Abstract
Scanning probe microscopy (SPM) has revolutionized the fields of materials, nano-science, chemistry, and biology, by enabling mapping of surface properties and surface manipulation with atomic precision. However, these achievements require constant human supervision; fully automated SPM has not been accomplished yet. Here we demonstrate an artificial intelligence framework based on machine learning for autonomous SPM operation (DeepSPM). DeepSPM includes an algorithmic search of good sample regions, a convolutional neural network to assess the quality of acquired images, and a deep reinforcement learning agent to reliably condition the state of the probe. DeepSPM is able to acquire and classify data continuously in multi-day scanning tunneling microscopy experiments, managing the probe quality in response to varying experimental conditions. Our approach paves the way for advanced methods hardly feasible by human operation (e.g., large dataset acquisition and SPM-based nanolithography). DeepSPM can be generalized to most SPM techniques, with the source code publicly available.
Similar content being viewed by others
Introduction
Scanning probe microscopy (SPM)1 consists of scanning an atomically sharp probe in close proximity (typically ≈1 nm) above a surface, while measuring a physical quantity [e.g., quantum tunneling current in scanning tunneling microscopy (STM)2 and force in atomic force microscopy (AFM)3] as a function of probe position. This allows for the construction of an image of the scanned surface. This method can be employed in a variety of environments, from ambient conditions to ultra-high vacuum at cryogenic temperatures, allowing for measurements on a wide range of samples2,4. Its unique capability to probe real-space physical properties (structural, electronic5 and chemical6) with atomic resolution and to manipulate adsorbates on a surface with atomic-scale precision7 makes it highly relevant for chemistry3 and biology4, and perhaps the most powerful characterization tool for materials, surface, and nano-science1,2,8.
The yield of SPM data acquisition—the fraction of good data usable for scientific analysis—is rarely reported and are generally low. There are two main factors limiting this yield: (i) the atomic-scale morphology of the probe can result in imaging artefacts and (ii) the state of the sample imaging regions (e.g., scanning excessively rough or contaminated regions rarely produces usable data and can result in damaging the probe). Both factors vary during the course of an experiment and need to be addressed.
In state-of-the-art SPM, a human operator selects sample regions to scan and assesses the acquired images (good or bad quality). This assessment is based on the operator’s experience. If she deems the image bad due to the state of the sample region or the probe, she changes the region or attempts to condition the probe. The de-facto procedure for the latter relies on trial-and-error. Conditioning actions (e.g., dipping the probe into the sample and applying a voltage pulse between probe and sample9) are performed until the probe morphology and image quality are restored. The probe atomic-scale structure dramatically influences image quality and outcomes of conditioning actions are uncertain; success depends on the microscopist’s experience and the time invested.
Previous studies have aimed to improve imaging efficiency, e.g., by linking probe morphology and image quality, via analytical simulations10, inverse imaging of the probe through sample features11,12,13, or probe characterization/manipulation (field ion microscopy14). These approaches can suppress the need for heuristic probe conditioning. However, they are difficult to implement for general use, in particular for large dataset acquisition.
Another avenue for minimizing the need of human operation is SPM automation15. Examples include scripted SPM operation16 to automatic imaging region selection17. A recent publication18 presents an AFM system capable of autonomous sample region selection and measurement. However, these methods are limited to specific applications in stable measurement conditions and do not manage probe quality under general operation.
The latter can be addressed via machine learning (ML), which allows for predictions, assessments, and decision-making in systems that are not fully understood, or too complex to be characterized analytically. Rather than following a well-established set of rules, ML methods derive decision strategies from training data. In image processing (e.g., object recognition19 and image segmentation20), ML approaches routinely outperform humans. These accomplishments are often based on convolutional neural networks (CNNs)19,21. Such CNNs use image or image-like data as input to perform classification and regression tasks (e.g., object recognition and image quality optimization). A CNN is controlled by millions of parameters that can be tuned via supervised learning. In this process, the network is trained using large sets of input data (e.g., images) to which a label is associated. This label corresponds to the desired output of the CNN (e.g., for object recognition and the name of the object in the image). Once the CNN is trained, it can label new unseen data.
Supervised learning has been applied to SPM in a recent study22 where ML assists a human operator in detecting and repairing a specific type of probe defect in the particular case of hydrogen-terminated silicon. In this work, a trained CNN assessed the quality of acquired SPM images; if necessary, a well-established probe-conditioning protocol was executed23. In another recent study24, ML was successfully used to determine imaging quality directly from a small number of acquired scan lines, without requiring complete images. However, fully autonomous operation for more general cases, where probe defects are varied and conditioning protocols are not well-defined, has not been demonstrated yet.
In ML applications where pre-labeled training data are not available, a CNN can still learn through trial-and-error, by receiving positive and negative feedback (rewards). This approach is known as (deep) reinforcement learning (RL)25. Such RL agents can learn to navigate complex environments26 (e.g., they excel in sophisticated games27,28).
