Abstract
Complex systems, ranging from developing embryos to systems of locally communicating agents, display an apparent capability of “programmable” pattern formation: They reproducibly form target patterns, but those targets can be readily changed. A distinguishing feature of such systems is that their subunits are capable of information processing. Here, we explore schemes for programmable pattern formation within a theoretical framework, in which subunits process local signals to update their discrete state following logical rules. We study systems with different update rules, topologies, and control schemes, assessing their capability of programmable pattern formation and their susceptibility to errors. Only a fraction permits local organizers to dictate any target pattern, by transcribing temporal patterns into spatial patterns, reminiscent of the principle underlying vertebrate somitogenesis. An alternative scheme employing variable rules cannot reach all patterns but is insensitive to the timing of organizer inputs. Our results establish a basis for designing synthetic systems and models of programmable pattern formation closer to real systems.
Similar content being viewed by others
Introduction
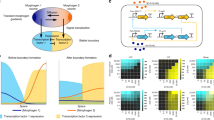
Programmable pattern formation is impressively exemplified in developmental biology, where relatively minor changes in the cis-regulatory regions of genes can reprogram the developmental process to yield dramatic changes in the morphology of the adult organism1,2. In these systems, the individual cells have internal states, but do not know the global state of the system. They process local cues according to their genetic program to determine how and when to change their internal state. Local organizers like the Spemann Mangold organizer3 can control the internal states of other cells, but there is no global agent overseeing the pattern formation process. The underlying biological concept is inductive signaling, whereby one cell can change the fate of another cell4. For instance, in Caenorhabditis elegans vulval patterning, the ‘anchor cell’ controls the cell fate pattern of six vulval precursor cells, involving three different cell fates5. Programmable pattern formation can also emerge on a higher level in agent-based systems, when e.g. groups of robots6,7 or humans8 coordinate their motion by local communication. These examples motivate the conceptual question: Which general schemes allow the same agents to produce different complex patterns by following rules to coordinate their behavior with their neighbors?
While natural systems consist of subunits that are already very complex, it is interesting to ask for the simplest model systems capable of programmable pattern formation. Such models would provide a conceptual framework and could reveal design principles, e.g., for synthetic molecular systems. DNA-based molecular systems, in particular, are readily programmable via the sequence-dependent interaction between DNA strands, which has been exploited to design self-assembling dynamic DNA devices9, neural network-like molecular computation10, coupled regulatory circuits11, and schemes for constructing molecular-scale cellular automata (CA)12. Here, we use minimal models to study the concept of programmable pattern formation using theoretical and computational tools. While the intention is not to model any particular system, DNA-based implementations of the model are an interesting perspective (see ‘Discussion’).
The framework of CA13,14,15 provides a suitable basis for minimal models of programmable pattern formation. A cellular automaton consists of discrete subunits, which have at least two distinguishable internal states. The subunits communicate with their neighbors to determine the update of their internal state. The dynamics of a subunit is thus governed by update ‘rules’ that depend on its state, as well as on the state of its neighbors. For our analysis, a useful feature of CA is that the number of possible update rules is finite—each rule is a different scheme for local information processing—and there are no additional model parameters. The CA framework is sufficiently flexible to describe a broad range of pattern formation processes that do not depend on long-range signaling between cells14,16. Furthermore, CA are not solely abstract computational models, but can faithfully describe the dynamics of real systems, also in developmental biology17. CA models typically assume synchronous updates of all cells, which however do not need to occur at constant time intervals in real time, e.g., to describe developmental dynamics that progressively slows down18. Synchronicity is also not strictly required, since the same dynamics can be obtained with an asynchronous CA system featuring subunits capable of local synchronization via interactions with their neighbors19. In molecular systems, synchronization can arise from a collectively produced long-range signal, or from a local coupling between oscillators11.
Within the conceptual models that we consider, organizer cells can emit time-dependent signals into their neighborhood, affecting the pattern formation process of the remaining ‘bulk cells’. Programmability of pattern formation then refers to the ability of the organizer cells to reproducibly steer the system toward different target patterns, using different signaling sequences. We define a given model to be completely programmable, if organizers can direct the system to all different target patterns from any initial pattern. We consider programmability of pattern formation to be a desirable property, since it is reminiscent of the ability of developmental systems to work with only a small number of signaling systems, which are highly homologous between morphologically very different animals4. Note that while some CA, including the paradigmatic ‘Game of Life’ introduced by Conway20, are well known to be universal computing devices, the question of programmable pattern formation is distinct from universal computing: In the context of computing, both the ‘program’ and the ‘input data’ are specified by the initial state of the CA. In contrast, for programmable pattern formation the initial state is arbitrary, while the target pattern is encoded in the state transitions of the organizer cells, and the patterning algorithm is specified by the update rule and the topology of the system.
Here we show how the dynamics of the bulk cells, as specified by their update rule, affects the programmability of pattern formation. For the minimal system with two-state cells, only ten update rules enable complete programmability. However, the number of such ‘programmable rules’ increases strongly with the number of internal states. Patterning errors, incurred by cells that do not always follow their update rule, can be strongly reduced by an error correction scheme. Furthermore, we find that the robustness against the timing of organizer inputs can be increased at the expense of a reduction in the extent of programmability if organizer cells are also able to induce changes in the update rules of bulk cells.
Results
A minimal model for programmable pattern formation controlled by organizer cells
To explore the programmability of global pattern formation from local sites, we combine concepts from control theory with a class of models for pattern formation. Multiple modeling frameworks for pattern formation processes are available, which treat time, space, and patterning state either as discrete or continuum quantities, and differ also with respect to the level of detail of the description16. Here, we choose the most coarse-grained level of description, known as CA models. Within this framework, a system consists of localized subunits referred to as ‘cells’. The patterning state of cell i at time t is denoted by \(x_i^t\), which can only take on a finite number k of different values, \(x_i^t \in \{ 0, \ldots ,k - 1\}\). In the simplest case of elementary CA, there are only two different states, \(x_i^t = 0\) or 1 and the cells are arranged in one dimension (1D). The dynamics of a CA model is governed by local rules specifying how the state of a cell is updated depending on the state of the cell itself and the states of the surrounding cells, see Fig. 1. The same update rule is applied synchronously to all cells of the CA at once. For elementary CA, only the immediate neighbors of a cell affect its update, via an update rule of the form \(x_i^{t + 1} = f\left( {x_{i - 1}^t,x_i^t,x_{i + 1}^t} \right)\). Depending on the update rule, patterning information emerging from a localized source can propagate through the system to affect the global patterning process.
