in cell lines and human tumor predicts response to sunitinib in renal cell carcinoma (RCC). J. Clin. Oncol. 26, 5008 (2008).
Salem, A. et al. Targeting hypoxia to improve non-small cell lung cancer outcome. J. Natl Cancer Inst. 110, 14–30 (2018).
Ramanathan, R. K. et al. A randomized phase II study of PX-12, an inhibitor of thioredoxin in patients with advanced cancer of the pancreas following progression after a gemcitabine-containing combination. Cancer Chemother. Pharmacol. 67, 503–509 (2011).
Williamson, S. K. et al. Phase III trial of paclitaxel plus carboplatin with or without tirapazamine in advanced non-small-cell lung cancer: Southwest Oncology Group Trial S0003. J. Clin. Oncol. 23, 9097–9104 (2005).
Davidson, A. et al. A phase III randomized trial of adding topical nitroglycerin to first-line chemotherapy for advanced nonsmall-cell lung cancer: the Australasian lung cancer trials group NITRO trial. Ann. Oncol. 26, 2280–2286 (2015).
Chang, K. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Xiang, Y., Ye, Y., Zhang, Z. & Han, L. Maximizing the utility of cancer transcriptomic data. Trends Cancer 4, 823–837 (2018).
Haider, S. et al. Genomic alterations underlie a pan-cancer metabolic shift associated with tumour hypoxia. Genome Biol. 17, 140 (2016).
Winter, S. C. et al. Relation of a hypoxia metagene derived from head and neck cancer to prognosis of multiple cancers. Cancer Res. 67, 3441–3449 (2007).
Hu, Z. et al. A compact VEGF signature associated with distant metastases and poor outcomes. BMC Med. 7, 9 (2009).
Bratslavsky, G., Sudarshan, S., Neckers, L. & Linehan, W. M. Pseudohypoxic pathways in renal cell carcinoma. Clin. Cancer Res. 13, 4667–4671 (2007).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Zhang, H. et al. Integrated proteogenomic characterization of human high-grade serous ovarian cancer. Cell 166, 755–765 (2016).
Li, L. & Greene, T. A weighting analogue to pair matching in propensity score analysis. Int. J. Biostat. 9, 215–234 (2013).
Yuan, Y. et al. Comprehensive characterization of molecular differences in cancer between male and female patients. Cancer Cell 29, 711–722 (2016).
Deng, J. et al. Comparative genomic analysis of esophageal squamous cell carcinoma between Asian and Caucasian patient populations. Nat. Commun. 8, 1533 (2017).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Eales, K. L., Hollinshead, K. E. R. & Tennant, D. A. Hypoxia and metabolic adaptation of cancer cells. Oncogenesis 5, e190 (2016).
Tang, X. et al. A joint analysis of metabolomics and genetics of breast cancer. Breast Cancer Res 16, 415 (2014).
Yang, W. et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–D961 (2013).
Glück, A. A., Aebersold, D. M., Zimmer, Y. & Medová, M. Interplay between receptor tyrosine kinases and hypoxia signaling in cancer. Int. J. Biochem. Cell Biol. 62, 101–114 (2015).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Qu, Y. et al. MiR-139-5p inhibits HGTD-P and regulates neuronal apoptosis induced by hypoxia-ischemia in neonatal rats. Neurobiol. Dis. 63, 184–193 (2014).
Tanizaki, J. et al. MET tyrosine kinase inhibitor crizotinib (PF-02341066) shows differential antitumor effects in non-small cell lung cancer according to MET alterations. J. Thorac. Oncol. 6, 1624–1631 (2011).
Li, C. et al. Fibronectin induces epithelial-mesenchymal transition in human breast cancer MCF-7 cells via activation of calpain. Oncol. Lett. 13, 3889–3895 (2017).
Georgescu, M. M. PTEN tumor suppressor network in PI3K-Akt pathway control. Genes Cancer 1, 1170–1177 (2010).
Kawauchi, K., Araki, K., Tobiume, K. & Tanaka, N. p53 regulates glucose metabolism through an IKK-NF-κB pathway and inhibits cell transformation. Nat. Cell Biol. 10, 611–618 (2008).
Chesnelong, C. et al. Lactate dehydrogenase A silencing in IDH mutant gliomas. Neuro-oncology 16, 686–695 (2014).
Engelman, J. A. et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 316, 1039–1043 (2007).
Schulte, A. et al. Erlotinib resistance in EGFR-amplified glioblastoma cells is associated with upregulation of EGFRvIII and PI3Kp110δ. Neuro-oncology 15, 1289–1301 (2013).
Topalian, S. L., Taube, J. M., Anders, R. A. & Pardoll, D. M. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat. Rev. Cancer 16, 275–287 (2016).
Van Allen, E. M. et al. Whole-exome sequencing and clinical interpretation of formalin-fixed, paraffin-embedded tumor samples to guide precision cancer medicine. Nat. Med. 20, 682–688 (2014).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Kazandjian, D. et al. FDA approval summary: nivolumab for the treatment of metastatic non-small cell lung cancer with progression on or after platinum-based chemotherapy. Oncologist 21, 634–642 (2016).