Here we present DeepSPM, an autonomous system capable of continuous SPM data acquisition. It consists of the following: (i) algorithmic solutions to select good imaging sample regions and perform measurements; (ii) a classifier CNN trained through supervised learning that assesses the state of the probe; and (iii) a deep RL agent that repairs the probe by choosing adequate conditioning actions. DeepSPM also addresses other typically arising issues (lost contact, crashed probe, and moving probe larger (macroscopic) distances to new approach areas).
Results
To train and evaluate DeepSPM, we used a low-temperature STM with a metallic probe (Pt/Ir) to image a model sample: magnesium phthalocyanine (MgPc) molecules adsorbed on a silver surface (Fig. 1 and Methods). Such molecular systems are scientifically and technologically relevant, owing to their electronic, optical, and chemical properties29,30. SPM provides an ideal tool for their characterization but also presents challenges (e.g., image-altering probe–molecule interactions). Although spatial resolution may be poorer in comparison with a semiconducting or functionalized probe23, a metallic probe is required for many SPM techniques [e.g., scanning tunneling spectroscopy (STS)5 and Kelvin probe force microscopy (KPFM)31].
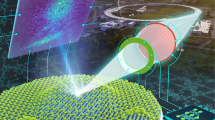
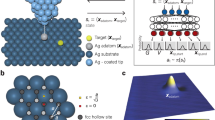
a Schematic of DeepSPM, a machine learning (ML)-based AI system for autonomous scanning probe microscopy operation [here, a low-temperature scanning tunneling microscope (STM)]. DeepSPM determines the control signals (measurement parameters) and acquires an STM image. After acquisition, DeepSPM assesses the image quality. If the image is deemed “good”, DeepSPM processes it and stores it, and performs the next measurement. If “bad”, DeepSPM detects and addresses possible issues. b STM images of MgPc molecules on Ag(100), acquired and assessed (green tick: good; red cross: bad) by DeepSPM. Examples of variable imaging conditions are shown: good probe and sample; sample area with excessive roughness; noisy image due to lost probe–sample contact; dull probe leading to blurry images; multiple-feature probe producing replicated images; contaminated probe resulting in artefacts; contaminated multiple-feature probe; unstable probe; c Good STM images processed and stored. The inset in c shows the chemical structure of MgPc. Scale bar: 2 nm. Color scale indicates the measured height, with bright colors indicating a higher surface.
DeepSPM overview
DeepSPM works as a control loop (Fig. 1a). The artificial intelligence system drives the SPM by selecting an appropriate scanning region (Supplementary Fig. 1), acquires an image, and assesses the acquired image data (Fig. 1b). If the image is deemed “good”, it is processed and stored (Fig. 1c), and DeepSPM proceeds with the next loop iteration. If the image is labeled “bad”, DeepSPM addresses the issues, maintaining continuous and stable operation (see Methods).
In Fig. 2, we give a graphical description of DeepSPM. The system is able to assess and identify causes of a defective acquisition (e.g., lost sample–probe contact, probe crash, bad sample region, bad probe). If sample–probe contact is lost or the probe crashes, DeepSPM re-establishes contact in a new scanning region (Methods). Images of bad sample regions (e.g., excessive roughness and contamination) can be identified algorithmically by the fact that measured heights span over a larger range compared with clean regions (see Methods). If a region is identified as a “bad sample”, DeepSPM selects a new one and performs a new measurement.
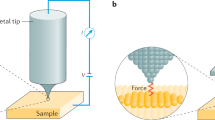
a DeepSPM approaches the SPM probe (e.g., metallic tip and oscillating cantilever) towards the sample until a measurement signal (e.g., tunneling current in a scanning tunneling microscope or frequency shift in an atomic force microscope) is detected, b selects a scanning region to image, and c performs an SPM image acquisition by scanning the probe and recording the measurement signal. d After an image is acquired, DeepSPM addresses (algorithmically) whether the sample region is overly rough, the probe–sample contact is lost, or the probe has crashed into the sample. If DeepSPM establishes that none of these events have occurred, the classifier convolutional neural network (CNN) then assesses the quality of the probe; if it is deemed good, the image is stored, and the measurement loop continues. e If the probe quality is deemed bad, the deep reinforcement learning agent attempts to repair the probe by selecting and applying a probe-conditioning action. f If DeepSPM does not find a region suitable for scanning, it re-positions the probe (macroscopically) to a new approach area.