a The model features bulk cells (spheres) and organizer cells (marked by red arrow). The state of cells is represented by their color. Fixed boundary cells (boxes) can also be regarded as organizer cells that never change their state. b We consider three types of cell arrangement: Linear, with either one or two organizer cells, and circular. c The patterning dynamics of bulk cells follows a cellular automaton rule. The time-dependent state \(x_{i}(t)\) of each of the L bulk cells is updated according to \(x_{i}(t+1)=f(x_{i-1}(t), \, x_{i}(t),\, x_{i+1}(t))\), with a rule f that maps every triple of input states \((x_{i-1}(t), \, x_{i}(t),\, x_{i+1}(t))\) to an output state \(x_{i}(t+1)\), as specified by a transition table (small spheres). The rules are enumerated by translating the pattern of output states, ordered by the descending binary equivalent of the input states, into a binary number (here: 010101102, i.e., rule 86). The patterning process is controlled by the local dynamics of the organizer cells, which supply the patterning input. Programmable pattern formation in a cell arrangement with a given CA update rule refers to the ability of the organizer cell(s) to reproducibly steer the bulk cells to different target patterns within T time steps, using appropriate sequences of signals.
Our model systems consist of ‘bulk cells’ and ‘organizer cells’ (Fig. 1a). Bulk cells simply follow their update rules, taking input signals from their neighbor(s), regardless of whether these are other bulk cells or organizer cells. Organizer cells do not take inputs but exert control on the pattern formation process by changing their internal state according to a specified protocol, which depends on the target pattern. We do not specify the origin of this protocol—it could be the result from an internal developmental program or could be the result of external manipulation. Since both natural and engineered patterning systems come in a variety of topologies, and the topology may affect the pattern formation process and its control, we consider two different topologies, linear and circular (Fig. 1b). In a circular topology, a single organizer cell is embedded in a ring of L bulk cells, while a linear array of bulk cells can be controlled by an organizer cell at one or both ends, or at any position within the array. We collectively denote the time-dependent patterning state of all bulk cells as \(X^t\), and the time-dependent state of the organizer cells as \(O^t\). The global patterning dynamics then follows \(X^{t+1}=F(X^{t},\,O^{t})\) with a global update function F.
We consider a patterning system of this type to be completely programmable, if (i) the organizer cells can steer any initial pattern \(X^0\) toward any desired target pattern Y in a finite time with a suitable time-dependent organizer input \(O^t\), and (ii) the time needed to reach the target pattern scales at most polynomially with the system size L. For instance, if the time to reach the target pattern increases linearly with L, we say that the system is completely programmable in linear time. In contrast, if the system would randomly generate different patterns, the expected time to produce a specified target pattern would increase exponentially with L. If complete programmability is not obtainable, we also consider partial programmability, where only a subset of target patterns is reachable. In either case, the sequence of organizer inputs \(O^t\), may depend on the initial pattern \(X^0\). We first focus on the question of whether target patterns can be reached and will then consider the issue of stabilizing target patterns.
Some update rules enable complete programmability
The discrete model facilitates an efficient computational test for complete programmability, which relies on the representation of the patterning dynamics by a directed graph (Fig. 2a–c). In this ‘patterning graph’, each patterning state of the system is represented by a node. A connecting arrow from node X to node Y indicates that an input signal B exists that takes the system from state X to state Y in one step, i.e., \(Y=F(X,B)\). The arrow is labeled with this input signal B (if multiple signals exist, the arrow gets multiple labels). Complete programmability is then equivalent to the statement that there is a path from every node to every other node in the patterning graph, i.e., that the graph is strongly connected21 (Fig. 2). Strong connectivity of directed graphs with tens of thousands of nodes can be rapidly tested with standard algorithms (see ‘Methods’).
a Illustration of the patterning dynamics of a small system \(L=3\), steered from initial pattern 101 to target pattern 011 (last row) in a linear topology with two organizer cells (cells with red arrows) under rule 86 (cf. Fig. 1). (b) Part of the corresponding patterning graph, with nodes representing patterns and arrows possible transitions. The organizer inputs that can trigger a transition are indicated next to the corresponding arrow, e.g., \([0,0]\) when the left and right organizer cell both supply a ‘0’ signal (sometimes multiple input combinations are possible). The path corresponding to (a) is highlighted in red. c A system with a given update rule permits programmable pattern formation if the corresponding full patterning graph is strongly connected, i.e., there is a path from every node to every other node (regardless of the required inputs). d The number of distinct programmable rules decreases monotonically with the system size and reaches a plateau-value (10 for linear topology, 7 for circular topology). e The update rules that remain programmable for all system sizes with circular (first 7) and linear topology (all shown rules), listed in their algebraic form with \(x_i^t\) denoting the state x at position i at time t, decimal rule number, and update map. Different shades indicate properties of the rules as indicated in the legend. f Statistical characterization of patterning graphs by their in-degree distribution (linear systems with two organizer cells, \(L=16\) bulk cells, and three different exemplary rules; distributions shown up to in-degree 20). All programmable rules have the same distribution as the shown rule 240, whereas the distribution of non-programmable rules is broad (shown examples: rule 233 and 58), except for the identity and complement rule (distributions for all rules shown in Fig. S2). For comparison, the in-degree distribution of a randomized graph is also shown (random redirection of arrows to any node with equal probability).
We applied this test in systems with different sizes and topologies, for the minimal model of \(K=2\) states, where the number of possible update functions f is only \(2^{\left( {2^3} \right)} = 256\). Given that our underlying model is symmetric with respect to the spatial directions ‘left’ and ‘right’, and also with respect to the internal states ‘0’ and ‘1’, the set of 256 rules can be split into 88 equivalence classes, which we refer to as ‘distinct rules’. We tested all rules and found that there are several which enable complete programmability of pattern formation. The number of such rules depends on the topology and decreases with the system size L (Fig. 2d). Strikingly, however, for systems of size \(L\geq9\) the numbers no longer decrease, and the set of remaining rules is unchanged up to the maximum size that we were able to test numerically. This observation suggests that a subset of ‘programmable rules’ enables complete programmability of pattern formation for systems with a fixed topology but any size. This subset contains ten distinct rules for linear topology and seven distinct rules for circular topology (Fig. 2d, e). In the linear topology, the optimal placement of organizer cell(s) with respect to the number of programmable rules is at the boundaries (Supplementary Notes S1.1 and Fig. S1). The ten distinct rules for linear topology were also independently22 identified by another study23,24.