Geeleher, P. et al. Discovering novel pharmacogenomic biomarkers by imputing drug response in cancer patients from large genomics studies. Genome Res. 27, 1743–1751 (2017).
Ye, Y. et al. The genomic landscape and pharmacogenomic interactions of clock genes in cancer chronotherapy. Cell Syst. 6, 314–328.e2 (2018).
Shen, Y. et al. MiR-375 is upregulated in acquired paclitaxel resistance in cervical cancer. Br. J. Cancer 109, 92–99 (2013).
Stegeman, H. et al. Interaction between hypoxia, AKT and HIF-1 signaling in HNSCC and NSCLC: implications for future treatment strategies. Future Sci. OA 2, FSO84 (2016).
Byers, L. A. et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin. Cancer Res. 19, 279–290 (2013).
Kim, E. S. et al. The BATTLE trial: personalizing therapy for lung cancer. Cancer Discov. 1, 44–53 (2011).
Li, B. et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 17, 174 (2016).
Zheng, X., Zhang, N., Wu, H. J. & Wu, H. Estimating and accounting for tumor purity in the analysis of DNA methylation data from cancer studies. Genome Biol. 18, 17 (2017).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinformatics 14, 7 (2013).
Xiang, Y. et al. Comprehensive characterization of alternative polyadenylation in human cancer. J. Natl Cancer Inst. 110, 379–389 (2018).
Acknowledgements
This work was supported by the Cancer Prevention & Research Institute of Texas (grant no. RR150085 to L.H., grant no. RP140462 to H.L., grant nos. RP150094 and RP180259 to C.L. and grant no. R1218 to L.Y.); the National Institutes of Health (grant nos. CA168394, CA098258 and CA143883 to G.B.M., grant no. CA175486 to H.L., grant no. CA209851 to H.L. and G.B.M., grant no. R00DK094981, 1R01CA218025 and 1R01CA231011 to C.L., grant no. R00CA166527 and 1R01CA218036 to L.Y. and grant no. R01 HL137990 and 1R01HL136969 to Y.X.). Department of Defense Breakthrough Awards were granted to C.L. and L.Y. (award no. BC180196 to C.L. and award no. BC151465 to L.Y.). The American Association for Cancer Research–Bayer Innovation and Discovery Grant (no. 18-80-44) and Andrew Sabin Family Foundation Fellows Award were awarded to L.Y., J.G. was awarded an MD Anderson Physician Scientist Award, a Khalifa Physician Scientist Award, an Andrew Sabin Family Foundation Fellows Award, an MD Anderson Faculty Scholar Award and a Doris Duke Charitable Foundation Career Development Award (award no. 2018097). The National Natural Science Foundation of China supported S.Z. with grant nos. 81822034 and 81773119. We gratefully acknowledge contributions from the TCGA Research Network. We thank L.-A. Chastain for editorial assistance.
Author information
Authors and Affiliations
Contributions
L.H. conceived and supervised the project. Y.Ye and L.H. designed and performed the research. Y.Ye, H.C., Y.Yuan, Y.Xiang, H.R., Z.Z., A.S., H.Z., L.L. and L.D. performed the data analysis. Y.Ye, Q.H. K.L., C.L., L.Y. and L.H. performed the drug tests. Y.Ye, Y.L., B.Z., S.Z., J.G., E.J., S.H.L., L.W., Y.Xia, L.Y., C.L., G.B.M., H.L. and L.H. interpreted the results. Y.Ye, Q.H., G.B.M., H.L. and L.H. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
G.B.M. has sponsored research support from AstraZeneca, Critical Outcomes Technologies, Karus Therapeutics, Illumina, Immunomet, NanoString, Tarveda Therapeutics and Immunomet. He is on the Scientific Advisory Boards of AstraZeneca, Critical Outcomes Technologies, Immunomet, Ionis Pharmaceuticals, Nuevolution, Symphogen and Tarveda Therapeutics. H.L. is a shareholder and scientific advisor of Precision Scientific and Eagle Nebula. J.G. serves as a consultant for ARMO Biosciences, AstraZeneca, Jounce Therapeutics, Nektar and Pfizer.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–10 and Supplementary Tables 1–3.
Supplementary Data 1
Hypoxia-associated features across 21 cancer types
Supplementary Data 2
Spearman correlation of hypoxia-associated genes and drug sensitivity of drugs in GDSC
Supplementary Data 3
Spearman correlation of hypoxia score and imputed drug response across cancer types
Rights and permissions
About this article
Cite this article
Ye, Y., Hu, Q., Chen, H. et al. Characterization of hypoxia-associated molecular features to aid hypoxia-targeted therapy. Nat Metab 1, 431–444 (2019). https://doi.org/10.1038/s42255-019-0045-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0045-8
This article is cited by
-
Ferroptosis in cancer: From molecular mechanisms to therapeutic strategies
Signal Transduction and Targeted Therapy (2024)
-
Characterization of hypoxia-responsive states in ovarian cancer to identify hot tumors and aid adjuvant therapy
Discover Oncology (2024)
-
Epithelial-to-mesenchymal transition in cancer progression: unraveling the immunosuppressive module driving therapy resistance
Cancer and Metastasis Reviews (2024)
-
Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
Four science stars on the fast-track to impact
Nature (2023)