Intelligent probe quality assessment
If DeepSPM concludes that the sample imaging region is “good”, it assesses the state of the probe: the classifier CNN (Supplementary Table 1) inspects the recorded image and predicts the probability of it being recorded with a bad probe (Fig. 3a). To train the classifier, we used a dataset of 7589 images of the MgPc/Ag(100) sample, labeled as acquired either with a “good” or “bad probe”. In addition, we used data augmentation to increase the amount of training data (see Methods). It is noteworthy that the category “bad probe” includes various kinds of probe defects (Fig. 1b). We tested its performance on an unseen test dataset, achieving an accuracy of ~94% (Supplementary Table 1), a positive predictive value ~87% and a negative predictive value ~96%. As point of reference, classification accuracy of a human in a benchmark visual object recognition challenge32 (ImageNet) ranges from 88% to 95%, on par with our CNN classifier. It is noteworthy that classification performance is intrinsic to the type of data and the specific datasets considered. We are, in concurrence with other work33, among the first to present a dataset of this kind and there are currently no available baselines for comparison. However, the performance achieved by DeepSPM enables autonomous data acquisition and long-term operation of DeepSPM.
a After image pre-processing, DeepSPM (our method) detects lost contact, probe crashes, and bad sample regions (excessive roughness) by measuring apparent height distributions. If none of these problems are detected, the classifier convolutional neural network (CNN) assesses the probe quality. b If the probe is deemed bad, the reinforcement learning agent selects and executes a conditioning action from a predefined list, with the aim of achieving the shortest possible conditioning sequence. This selection is achieved via a second (action) CNN, which predicts the cumulative future reward (Q-value) for each conditioning action. DeepSPM executes it at the center of the largest empty area found within the scanned region. c Example of probe-conditioning episode. DeepSPM repairs a dull STM probe in three conditioning steps. The conditioning actions selected by DeepSPM are displayed below each image and were executed at the white crosses. d Distribution of the mean episode length (bootstrapping, 100,000 draws43) required to restore a good probe, for random testing (random selection of one of the 12 possible conditioning actions; 184 conditioning episodes), RL agent testing (189 conditioning episodes), and RL agent autonomous operation (117 episodes). During testing, the probe was actively damaged after each conditioning episode. The RL agent continued to train during data acquisition, for both testing and autonomous operation.
Intelligent probe conditioning
If the classifier CNN concludes that the probe is bad, DeepSPM uses a deep RL agent to condition it (Fig. 3b). This RL agent is controlled by a second CNN (action CNN), which is trained by interacting with the SPM setup: the RL agent inspects the last recorded image and performs a probe-conditioning action, selected from a list of 12 actions. We determined this list by considering actions commonly used for probe conditioning by expert human operators (Methods, Supplementary Note 1, and Supplementary Table 2); they consist of either a voltage pulse applied between probe and sample, or a dip of the probe into the sample9. After each conditioning step, DeepSPM evaluates the outcome of the conditioning process by acquiring the next image, which is then assessed by the classifier CNN. If the new image is classified as “bad probe”, the agent receives a negative reward (r = −1) and proceeds with another action. If the image is classified as “good probe”, the conditioning episode (sequence of conditioning steps; Fig. 3c) is terminated and the agent receives a positive reward (r = 10; Methods).
The RL agent learns an approximately optimal conditioning procedure by attempting to maximize the cumulative reward received for each conditioning episode, thus minimizing the number of required conditioning steps (Supplementary Note 2). To achieve this, we relied on Q-learning34 (see Methods): the action CNN processes each recorded image and predicts the expected future reward (Q-value) resulting from each possible conditioning action. The RL agent then selects the action with the highest Q-value (ε-greedy policy; Methods and Supplementary Fig. 2).
To test the RL agent’s performance, we compared it with a baseline case where conditioning actions are selected randomly from the list of common actions (Fig. 3b). During testing, we actively damaged the probe after each conditioning episode (see Methods). The trained RL agent is able to condition the probe efficiently (Fig. 3d and Supplementary Figs. 3 and 4) and does so in an average number of conditioning steps ~28% smaller than in the random case. Intelligent selection of conditioning actions by the RL agent significantly outperforms random selection.
Autonomous SPM operation
We next demonstrate long-term autonomous operation of the entire DeepSPM system. For a period of 86 h, we let DeepSPM control the microscope. Figure 4a shows DeepSPM’s behavior in an approach area, highlighting occurrences of bad probe detection and conditioning, and avoiding “bad sample” regions. In Fig. 4b, we show the area scanned by the system as a function of time. In total, DeepSPM scanned a sample area of 1.2 μm2 (Fig. 4), recorded >16,000 images, handled 2 lost contacts, identified and avoided 1075 regions of excessive roughness, and repaired the probe 117 times (Supplementary Table 3).
a Example of DeepSPM’s behavior within one approach area (850 × 850 nm2) during an autonomous data acquisition run. DeepSPM approached the probe and initialized data acquisition at the center of the area (1). At (2), DeepSPM stopped the data acquisition and moved to the next approach area. The black curve indicates the probe trajectory. The plot shows valid measurement regions scanned with a good probe (green), regions deemed bad due to excessive roughness (magenta), locations where a bad probe was detected (orange) and a probe-conditioning action was performed accordingly (blue), and regions deemed bad due to proximity of excessively rough or probe-conditioning areas (gray). To account for sample variability, DeepSPM triggered probe conditioning only when it detected ten consecutive images recorded with a bad probe (see Methods). b Total sample area imaged by DeepSPM during the 86 h autonomous data acquisition run [only scanned sample areas are included; areas inferred bad (gray) were omitted]. Vertical blue lines indicate probe-conditioning events. Shaded time window corresponds to the approach area in a. (Vbias = −1V, It = 25 pA, scan speed of 80 nm s−1, 6.4 px nm−1).