While complete programmability, as defined above, is a global property of the patterning graph (strong connectivity), the ten rules of Fig. 2e also stand out in a local property of the patterning graph, the in-degree distribution. In a directed graph, the in-degree of a node corresponds to the number of incoming arrows. Figure 2f shows histograms of the in-degrees of all nodes in the patterning graphs of one programmable and two non-programmable rules. For comparison, Fig. 2f also shows the in-degree distribution of a randomized graph, in which the outgoing links from each node are randomly reassigned to any target node. These examples illustrate an empirical property (Fig. S2 and Supplementary Notes S2): Whereas programmable rules have an in-degree distribution with only a single peak at in-degree two (for one organizer cell) or four (for two organizer cells), non-programmable rules display a broad in-degree distribution, which depends on the specific rule and differs from the distribution for a randomized graph. This indicates that one can distinguish programmable from non-programmable rules already based on the local structural statistics of the patterning graph, which for large systems could also be sampled from a randomly chosen subset of nodes. We will see further below that the number of programmable rules increases rapidly with the number k of cell states.
Different mechanisms for complete programmability of patterning
To illustrate the mechanisms by which complete programmability is achieved, Fig. 3 displays the spatio-temporal dynamics for several rules, topologies, and target patterns. The simplest mechanism is that of Fig. 3a, where a single organizer cell feeds its changing state into a linear array of bulk cells, in this case from the left edge. The bulk cells propagate the received information to their neighbors, effectively writing a temporal signal into a spatial pattern. This mechanism is reminiscent of the clock-and-wavefront mechanism in vertebrate somitogenesis, which relies on a temporal oscillation that is converted into a spatial stripe pattern25,26,27,28.
Exemplary kymographs show the patterning dynamics of different systems toward a target pattern (last line in temporal direction), each with \(L=8\) bulk cells, but different cell arrangements and update rules: a linear system with one organizer cell (red arrow) on the left, fixed cell (box) on the right and rule 240, b rule 15, and c rule 105, d linear system with two organizer cells with rule 90, e two-sided control with rule 30 and a different target pattern, f circular system with embedded organizer cell and rule 240, g rule 30 starting from a non-homogeneous initial state, and h from a homogeneous initial state takes more time to reach the target pattern. i Characterization of the patterning dynamics by the entropy-like observable \(S(t)\), a logarithmic measure of the number of different patterns that remain after t update steps, if the patterning process is started from the ensemble of all possible initial states (see main text). The data show the time-dependence of \(S(t)\) for different update rules (symbols) in a circular topology with the same target pattern as in (f). The dashed line marks \(S=-t+L\) for comparison. Circles indicate rule insensitive to the initial condition, triangles nonlinear rules, squares both sides bijective rules and stars the remaining rule (cp. Fig. 2e). j As in (i), but for linear topology with two organizers. The solid line additionally marks \(S=-2t+L\) for comparison. The behavior shown for this particular target pattern is generic, as can be seen from Figs. S3–S5, which show the minimum, maximum and average \(S(t)\) over all patterns.
The examples in Fig. 3b–e display more complex behavior, suggesting alternative modes of programmable pattern formation. Figure 3b displays a variant of the mechanism in Fig. 3a, which updates a cell to the inverted state of its left neighbor. Thereby, this rule produces a dynamics where the pattern oscillates as it is pushed from the organizer cell into the bulk. In both cases, the initial state of the system is completely erased during the patterning process, such that the sequence \(O^t\) of organizer inputs is independent of the initial pattern \(X^0\). In contrast, a third rule (Fig. 3c) generates the same target pattern partially from the initial state, exploiting the computational power of the update rule. As a consequence, the target pattern is reached more rapidly. Figure 3d, e illustrate programmable pattern formation with simultaneous input from two organizer cells. Only update rules that are affected by input signals from both the left and the right side can simultaneously process information from two organizer cells. In the case of Fig. 3d, rule 90 produces the target pattern by symmetrically using information from both organizer cells, while Fig. 3e illustrates an asymmetric pattern formation process with rule 30, where information from the left side is preferentially used. Finally, Fig. 3f–h displays kymographs for cases where a single organizer cell is embedded in a ring of bulk cells. In Fig. 3f, the information from the organizer cell is pushed only in one direction, such that the patterning process is analogous to that of Fig. 3a for the linear cell array. In contrast, Fig. 3g illustrates a case where the target pattern is computed (by rule 30) from the initial pattern. The most complex example is that of Fig. 3h, where rule 30 propagates information from the organizer cell to both sides, producing an ‘interference’ phenomenon when the two signals meet, which results in a much longer time required to reach the target pattern.
The unifying principle underlying these examples is linear transport of patterning information from one or multiple sources, with concurrent processing of this information by the bulk cells. The behavior is visually simple only if the bulk cells merely pass on the information they receive, while additional signal integration with their internal states typically generates complex spatio-temporal dynamics.
Time to reach the target pattern
The examples in Fig. 3 suggest that systems with programmable rules are completely programmable in linear time. With simple unidirectional transport of patterning information (as in Fig. 3a, b, f), the maximum number of update steps in an optimal path is equal to the number L of bulk cells. With two organizer cells, this maximum can be cut in half, as seen in Fig. 3d. Furthermore, in some cases the update rule can construct a portion of the target pattern from the initial pattern to speed up the pattern formation process (Fig. 3c, g).
We obtain a global view of the patterning dynamics by considering the ensemble of all possible initial states of the system and monitoring how this ensemble progressively shrinks toward a single point in state space (the target pattern). A convenient observable to characterize these dynamics is the time-dependent ‘entropy’ \(S(t)={\mathrm{log}}_{2}({\it{\Omega}}(t))\), where \({\it{\Omega}}(t)\) denotes the number of points in state space occupied by the ensemble at time t. As the patterning process proceeds from every possible initial state along every possible shortest path to the target state \(S(t)\), decreases from L to zero. The computed time traces S(t) for a system of size \(L=8\) with the different programmable rules are shown in Fig. 3i (circular topology) and Fig. 3j (linear topology with organizer cells at both ends). In the latter case, \(S(t)\) decreases roughly linearly for all rules, corresponding to an exponentially shrinking volume of the pattern ensemble in state space. The velocity of this ‘entropy reduction’ is either \(\frac{{\Delta S}}{{\Delta t}} = - 1\) or \(-2\) (dashed and solid line, respectively). In the circular topology, \(S(t)\) either decreases linearly with slope −1, or displays a slower decrease with variable slope. Taken together, the dynamics of \(S(t)\) is consistent with the spectrum of behaviors observed in the examples of Fig. 3a–h. It is also consistent with the behavior of the average shortest path length in the patterning graph (Supplementary Notes S3).
Conservation principle and programmable rules for k-state systems
Intuitively it is clear that faithful transport of patterning information from organizer cells into the bulk requires a conservation principle. This notion is formalized by the concept of bijectivity. We define a rule f to be left-bijective, if the mapping \(x\rightarrow{y}\) with \(y = f\left( {x,x_i^t,x_{i + 1}^t} \right)\) is bijective for each combination of \(x_i^t\), \(x_{i + 1}^t\) values (Fig. 4a, b). For a left-bijective rule every possible output \(x_i^{t + 1}\) can be reached by choosing an appropriate left input \(x_{i - 1}^t\), irrespective of \(x_i^t\) and \(x_{i + 1}^t\). This property suffices to guarantee that one can find a series of inputs \(O^{t}\) from an organizer cell on the left to produce any target pattern in the bulk cell array (Fig. 4c and Supplementary Notes S1.2, S1.3). Similarly, if an update rule is right-bijective, it permits complete programmability of pattern formation from an organizer cell on the right. The argument of Fig. 4c is constructive in the sense that it not only guarantees the existence of a suitable organizer sequence \(O^{t}\) to reach the target pattern, but it provides a recipe to explicitly construct \(O^{t}\), given the update rule as well as the initial and the target pattern (Supplementary Notes S1.4). This recipe confirms the distinction between the simple rules of Fig. 3a, b and the other programmable rules with more complex behavior: For the simple rules, the organizer sequences \(O^{t}\) can be chosen independent of the initial state of the system, whereas for the complex rules, the construction of \(O^{t}\) requires knowledge of the initial state (see the classification of rules in Fig. 2e). Rules that are both left- and right-bijective can faithfully transport information from both sides, which can speed up the patterning process with two organizer cells, as seen in Fig. 3d, e. However, in the circular topology, rules that are both left- and right-bijective do not enable complete programmability, since they are unable to convert initially symmetric patterns into an asymmetric one (Supplementary Notes S1.5).
a Illustration of a left-bijective rule (black box) for an elementary two-state cellular automaton. The eight possible input configurations are grouped into four pairs (blue boxes) according to the states of the middle and right-hand cells (dark boundaries) of the rule inputs (light blue shading). For each of these pairs, the rule establishes a one-to-one mapping between the state of its left input cell (light boundary) and its output (dark blue background). Figure S6 illustrates the bijectivity of all programmable rules in a similar way. b In contrast, a rule that maps at least one pair of input states onto the same output state is not left-bijective (first and second pair in the shown example). c Illustration of the construction scheme for the organizer sequence \(O(t)\) that steers an initial pattern to a desired target pattern. In this example, the system has \(L=5\) bulk cells and an organizer cell on the left (red boundary). The construction scheme determines the organizer sequence by backward propagation from the target pattern, and explicitly demonstrates that bijectivity implies programmability. All white cells are not influenced by the organizer sequence \(O(t)\), so their states can be computed from the initial pattern with the update rule. Back propagation then begins by setting the value of the rightmost cell in the final pattern to the desired value \(x_{4}(5)\). Since \(x_{4}(4)\) and \(x_{5}(4)\) are known, left-bijectivity guarantees that there is a value \(x_{3}(4)\) such that \(x_{4}(5)\) has the desired value. Similarly, it is possible to set \(x_{2}(3)\) such that \(x_{3}(4)\) takes on the required value determined in the previous step. Iterating along the diagonal with the lightest blue shade then fixes the first organizer input, \(O(0)\). Back propagation of \(x_{3}(5)\) (second lightest blue shade) then fixes \(O(1)\) and so forth (each back propagation indicated by a different blue shade). Thus, bijectivity of the update rule suffices to construct an organizer sequence \(O(t)\) to steer a given initial pattern into any desired target pattern.
Importantly, the argument of Fig. 4c is valid for any length L of the system, for any number k of cell states, and it can also be generalized to obtain a constructive recipe for update rules that depend on larger neighborhoods of cells (Supplementary Notes S1.6). The bijectivity property can be used to show that with k cell states there are at least \(k!^{k^2}\) programmable rules before taking into account symmetries, but that the fraction of bijective rules within all rules decreases rapidly with k (Supplementary Notes S1.7). Furthermore, it follows (Supplementary Notes S1.8) that the maximal length of the shortest path from a given initial pattern to a desired target pattern is L in the case of one organizer cell, and \(\frac{L}{2}\) for two organizer cells and both left- and right-bijective rules (or \(\frac{{L + 1}}{2}\) when \(\frac{L}{2}\) is not an integer), as we had empirically seen above.
Robustness against errors and error correction
In real systems, the communication between subunits, as well as the information processing within subunits, are exposed to noise, causing some level of stochasticity in the dynamics. How sensitive programmable pattern formation is to such stochasticity is therefore a crucial question. To explore this question, we introduce an error process into the model. After executing the deterministic update rule, each cell stochastically switches to the opposite state with probability p, or remains in its state with probability \(1 - p\), independent of the state of its neighbors. This stochastic update models effects such as loss of memory (of the prior cell state), noise in the internal regulatory circuit that encodes the update rule, unreliable signal transmission from neighboring cells, and noise in the exact timing of state transitions. Since we consider a spatially and temporally homogenous system, we take p to be constant in time and space.