To evaluate the overall performance of DeepSPM, we manually inspected the recorded images (Supplementary Note 3). Out of all images labeled “good” by DeepSPM, ~87% were found to be without defects or imaging artifacts. Out of all conditioning episodes initiated by DeepSPM, ~86% were found to be really necessary (Supplementary Fig. 5, Methods, and Supplementary Fig. 6). It is noteworthy that these performance metrics are not related to static classification (as for the classifier CNN testing), as the state of the STM/sample system and the recorded images depend dynamically on the decisions made by DeepSPM.
During autonomous operation, the RL agent achieved an average conditioning episode length of 4.93, ~34% shorter than during testing (Fig. 3d). We attribute this to the fact that, during testing, the probe was actively damaged after each conditioning episode. This was not the case during autonomous operation, where arguably the state of the probe remains closer to a good one (Supplementary Note 3).
Discussion
The available conditioning actions do not allow the RL agent to control the atomic-scale structure of the probe, which determines imaging quality. Their outcome is probabilistic and conditioning episode lengths vary (Fig. 3d). Nonetheless, our RL agent’s better-than-random performance shows that: (i) at each step of the conditioning process, it is in principle possible to intelligently choose an action that is likely to improve the probe, and (ii) that an ML system can learn to make this choice (Supplementary Note 4).
In our specific case here, the single images that DeepSPM records and that determine the RL agent’s conditioning action selection do not enable the retrieval of the atomistic morphology of the probe. Therefore, the conditioning process behaves effectively as if it had memory; the probe state depends on the specific sequence of previous actions and images. Continuous training of the RL agent during operation allows it to follow the evolution of the probe state and is hence essential to achieve better-than-random performance during operation (Supplementary Fig. 4).
Here we used RL to optimize a data acquisition protocol. The benefit of RL for such a task is that the rules for optimization (optimal choices for probe conditioning) do not need to be known in advance; an agent with sufficient training can establish them by interacting with the experimental setup, without human guidance.
Automation of complex experimental procedures such as SPM frees valuable researcher time. DeepSPM brings state-of-the-art SPM closer to a turnkey application, enabling non-expert users to achieve optimal performance.
DeepSPM can be applied directly to any sample, probe material, or STM/AFM setup, as long as specific training datasets are available (see Methods). It can further be expanded to other SPM spectroscopy techniques (e.g., STS and KPFM), where probe quality conditions would need to include additional spectroscopic requirements5,31 (Supplementary Note 5). Fully autonomous SPM also opens the door to high-throughput and scalable atomically precise nano-fabrication35, hardly feasible via manual operation.
Future work can further extend our method by combining it with semi-automatic ML approaches used for, e.g., the identification of adverse imaging conditions24,33 or imaging regions of interest18.
Methods
Sample preparation
The samples were prepared in-situ by sublimation of MgPc molecules (Sigma-Aldrich) at 650 K (deposition rate ≈ 0.014 molecules nm−2 min−1; sub-monolayer coverages) onto a clean Ag(100) surface (Mateck GmbH) held at room temperature. The Ag surface was prepared in ultra-high vacuum (UHV) by repeated cycles of Ar+ sputtering and annealing at 720 K. The base pressure was below 1 × 10−9 mbar during molecular deposition.
STM measurements
The STM measurements were performed using a commercial scanning probe microscope (Createc) capable of both STM and non-contact AFM at low temperature (down to 4.6 K) and in UHV. This setup includes two probe-positioning systems: a coarse one for macroscopic approaching and lateral positioning of the probe above the sample; a fine one consisting of a piezo scanner that allows for high-resolution imaging. The lateral range of the fine piezo scanner at 4.6 K is ±425 nm, i.e., each approach of the probe to the sample with the coarse system defines a scanning region area of 850 × 850 nm2). Nanonis electronics and SPM software (SPECS) were used to operate the setup. All measurements were performed at 4.6 K with a Pt/Ir tip. All topographic images were acquired in constant-current mode (Vbias = 1V, It = 25 pA) at a scan speed of 80 nm s−1.
After moving the probe macroscopically to a new sample area, DeepSPM extends the z piezo scanner to ~80% of its maximum extension, to maximize the range of tunneling current feedback controlling the z position of the scanner without crashing or losing contact. If tunneling contact is lost, DeepSPM re-approaches the probe. DeepSPM also handles probe crashes (see section below “Detection and fixing of lost contact/probe crash”).
After approaching the probe to the sample, DeepSPM waits ~120 s before starting a new scan, to let thermal drift and (mainly) creep of the z piezo settle. New measurements start at the neutral position of the xy scan piezo (that is, x = y = 0), where no voltages are applied to the xy piezo scanner, to minimize the piezo creep in the xy scanning plane. DeepSPM selects the next imaging region by minimizing the distance that the probe travels between regions (see Fig. 4, section “Finding the next imaging region” and Supplementary Fig. 1), further reducing xy piezo creep. After moving the probe to a new imaging region, DeepSPM first records a partial image that is usually severely affected by distortions due to xy piezo creep. This image is discarded and a second image, not or only minimally affected, is recorded at the same position.
During autonomous operation, the RL agent of DeepSPM continues to learn about probe conditioning (see main text and below). To avoid damaging a good probe by unnecessary conditioning during autonomous operation, DeepSPM initiates a probe-conditioning episode only after ten consecutive images have been classified as “bad probe” by the classifier CNN (Supplementary Fig. 6). The episode is terminated as soon as the first image is classified as “good probe” by the classifier CNN. Images that are not part of a conditioning episode (including the ten consecutive images that triggered it), or that have been disregarded due to “bad sample”, lost probe–sample contact or crashed probe, are labeled as “good image”.
Finding the next imaging region
For each new approach area, DeepSPM starts acquiring data at the center of the scanning range (Fig. 4a). DeepSPM uses a binary map to block the imaging regions that have already been scanned (Supplementary Fig. 1). If a region is identified as “bad sample” (e.g., excessive roughness is detected), DeepSPM defines a larger circular area around it (as further roughness is expected in the vicinity), and avoids scanning this area. The radius rforbidden of this region is increased as consecutive imaging regions are identified as “bad sample”:
where t is the number of consecutive times an area with excessive roughness was detected.
As probe-conditioning actions can cause debris and roughness on the sample, DeepSPM blocks a similar circular area to avoid around the location of each performed conditioning action. The size of this area depends on the executed action (Supplementary Table 2). DeepSPM chooses the next imaging region centered at position \({\mathbf{v}}_{\mathrm{t}} = x_{\mathrm{t}}{\hat{\mathbf{x}}} + y_{\mathrm{t}}{\hat{\mathbf{y}}}\) (xt, yt are coordinates with respect to the center of the approach area) that minimizes
provided that this is a valid imaging region according to the established binary map. Here, vt−1 denotes the position of the center of the last imaging region, ||…||1 is the Manhattan norm, and ||…||2 is the standard Euclidian norm. The parameter α controls the relative weight of the two distances, i.e., to the center of the last scanned region (to minimize travel distance), and to the center of the approach area (to efficiently use the entire available area). We found α = 1 works well. This algorithm minimizes the distance the probe travels between consecutive imaging regions, reducing the impact of xy piezo creep. Once the area defined by the fine piezo scanner range has been filled, or the distance to the center of the next available scanning region is larger than 500 nm, DeepSPM moves the probe (macroscopically, with the coarse positioning system) to a new approach area.
DeepSPM architecture
The DeepSPM framework consists of two components: (i) the controller, written in Python and TensorFlow, and (ii) a TCP server, written in Labview. The controller contains the image processing, classifier CNN, and RL agent. The TCP server creates an interface between the controller and the Nanonis SPM software. The controller sends commands via TCP, e.g., for acquiring and recording an image, executing a conditioning action at a certain location. The server receives these commands and executes them on the Nanonis/SPM. It returns the resulting imaging data via TCP to the controller, where it is processed to determine the next command. Based on this design, the agent can operate on hardware decoupled from the Nanonis SPM software.
Training and test dataset for classifier CNN
We compiled a dataset of 7589 images (constant-current STM topography, 64 × 64 pixels) of MgPc molecules on Ag(100), acquired via human operation. We assigned to each image a ground truth label of the categories “good probe” (25%) or “bad probe” (75%). We randomly split the data into a training (76%) and test set (24%). We used the latter to test the performance of the classifier CNN on unseen data (i.e., not used for training). The dataset is available online at https://alex-krull.github.io/stm-data.html.
It is important to note that the classifier CNN was trained to distinguish a “good probe” from a “bad probe”. The classifier CNN was not trained to identify a specific type of probe defect in the case of a “bad probe”. Figure 1b shows examples of possible imaging defects, including different types of probe defects (recognized as “bad probe” by the classifier CNN) and other image acquisition issues (e.g., lost contact and excessive sample roughness) that are detected algorithmically.
CNN architecture
We used the same sequential architecture for both the classifier CNN and the action CNN of the RL agent, differing only in their output layer and specific hyper-parameters (see below). The basic structure is adapted from the VGG network21. We used a total of 12 convolutional layers: four sets of three 3 × 3 layers (with 64, 128, 256, and 512 feature maps, respectively) and 2 × 2 max-pooling after the first two sets. The convolutional layers are followed by two fully connected layers, each consisting of 4096 neurons. Each layer, except the output layer, uses a ReLU activation function and batch normalization36. The input in all networks consisted of 64 × 64 pixel constant-current STM topography images. We used Dropout37 with a probability of 0.5 after each fully connected layer to reduce overfitting. The network weights were initialized using Xavier initialization38.