To monitor the impact of the stochasticity, we measure the reliability of the pattern formation process as a function of p. Specifically, we determine the probability that the final pattern has no error, i.e., that it matches the desired target pattern (see ‘Methods’). For programmable rules that take input from only one neighboring cell, this probability can be estimated as
since a system of size L reaches its target state after at most L steps, and errors in \(L(L-1)/2\) individual cell updates can have an influence on the final pattern (see Fig. 4). This error estimate helps to interpret our simulations of the model (Fig. 5). The reliability as a function of the error rate p in a system of fixed size is shown in Fig. 5c for all programmable rules (red symbols), confirming that the estimate (solid red line) captures the essential behavior of the model. In particular, the reliability decreases linearly with p for small error rates (dashed red line), implying that all patterning schemes considered so far are very sensitive to errors.
a To explore a mechanism for error correction, we consider a cylindrical system with an array of organizer cells (white cells with red border) along the left edge and a fixed boundary condition (white squares) along the right edge. Cells are indexed by i in horizontal direction and j in vertical direction. The update rule now takes input from a nine-cell neighborhood (shaded blue). b Given a nine-cell neighborhood, the update rule applies ‘majority voting’ in the vertical direction (index j), establishing a consensus triplet to which one of the 1D programmable rules f is applied, yielding the final output. c Computer simulations (symbols) and analytical theory (lines) for systems with error correction by majority voting (blue) compared to the case without error correction (red). Circles indicate rules insensitive to the initial condition, triangles nonlinear rules, squares both sides bijective rules and stars the remaining rule (cp. Fig. 2e). In each case, a system of horizontal and vertical size of \(L=K=9\) was used. After each update cells switch to a different state with probability p. Sufficient simulation runs were performed to estimate the plotted probability of arriving at the correct final pattern with a statistical error smaller than the symbol size (see ‘Methods’). Convergence is demonstrated by the observation that rules for which the same error behavior is expected (Rules 15 and 240) yield data points lying on top of each other. See Supplementary Note S4 for the analytical approximations and Supplementary Fig. S14 for a corresponding analysis of a planar system.
The root cause of the high sensitivity to errors is the one-dimensional geometry of our model systems: A single failure breaks the ‘chain of command’ from the organizer cells to the distant bulk cells. Given that most real systems have two- or three-dimensional arrangements of subunits, it is natural to extend the spatial dimension of our model. We focus on a two-dimensional extension of our model, in which K parallel cell lanes, each of length L, are connected to form a tube (see Fig. 5a, where periodic boundary conditions are applied in the vertical direction). The parallel lanes offer redundancy, which the cells can leverage to increase the reliability: They communicate with their lateral neighbors and apply a majority voting rule for their update (Fig. 5b), which in tissues could be mediated by diffusible signaling molecules. Along the axis of the tube cells follow the same rules as in the one-dimensional model above. The blue symbols in Fig. 5c show the reliability as a function of p for a tube with the same length L as the one-dimensional system (see caption for parameters and methods section for the numerical procedure). We observe a dramatic increase in reliability, caused by the ability of the lateral majority voting rule to correct isolated errors. The error correction changes the scaling of the reliability with p from linear to quadratic (dashed blue line). In fact, the observed behavior can be captured by an estimate (solid blue line) based on counting the number of arrangements of errors that cannot be corrected (Supplementary Notes S4.1). The expansion of this estimate for small p shows that
for the tube. The rules which only shift the state of the cell to the next cell perform best since they spread errors the least (Supplementary Notes S4.2).
Robustness against variable timing of organizer signals
The above analysis showed that local organizers can steer the bulk cells into any one-dimensional target pattern using only local signals processed according to simple rules. However, this requires precise timing in the switching of the organizer signals. Precise timing is also needed for the arrest of the patterning process when the target pattern is reached, because the target pattern is generally not a fixed point of the dynamics (programmable rules have only trivial stationary patterns). To explore the degree of programmability that can be achieved with less precise timing, we consider an alternative scheme, which uses update rules with nontrivial stationary patterns: For each organizer input, we let the system evolve until the pattern no longer changes before applying a new input. Together with each input, we also allow a global change of the update rule (same for all cells). In a developmental system, this would amount to a change in the interpretation of intercellular signals in different developmental stages, which is a known phenomenon, e.g., for the Toll signaling pathway of Drosophila1. The change could be triggered by a global signal, which does not need to be timed precisely, since the system runs into a stationary pattern at which it can stay for an extended time. Global changes of the update rule could in principle also be implemented in a synthetic DNA-based system (Supplementary Notes S5).
For simplicity, we refer to the combination of an input with an update rule as an ‘instruction’. We only use instructions that lead the system to a stationary state, avoiding those that lead to limit cycles. To construct an efficient search method for a protocol that steers the system from a given initial pattern to a desired target pattern, we first analyze the patterning graphs of all CA rules. For each rule and organizer input, we identify all attractors and their basins of attraction, which consist of all patterns from which the attractor is reachable (Fig. 6a). We then construct a single ‘attractor graph’ from all basins of attraction, by adding a directed link \(X\rightarrow{Y}\) for each pattern X in the attraction basin of pattern Y (Fig. 6b). Each link has an associated instruction. Using the attractor graph, we determine the instruction sequence by extracting the shortest path connecting two patterns. This recipe minimizes the number of instructions, but other objective functions, such as minimizing the number of changes in the rule or the total time needed to reach the final attractor, could be implemented with similar methods. Each instruction of the instruction sequence is then applied sufficiently long to reach the respective steady state, ultimately steering the system from the initial pattern to the desired target pattern (Fig. 6c).
To explore a mechanism for programmable pattern formation that is robust against variable timing of organizer signals, we consider an alternative scheme for programmable pattern formation, which uses all update rules with nontrivial stationary patterns (see main text). a Example of a patterning graph of rule 40 with \(L=3\) cells in a linear topology with two organizer cells. In dark red a sample path is shown leading from pattern ‘101’ to the fixed point ‘001’ using the instruction (rule 40, \([1, 1]\)) – i.e., left and right organizer cell have both state 1. The light red shaded areas show the attraction basins of the instruction (rule 40, \([1, 1]\)) with the attractors ‘000’ and ‘001’. Using also the other possible instructions with rule 40 all configurations are in the attraction basin of ‘000’, while only the right shaded subset is in the attraction basin of ‘001’. b The contributions of rule 40 with all possible inputs to the attractor graph are calculated by adding a directed edge toward a node if the pattern corresponding to the origin of the arrow is in the attraction basin of the target node. The red arrow corresponds to the red path on the left. The other contributions shown are from rule 86 depicted in Fig. 2. All contributions from all rules generate the attractor graph. c Example of spatio-temporal evolution for \(L=10\) with organizer cells on both sides. The target pattern is reached with 5 instructions. After each instruction, the system is allowed to reach its steady state which may last as long as desired (dots in timeline). d Number of reachable patterns as a function of L for a linear system with one (light blue squares) and two (dark blue circles) organizer cells and a circular system (red triangles) compared to the total number of patterns \(2^{L}\). The number of reachable patterns is determined by calculating how many nodes (patterns) of the attractor graph can be reached on a directed path from the 0 pattern. Dashed lines show a fit of the form \(a^{L}\) to the respectively colored data. e Average number of instructions necessary to generate reachable patterns with colors as defined in (d), with standard deviation as error bars, as a function of L, with a linear fit (dashed lines) in the range \(L\!\in\) [8, 16] to exclude finite-size effects.