Classifier CNN
The classifier CNN uses the architecture above. It has a single neuron output layer with a sigmoid activation function. This output (ranging from 0 to 1) gives the classifier CNN’s estimate of the probability that the input image was recorded with a “good probe”. The decision threshold was set to 0.9. It is noteworthy that DeepSPM requires ten consecutive images classified as “bad probe” to start a conditioning episode (Supplementary Fig. 6). We trained the classifier CNN using the ADAM39 optimizer with a cross-entropy loss and L2 weight decay with a value of 5 × 10−5 and a learning rate of 10−3. To account for the imbalance of our training set (“good probe” 25% and “bad probe” 75%), we weighed STM images labeled as “good probe” by a factor of 8 when computing the loss40. In addition, we increased the available amount of training data via data augmentation, randomly flipping the input SPM images horizontally or vertically. It is noteworthy that all training data consisted of experimental data previously acquired and labeled manually.
Reinforcement learning agent and action CNN
Our RL agent responsible for the selection of probe-conditioning actions is based on double DQN34, which is an extension of DQN28. We modified the double DQN algorithm to suit the requirements of DeepSPM as follows. The action CNN controlling the RL agent uses the architecture above, with a single constant-current STM image as input. It is noteworthy that the original DQN uses a stack of four subsequent images. Our action CNN has an output layer consisting of 12 nodes, one for each conditioning action. The output of each node is interpreted as the Q-value of the corresponding action, i.e., the expected future reward to be received after executing it. We initialized the weights of the action CNN (excluding the output layer) with those of the previously trained classifier CNN, based on the assumption that the features learned by the latter are useful for the action CNN41. The output layer, which has a different size in both networks, is initialized with the Xavier initialization38. To train the action CNN, we let it operate the SPM, acquiring images, and selecting and executing probe-conditioning actions repeatedly when deemed necessary (Figs. 1 and 2). Once sufficient probe quality was reached (i.e., the probability predicted by the classifier CNN exceeded 0.9), the conditioning episode was terminated—a conditioning episode consists of the sequence of probe-conditioning actions required to obtain a good probe. Random conditioning actions (up to five) were then applied to reset (i.e., re-damage the probe), until the predicted probability drops below 0.1. The RL agent received a constant reward of −1 for every executed probe-conditioning action. It received a reward of +10 for each terminated training episode, i.e., each time the probe was deemed good again. We chose these reward values heuristically by testing them in a simulated environment. In these simulations, the RL agent executed conditioning actions and the reward protocol was applied based on images resulting from the convolution of a good, clean synthetic image with a model kernel representing the probe morphology. Following a conditioning action, this kernel was updated stochastically. In this reward scheme, the RL agent receives a positive cumulative reward for and favors short conditioning episodes, whereas it receives a negative cumulative reward and is punished for longer episodes.
The RL agent uses ε-greedy exploration to gather experience25. For each conditioning step, the agent chooses a conditioning action probabilistically based on parameter ε (0 < ε < 1): it chooses randomly with a probability ε, and it chooses the action with the largest predicted future reward (Q-value) with probability (1 − ε). For example, if ε = 1, action selection is strictly random; if ε = 0, action selection is based strictly on predicted Q-value. We start training (Supplementary Fig. 2) with 500 random steps (ε = 1) that are used to pre-fill an experience replay buffer28. This buffer contains all experiences the agent has gathered so far, each consisting of an input image, the chosen action and its outcome (the next image assessed by the classifier CNN, as well as the reward received). We used data augmentation, adding four experiences to the buffer for each step. These additional experiences consisted of images flipped horizontally and vertically. After 500 steps (i.e., 2000 experiences in the buffer), we started training the action CNN with the buffer data. We used the ADAM optimizer39 with a batch size of 64 images processed simultaneously and with a constant learning rate of 5 × 10−4. We limited the buffer size to 15,000, with new experiences replacing the old ones (first-in, first out). To allow parallel execution and increase the overall performance of the training, we decoupled the gathering of experience and the learning into separate threads. During training, we decreased ε linearly over 500 steps, from 1.0 to 0.05. After reaching ε = 0.05, we continued training with additional 4360 steps, during which we kept ε = 0.05 constant34. We used a constant discount factor of γ = 0.9525.
Testing of the RL agent
After training the RL agent, we tested its performance in operating the STM by comparing it with the probe-conditioning performance achieved via random conditioning action selection. During this evaluation, we allowed the action CNN to continue learning from continuous data acquisition with a constant ε = 0.05. Except for this value of ε, the testing process matches that of the RL agent training above. To achieve a meaningful comparison, we accounted for the fact that the state of the sample and the probe changes after each executed conditioning action (Supplementary Figs. 3 and 4), by adopting an interleaved evaluation scheme. That is, RL agent action selection and random selection alternate in conditioning the probe, switching after each completed probe-conditioning episode.
STM image pre-processing
The scanning plane of the probe is never perfectly parallel to the local surface of the sample. This results in a background gradient in the SPM images that depends on the macroscopic position and, to a lesser extent, on the nanoscopic shape of the probe. This gradient was removed in each image by fitting and subtracting a plane using RANSAC42 (Python scikit-learn implementation; polynomial of degree 1, residual threshold of 5 × 10−12, max trials of 1000). The acquired STM data were further normalized and offset to the range [−1; 1], i.e., such that pixels corresponding to the flat Ag(100) had values of −1 and those corresponding to the maximum apparent height of MgPc (~2 Å) had values of 1. In addition, we limited the range of values to [−1.5, 1.5], shifting any values outside this range to the closest one inside the interval.