We only consider the homogeneous initial condition (all cells in state ‘0’) with no prior spatial information that could seed the generation of patterns. Not all patterns can be reached by this scheme for larger system sizes (Supplementary Notes S6.1). We performed an exhaustive analysis to determine the reachable patterns for grid sizes up to length \(L=16\). We fit the resulting data in the saturated range \(L\!\in\) [8, 16] and determined the number of reachable patterns to scale as \(1.89^L\) (circular and linear topology with two organizer cells) and \(1.82^L\) (linear topology with one organizer cell), showing that even if not all patterns are reachable, an exponentially growing number is (Fig. 6d). Interestingly, approximately the same scaling applies for the linear topology if we weaken the assumption that the CA can distinguish left from right, i.e., include only outer-totalistic rules in the attractor graph, which are agnostic to the directionality of the signals (Supplementary Notes S6.2). These empirical observations are consistent with the results of an analytical approach to determine the number of attractors of finite CA, which also indicates that the number of attractors for individual rules (except the identity rule 204) grows more slowly with L than the total number \(2^L\) of possible patterns29.
To characterize the patterning dynamics, we calculated the average shortest path length in the attractor graph, i.e., the average minimal number of required instructions, to reach the accessible target patterns as a function of the system size L. The empirically observed linear dependence (Fig. 6e) indicates that, even as the number of reachable patterns increases exponentially, the time, measured in number of instructions, to reach a target pattern increases only linearly with system size, as in our original scheme for programmable patterning (Fig. 3i, j).
Models for pattern formation processes can also be regarded as a means to compress the information required to specify a pattern. This notion is formalized by the concept of Kolmogorov complexity of a pattern, defined as the length of the shortest program for a Turing machine, which outputs that pattern and halts30. Within our scheme, we can say that the complexity of a pattern is measured by the number of instructions needed to generate it starting from the homogeneous initial condition. Empirically, the patterns which require the fewest instructions exhibit some periodicity, which makes them amenable to compression, while there is no obvious visible difference between the most complex reachable patterns and the unreachable patterns (Supplementary Notes S6.3, Fig. S15).
Discussion
Programmable pattern formation in cellular systems is a remarkable phenomenon in biology and a long-term goal for the design of synthetic multicellular systems31. Here, our objective was not to study any specific system, but to identify general schemes whereby local signals from organizer cells can direct global pattern formation. We chose a CA-based modeling framework, which is sufficiently general to encompass a broad class of model systems, yet simple enough for explorative studies. For elementary CA, in which cells have only two states and two neighbors, we performed an unbiased exhaustive analysis of all dynamical rules. We were then able to generalize some of our results to more complex systems. In particular, our approach led to the following findings: (i) complete programmability of pattern formation by isolated organizer cells is possible only with a small fraction of distinct rules (Fig. 2), which fulfill a conservation principle for the transmission of patterning information (Fig. 4). (ii) While the detailed patterning dynamics implemented by each programmable rule is different (Fig. 3), the unifying principle can be interpreted as a generalization of the ‘clock-and-wavefront’ scheme underlying vertebrate somitogenesis, where a temporal signal is converted into a spatial pattern. (iii) Global pattern formation controlled by isolated locally acting organizers is intrinsically susceptible to errors, but the accuracy of pattern formation can be substantially improved with a simple error-correcting scheme based on local majority voting (Fig. 5). (iv) Programmable pattern formation controlled by organizer cells is generally sensitive to the timing of organizer inputs, but robustness against variable timing is achievable with organizers that have the additional ability to change the update rule of bulk cells, i.e., the way in which bulk cells interpret the received signals (Fig. 6).
Our results contribute toward a conceptual framework for constructing molecular or cellular systems with the ability of programmable pattern formation. DNA-based systems form a promising platform for molecular realizations of programmable pattern formation due to their programmability and information processing ability32,33. An elementary CA with the programmable rule 90 has already been implemented with DNA tiles34,35 albeit not in a way that permits inputs from an organizer. Other DNA-based implementations, which are more complex but offer more flexibility, have also been proposed12. On the basis of these proposed designs, a biomolecular CA that allows for input signals controlling its patterning process could be implemented as described in Supplementary Notes S5.
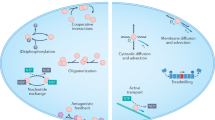
Synthetic cell-like systems with the capability to communicate and process information have also been implemented, based on emulsion droplets11 and liposomes36. Information processing within such synthetic cells is realized with artificial gene circuits, based on in vitro transcription or transcription-translation systems, whereas communication between neighboring cells is enabled, for instance, by dedicated protein pores31. A complementary path to achieve programmable pattern formation in cellular systems is to equip biological cells with engineered sensing and response systems37,38.
Given that our model was not designed to mimic any specific system, it is noteworthy that it led us to a principle of programmable pattern formation, which can be regarded as a generalization of the clock-and-wavefront scheme underlying vertebrate somitogenesis25,26,27. The basic principle is the same as that of a tape recorder, where a temporal audio signal is written into a spatial magnetic pattern. In the case of the clock-and-wavefront scheme, a periodic gene expression signal generates a stripe pattern via a determination front, which sweeps the tissue and arrests cells in their current state. The patterning dynamics displayed in Fig. 3 generalizes the clock-and-wavefront scheme by allowing for (i) any target pattern, not just regular stripes, and (ii) simultaneous transport and processing of patterning information. While our model does not implement a determination front, we considered an alternative scheme, in which the update rule of the bulk cells is also controlled by the organizers and the target pattern is stabilized dynamically. CA with changing update rules are interesting also from the computational perspective since they were previously found to display capabilities linked to the computational problem of open-ended evolution39.
The simplicity of our model was key to obtaining rigorous results, but also poses limitations. One important limitation is the restriction to discrete internal cell states. Pattern formation processes are typically described by nonlinear dynamical systems with multistable behavior, such that qualitatively distinct patterning states, e.g., gene expression ‘on’ or ‘off’, can emerge in spatially adjacent regions. Our model adopts a coarse-grained level of description, which already assumes the existence of such discrete states and ignores all intermediate states. For a biological system, discrete update rules represent logic-based models of a biochemical signaling network40. For other types of systems, discrete update rules typically also represent ‘digital’ approximations of the underlying ‘analog’ dynamics.