Finding an appropriate action location
For a given acquired STM image, DeepSPM executes a probe-conditioning action at the center of the largest clean Ag(100) square area (Fig. 3). This center is found by calculating a binary map from the pre-processed image (see above), where pixels close (≤0.1 Å) to the surface fitted plane are considered empty, i.e., belong to a clean Ag(100) patch, and all others as occupied. The center of the largest clean Ag(100) square area within this binary map was chosen as the conditioning location. We defined an area requirement for each conditioning action (Supplementary Table 1). A conditioning action is allowed and can be selected by the agent only if the available square area is within this specified requirement.
Detection and fixing of lost contact/probe crash
DeepSPM is able to detect and fix any potential loss of probe–sample contact during scanning. It does so by monitoring the extension (z-range) of the fine piezo scanner; if the fine piezo scanner extends in the z-direction beyond a specified threshold (towards the sample surface), DeepSPM prevents the potential loss of probe–sample contact by re-approaching the probe towards the sample with the coarse probe-positioning system (until the probe is within an acceptable distance range from the sample). Data acquisition can then continue at the same position. Similarly, DeepSPM can prevent probe–sample crashes, i.e., by increasing the probe–sample distance with the coarse probe-positioning system if the fine piezo scanner retracts in the z-direction beyond a specified threshold (away from the sample surface).
Data availability
Relevant data are made available by the authors. Our classification dataset is available online at https://alex-krull.github.io/stm-data.html. A visual representation of the probe-shaping episodes of the autonomous operation is available as Supplementary Data 1.
Code availability
The source code will be made available at https://github.com/abred/DeepSPM.
References
Meyer, E., Hug, H. J. & Bennewitz, R. Scanning Probe Microscopy: The Lab on a Tip (Springer, Berlin, 2004). .
Ramachandra Rao, M. S. & Margaritondo, G. Three decades of scanning tunnelling microscopy that changed the course of surface science. J. Phys. D. Appl. Phys. 44, 460301 (2011).
Pavliček, N. & Gross, L. Generation, manipulation and characterization of molecules by atomic force microscopy. Nat. Rev. Chem. 1, 0005 (2017).
Dufrêne, Y. F. et al. Imaging modes of atomic force microscopy for application in molecular and cell biology. Nat. Nanotechnol. 12, 295–307 (2017).
Wilder, J. W. G., Venema, L. C., Rinzler, A. G., Smalley, R. E. & Dekker, C. Electronic structure of atomically resolved carbon nanotubes. Nature 391, 59 (1998).
Gross, L., Mohn, F., Moll, N., Liljeroth, P. & Meyer, G. The chemical structure of a molecule resolved by atomic force microscopy. Science 325, 1110–1114 (2009).
Crommie, M. F., Lutz, C. P. & Eigler, D. M. Confinement of electrons to quantum corrals on a metal surface. Science 262, 218–220 (1993).
Binnig, G. & Rohrer, H. Scanning tunneling microscopy—from birth to adolescence. Rev. Mod. Phys. 59, 615–625 (1987).
Tewari, S., Bastiaans, K. M., Allan, M. P. & van Ruitenbeek, J. M. Robust procedure for creating and characterizing the atomic structure of scanning tunneling microscope tips. Beilstein J. Nanotechnol. 8, 2389–2395 (2017).
Villarrubia, J. S. Algorithms for scanned probe microscope image simulation, surface reconstruction, and tip estimation. J. Res. Natl Inst. Stand. Technol. 102, 425 (1997).
Welker, J. & Giessibl, F. J. Revealing the angular symmetry of chemical bonds by atomic force microscopy. Science 336, 444–449 (2012).
Chiutu, C. et al. Precise orientation of a single C_60 molecule on the tip of a scanning probe microscope. Phys. Rev. Lett. 108, 268302 (2012).
Schull, G., Frederiksen, T., Arnau, A., Sánchez-Portal, D. & Berndt, R. Atomic-scale engineering of electrodes for single-molecule contacts. Nat. Nanotechnol. 6, 23–27 (2011).
Paul, W., Oliver, D., Miyahara, Y. & Grütter, P. FIM tips in SPM: Apex orientation and temperature considerations on atom transfer and diffusion. Appl. Surf. Sci. 305, 124–132 (2014).
Extance, A. How atomic imaging is being pushed to its limit. Nature 555, 545–547 (2018).
Krenner, W., Kühne, D., Klappenberger, F. & Barth, J. V. Assessment of scanning tunneling spectroscopy modes inspecting electron confinement in surface-confined supramolecular networks. Sci. Rep. 3, 1454 (2013).
Rusimova, K. R. et al. Regulating the femtosecond excited-state lifetime of a single molecule. Science 361, 1012–1016 (2018).