Another limitation is the one-dimensional arrangement of cells within our model, which permits only linear propagation of patterning information in space. This restriction is somewhat relaxed in our quasi-1D extension of the model (Fig. 5), where lateral signaling between cells is used for error correction. However, this extension does not address the more general question of programmable pattern formation in two or more dimensions, which remains open for future work.
We also did not include biological processes like cell growth, cell division, and death, but assumed that the patterning process occurs in a group of cells with a static arrangement, as for example in Caenorhabditis vulva development5. Finally, we limited our study to a patterning scenario based on organizer cells. However, we found that the dynamical update rules of the bulk cells can also generate parts of the target pattern (Fig. 3), and considered an alternative scheme for programmable pattern formation, which combines patterning information from local organizers with ‘distributed computation’ of patterning information (Fig. 6). Bulk cells with more states, or larger neighborhoods for the update rule, will have more computational ability and will therefore enhance the potential for programmable pattern formation via distributed computation. Indeed, it is well known that CA can serve as computing devices, with some even shown to be computationally universal15. In those cases, the initial state of the cellular automaton serves both as the program and the input data, while the update rule specifies the mechanism of the computer and the result of the computation is obtained from the state after time evolution. The situation is different for programmable pattern formation: In our ‘organizer scenario’, the initial state of the system can be simple, e.g., homogeneous, while the input data (patterning information) is supplied as a time-dependent local signal. Bijective update rules enable universal pattern formation with this scenario. Interestingly, these bijective rules were among the ‘illegal’ rules excluded in Wolfram’s pioneering study on the statistical mechanics of CA14, due to their violation of the quiescence and isotropy conditions.
Our question of programmability is closely related to the question of ‘controllability’ in the field of control theory. Control theory provides a general mathematical framework to analyze the control of dynamical systems41. It formalizes the intuitive notion of ‘controllability’ as the ability to steer a dynamical system to any desired state from any initial state by appropriate signals. A focus of recent research has been on the control of complex networks42,43, a broad class of dynamical systems ranging from networks of protein interactions44 and neurons45 to power grids46. Application of control theory concepts to linear dynamics on networks with complex topologies led to insights about the relation between network topology and the controllability of its dynamics42,43. Here, we focused instead on systems with simple topologies but more complex dynamics. Closely related control problems have independently22 been considered under the name of ‘boundary regional control’23,24,47. Several examples of CA systems were tested for regional controllability24, and the framework of Boolean derivatives was used to establish that a property named ‘peripheral linearity’, closely related to our left/right bijectivity, suffices to guarantee controllability23. The control problem can also be extended to probabilistic CA by demanding that the probability of reaching the desired target state is maximized, as shown for a specific example based on an extension of the Domany-Kinzel CA47.
In conclusion, programmable pattern formation connects the experimental fields of synthetic and systems biology to theoretical research on self-organization, computation, and control. We established simple scenarios for programmable pattern formation in cellular systems based on local organizers. Our results provide a rigorous basis for the analysis of more complex patterning scenarios, and for a conceptual framework to design synthetic molecular and cellular systems.
Methods
Computational test for programmability of an update rule
In order to test an update rule for programmability, we first construct the patterning graph. For a given rule and boundary states B, we perform one update step for a pattern X, which takes the system to pattern Y. This corresponds to an arrow in the patterning graph from node X to node Y. We now construct the patterning graph by creating a graph in the ‘python-igraph’ library48 and inserting directed links between all nodes which are connected by one step in the dynamics. Having constructed the patterning graph, the igraph library provides a direct test if the graph is strongly connected. For exemplary cases, we also checked the results by performing a similar procedure with the computationally less efficient networkx library49.
Numerical simulations with errors
For a given update rule and 1D initial and target patterns of length L, the shortest path and its corresponding organizer sequence are calculated. The deterministic update on the 2D grid (with given vertical size K) is obtained by combining the majority voting rule in the vertical direction with the 1D update rule in the horizontal direction (Fig. 5b). After each deterministic update, each cell is stochastically switched to the opposite state with a probability p, as described above. The final pattern is compared with the desired target pattern and, if they coincide, counted as having no error in the final pattern. This procedure is carried out Ntrial times for each possible combination of initial and target pattern. The probability for an error in the final pattern, which depends on the rule R, the grid dimensions L and K, and the error probability p, is then approximated as
where Ntrial = 5 is already large enough to achieve convergence.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The code that supports the findings of this study is available from the corresponding author upon reasonable request.
References
Wolpert, L. Principles of Development. 5th edn. (Oxford University Press, Oxford, 2015).
Prud’homme, B., Gompel, N. & Carroll, S. B. Emerging principles of regulatory evolution. PNAS 104 Suppl 1, 8605–8612 (2007).
Robertis, E. M. de. Spemann’s organizer and self-regulation in amphibian embryos. Nat. Rev. Mol. Cell Biol. 7, 296–302 (2006).
Perrimon, N., Pitsouli, C. & Shilo, B.-Z. Signaling mechanisms controlling cell fate and embryonic patterning. Cold Spring Harb. Perspect. Biol. 4, a005975 (2012).
Hoyos, E. et al. Quantitative variation in autocrine signaling and pathway crosstalk in the Caenorhabditis vulval network. Curr. Biol. 21, 527–538 (2011).
Rubenstein, M., Cornejo, A. & Nagpal, R. Programmable self-assembly in a thousand-robot swarm. Science 345, 795–799 (2014).
Slavkov, I. et al. Morphogenesis in robot swarms. Sci. Robot. 3, eaau9178 (2018).
Farkas, I., Helbing, D. & Vicsek, T. Mexican waves in an excitable medium. Nature 419, 131–132 (2002).
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Yin, P., Sahu, S., Turberfield, A. J. & Reif, J. H. Design of autonomous DNA cellular automata. In DNA computing. 11th International Workshop on DNA Computing. DNA11, London, ON, Canada, June 6–9, 2005; revised selected papers, Vol 3892 (eds. Carbone, A. & Pierce, N. A.) 399–416 (Springer, Berlin, 2006).
Ulam, S. On some mathematical problems connected with patterns of growth of figures. In Proceedings of Symposium in Applied Mathematics, 215–224 (1962).
Wolfram, S. Statistical mechanics of cellular automata. Rev. Mod. Phys. 55, 601–644 (1983).
Wolfram, S. Computation theory of cellular automata. Commun. Math. Phys. 96, 15–57 (1984).
Deutsch, A. & Dormann, S. Cellular Automaton Modeling of Biological Pattern Formation. Characterization, Applications, and Analysis (Birkhäuser Boston, Boston, MA, 2005).
Manukyan, L., Montandon, S. A., Fofonjka, A., Smirnov, S. & Milinkovitch, M. C. A living mesoscopic cellular automaton made of skin scales. Nature 544, 173–179 (2017).
Padgett, J. & Santos, S. D. M. From clocks to dominoes: lessons on cell cycle remodelling from embryonic stem cells. FEBS Lett. https://doi.org/10.1002/1873-3468.13862 (2020).
Nehaniv, C. L. Asynchronous automata networks can emulate any synchronous automata network. Int. J. Algebra Comput. 14, 719–739 (2004).
Gardner, M. Mathematical games. The fantastic combinations of John Conway’s new solitaire game of “life”. Sci. Am. 223, 120–123 (1970).
Newman, M. E. J. Networks. An introduction (Oxford Univ. Press, Oxford, 2010).
Ramalho, T. Information Processing in Biology: A Study on Signaling and Emergent Computation. Dissertation (Ludwig-Maximilians-Universität, 2015).
Bagnoli, F., El Yacoubi, S. & Rechtman, R. Toward a boundary regional control problem for Boolean cellular automata. Nat. Comput. 17, 479–486 (2018).
Dridi, S., El Yacoubi, S., Bagnoli, F. & Fontaine, A. A graph theory approach for regional controllability of Boolean cellular automata. Int J. Parallel Emergent Distrib. Syst. 35, 499–513 (2020).
Oates, A. C., Morelli, L. G. & Ares, S. Patterning embryos with oscillations: structure, function and dynamics of the vertebrate segmentation clock. Development 139, 625–639 (2012).
Cooke, J. & Zeeman, E. C. A clock and wavefront model for control of the number of repeated structures during animal morphogenesis. J. Theor. Biol. 58, 455–476 (1976).
Hubaud, A. & Pourquié, O. Signalling dynamics in vertebrate segmentation. Nat. Rev. Mol. Cell Biol. 15, 709–721 (2014).
Naoki, H. et al. Noise-resistant developmental reproducibility in vertebrate somite formation. PLoS Comput. Biol. 15, e1006579 (2019).
Twining, C. J. & Binder, P.-M. Enumeration of limit cycles in noncylindrical cellular automata. J. Stat. Phys. 66, 385–401 (1992).
Cover, T. M. & Thomas, J. A. Elements of Information Theory. 2nd edn. (Wiley-Interscience, Hoboken, NJ, 2006).
Dupin, A. & Simmel, F. C. Signalling and differentiation in emulsion-based multi-compartmentalized in vitro gene circuits. Nat. Chem. 11, 32–39 (2019).
Chatterjee, G., Dalchau, N., Muscat, R. A., Phillips, A. & Seelig, G. A spatially localized architecture for fast and modular DNA computing. Nat. Nanotechnol. 12, 920–927 (2017).
Zadorin, A. S. et al. Synthesis and materialization of a reaction-diffusion French flag pattern. Nat. Chem. 9, 990–996 (2017).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol. 2, e424 (2004).
Barish, R. D., Schulman, R., Rothemund, P. W. K. & Winfree, E. An information-bearing seed for nucleating algorithmic self-assembly. PNAS 106, 6054–6059 (2009).
Adamala, K. P., Martin-Alarcon, D. A., Guthrie-Honea, K. R. & Boyden, E. S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 9, 431–439 (2017).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell 164, 780–791 (2016).
Toda, S., Blauch, L. R., Tang, S. K. Y., Morsut, L. & Lim, W. A. Programming self-organizing multicellular structures with synthetic cell-cell signaling. Science 361, 156–162 (2018).
Adams, A., Zenil, H., Davies, P. C. W. & Walker, S. I. Formal definitions of unbounded evolution and innovation reveal universal mechanisms for open-ended evolution in dynamical systems. Sci. Rep. 7, 997 (2017).
Morris, M. K., Saez-Rodriguez, J., Sorger, P. K. & Lauffenburger, D. A. Logic-based models for the analysis of cell signaling networks. Biochemistry 49, 3216–3224 (2010).
Schulz, M. (ed.). Control Theory in Physics and Other Fields of Science. Concepts, Tools, and Applications. (Springer-Verlag, Berlin Heidelberg, 2006).
Liu, Y.-Y., Slotine, J.-J. & Barabasi, A.-L. Controllability of complex networks. Nature 473, 167–173 (2011).
Liu, Y.-Y. & Barabási, A.-L. Control principles of complex systems. Rev. Mod. Phys. 88, https://doi.org/10.1103/RevModPhys.88.035006 (2016).
Wuchty, S. Controllability in protein interaction networks. PNAS 111, 7156–7160 (2014).
Schiff, S. J. Neural Control Engineering. The Emerging Intersection between Control Theory and Neuroscience (MIT Press, Cambridge, MA, 2012).
Cornelius, S. P., Kath, W. L. & Motter, A. E. Realistic control of network dynamics. Nat. Commun. 4, 1942 (2013).
Bagnoli, F., Dridi, S., El Yacoubi, S. & Rechtman, R. Optimal and suboptimal regional control of probabilistic cellular automata. Nat. Comput. 3, 307 (2019).
Csárdi, G. & Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 1695, https://igraph.org/ (2006).
Hagberg, A. A., Schult, D. A. & Swart, P. J. Exploring network structure, dynamics, and function using NetworkX. In Pro 7th Python in Science Conference (SciPy2008) (eds. Varoquaux, G. et al.) 11–15 (Passadena, 2008).
Acknowledgements
We are grateful to Kilian Vogele and Friedrich Simmel for helpful discussions. This work is supported by the German Research Foundation via the collaborative research center SFB1032 and the Excellence Cluster ‘ORIGINS’ through U.G.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
All authors designed the research and analyzed the data. T.R., S.K., H.W. performed simulations, T.R., S.K., U.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Communications Physics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ramalho, T., Kremser, S., Wu, H. et al. Programmable pattern formation in cellular systems with local signaling. Commun Phys 4, 140 (2021). https://doi.org/10.1038/s42005-021-00639-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-021-00639-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.