Huang, B., Li, Z. & Li, J. An artificial intelligence atomic force microscope enabled by machine learning. Nanoscale 10, 21320–21326 (2018).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. in Advances in Neural Information Processing Systems 25 1097–1105 (Curran Associates, Inc., 2012).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. Preprint at https://arxiv.org/abs/1409.1556 (2014).
Rashidi, M. & Wolkow, R. A. Autonomous scanning probe microscopy in situ tip conditioning through machine learning. ACS Nano 12, 5185–5189 (2018).
Wang, J. et al. Direct imaging of surface states hidden in the third layer of Si (111)-7x7 surface by pz-wave tip. Appl. Phys. Lett. 113, 031604 (2018).
Gordon, O., Junqueira, F. & Moriarty, P. Embedding human heuristics in machine-learning-enabled probe microscopy. Preprint at https://arxiv.org/abs/1907.13401 (2019).
Sutton, R. S. & Barto, A. G. Reinforcement Learning: An Introduction. Second Edition (MIT Press, Cambridge, 2018).
Duan, Y., Chen, X., Houthooft, R., Schulman, J. & Abbeel, P. Benchmarking deep reinforcement learning for continuous control. In Proc. Int. Conference on Machine Learning (ICML) 48, 1329–1338 (International Machine Learning Society (IMLS), 2016).
Silver, D. et al. Mastering the game of Go without human knowledge. Nature 550, 354–359 (2017).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Auwärter, W., Ecija, D., Klappenberger, F. & Barth, J. V. Porphyrins at interfaces. Nat. Chem. 7, 105–120 (2015).
Li, C., Wang, Z., Lu, Y., Liu, X. & Wang, L. Conformation-based signal transfer and processing at the single-molecule level. Nat. Nanotechnol. 12, 1071–1076 (2017).
Mohn, F., Gross, L., Moll, N. & Meyer, G. Imaging the charge distribution within a single molecule. Nat. Nanotechnol. 7, 227–231 (2012).
Russakovsky, O. et al. ImageNet large scale visual recognition challenge. Int. J. Comp. Vis. 115, 211–252 (2015).
Gordon, O. et al. Scanning tunneling state recognition with multi-class neural network ensembles. Rev. Sci. Instrum. 90, 103704 (2019).
Hasselt, H. van, Guez, A. & Silver, D. Deep reinforcement learning with double Q-learning. In Proc. AAAI Conference on Artificial Intelligence 2094–2100 (AAAI Press, 2016).
Achal, R. et al. Lithography for robust and editable atomic-scale silicon devices and memories. Nat. Commun. 9, 2778 (2018).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. In Proc. Int. Conference on Machine Learning (ICML) (International Machine Learning Society (IMLS), 2015).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Glorot, X. & Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. In Proc. Int. Conference on Artificial Intelligence and Statistics 9, 249–256 (Journal of Machine Learning Research (JMLR), 2010).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Panchapagesan, S. et al. Multi-task learning and weighted cross-entropy for DNN-based keyword spotting. In Proc. Annual Conference of the International Speech Communication Association (Interspeech) (International Speech Communication Association (ISCA), 2016).
Zamir, A. R. et al. Taskonomy: disentangling task transfer learning. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 3712–3722 (Institute of Electrical and Electronics Engineers (IEEE), 2018).
Fischler, M. A. & Bolles, R. C. Random sample consensus: a paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 24, 381–395 (1981).
Efron, B. & Tibshirani, R. An Introduction to the Bootstrap (Chapman & Hall Boca Raton, 1993).
Acknowledgements
We thank Florian Jug for his feedback on the manuscript. A.S. acknowledges support from the Australian Research Council (ARC) Future Fellowship scheme (FT150100426). C.K. acknowledges support from the ARC Center of Excellence in Future Low-Energy Electronics Technologies. Part of the computations were performed on an HPC Cluster at the Center for Information Services and High Performance Computing (ZIH) at TU Dresden. This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 program (grant number 647769).
Author information
Authors and Affiliations
Contributions
A.K., P.H., and C.K. performed all experiments and analyzed all data. A.K., P.H., A.S., C.R., and C.K. designed the experiments, interpreted the data, and wrote the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Krull, A., Hirsch, P., Rother, C. et al. Artificial-intelligence-driven scanning probe microscopy. Commun Phys 3, 54 (2020). https://doi.org/10.1038/s42005-020-0317-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-020-0317-3
This article is cited by
-
Holotomography and atomic force microscopy: a powerful combination to enhance cancer, microbiology and nanotoxicology research
Discover Nano (2024)
-
Single-molecule chemistry with a smart robot
Nature Synthesis (2024)
-
A dynamic Bayesian optimized active recommender system for curiosity-driven partially Human-in-the-loop automated experiments
npj Computational Materials (2024)
-
Intelligent synthesis of magnetic nanographenes via chemist-intuited atomic robotic probe
Nature Synthesis (2024)
-
Deep learning based atomic defect detection framework for two-dimensional materials
Scientific Data (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.