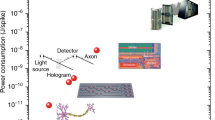
Abstract
Optical neural networks process information at the speed of light and are energetically efficient. Photonic artificial intelligence allows speech recognition, image classification, and Ising machines. Modern machine learning paradigms, as extreme learning machines, reveal that disordered and biological materials may realize optical neural networks with thousands of nodes trained only at the input and at the readout. May we use living matter for machine learning? Here, we employ living three-dimensional tumor brain models to demonstrate a random optical learning machine (ROM) for the investigation of glioblastoma. The tumor spheroid act as a computational reservoir. The ROM detects cancer morphodynamics by laser-induced hyperthermia, quantifies chemotherapy, and cell metabolism. The ROM is a sensitive noninvasive smart probe for cytotoxicity assay and enables real-time investigation of tumor dynamics. We hence design and demonstrate a novel bio-hardware for optical computing and the study of light/complex matter interaction.
Similar content being viewed by others
Introduction
Brain-inspired complex and disordered systems are emerging as computing reservoirs in machine learning, neuromorphic computing, image processing, and cryptography1. In conventional deep neural networks (D-NNs), the training becomes demanding as the number of nodes growths2,3. For this reason, new architectures containing a large number of untrained random nodes—a computing reservoir—are very appealing. Theorems and numerical experiments show that random NNs preserve the functionalities of D-NNs4,5. Random NNs include the extreme-learning machine6 (ELM), and deep reservoir computing (D-RC)7. Random NNs do not require the precise control of each of the deep computational layers and are trained only at the input and output layers. Random NNs enable new hardware with thousands of nodes and are rapidly emerging in classical and quantum optical computing8,9,10,11,12,13,14. Optical neuromorphic computing processes information at the speed of light15,16,17,18,19, and recently, photonic spin-glasses and Ising machines have been experimentally demonstrated20,21,22,23,24,25. Random optical-learning machines (ROMs) may also have innovative and surprising applications in biophysics and medicine, because any biological system may act as a reservoir for mixing light signals and perform computations26.
Here, we demonstrate that a tumor-based ROM is a novel approach to investigate cancer morphodynamics under the action of external stimuli as hyperthermia and chemotherapy. We show that a ROM may output information for tumor diagnosis and drug efficacy, which seems competitive with more conventional approaches, as biochemical assays and confocal imaging. Our living random optical neural network has thousands of cells acting as wave-mixing nodes made of glioblastoma cells, which form a large-scale computing reservoir and enable the detection of tumor morphodynamics. The biocomputing optical system measures biophysical quantities, as cancer metabolism, which commonly requires invasive methods.
Results
Tumor models as optical computing reservoirs
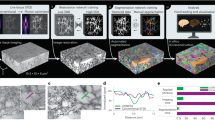
A tumor is a population of abnormal cells with temporally unrestricted growth. Most tumors are composed of different types of cells with remarkable variety in metastatic ability and chemotherapy resistance27. The spatial features of the tumor architecture are fundamental markers of cancer growth and invasiveness behavior28. We use three-dimensional (3D) tumor models (3DTMs) or spheroids29, which are used in oncology and are a platform for studying complex cell-to-cell interactions and anti-cancer therapeutics. For light, a 3DTM is a time-evolving complex assembly of disordered scattering cells and—as we demonstrate here—discloses applications for optical computation. In our biophotonic hardware, the 3DTM cellular layers are the diffractive deep layers of the ROM (Fig. 1). By exploiting structured light propagation30,31, we show that the ROM can perform programmed functions6.
a Conventional optical deep neural network (D-NN) with multiple layers (L1, …, LN). Each point is a neuron trained to perform a function between input and output planes with data Xn, as in diffractive optical networks16 and nanophotonic neural networks17. Bn and Wn are the bias and the linear transfer matrix at each layer n. F is a nonlinear activation function. The network is ordered with bias at each node (indicated by different colors). b Biophotonic random optical-learning machine (ROM): a random neural network with the tumor spheroid as a computing reservoir. Three-dimensional (3D) representation of the reservoir layers by z-stack confocal microscopy images of the tumor spheroid. In the input, a spatial light modulator (SLM) with feedback by the output layer tailors the input signal. The internal layers are random multicellular assemblies, which are not controlled or trained. Each cell is a scattering center with a complex transfer function. Owing to the biological nature of the reservoir, the internal weights change because of tumor spontaneous and stimulated evolution.
In optical NNs, the many signals composing a laser beam32,33 are mixed by using waveguides17, or in free-space16, and processed by a nonlinear operation at detection. Figure 1 compares the conventional optical D-NN architectures with the ROM. In the reported optical-learning machines16 (Fig. 1a), the trained internal matrix W and the biases B are implemented by tunable waveguide devices17 or customized diffractive layers16. In our design, we realize the input layer by an iteratively trained spatial light modulator (SLM), and the light propagates through the random layers of the 3DTM (Fig. 1b). The number, position, size, and optical properties (such as absorption and refraction) of the complex assembly of tumor cells determine the time-dependent internal weights of the reservoir.
Tumor model fabrication and laser-induced hyperthermia
We use glioblastoma cells to form brain cancer models fabricated following a common protocol29 for 3D spheroids (see “Methods”). Among 3D cell culture systems, we use non-scaffolded 3D spheroid cultures34. If compared with in vivo tumors, the spheroid mimics heterogeneity, internal structures growth kinetic, and, remarkably, drug response28. We consider both static and time-evolving 3DTMs. The evolution is either induced all-optically by nonlinear optical (thermal) effects due to an infrared laser pump (used to simulate hyperthermia) or freely evolving, due to the spontaneous cancer growth (eventually hampered by chemotherapy).
Optical reservoir computing with tumor spheroids
Figure 2 details the internal organization of the ROM. One can identify different kind of layers that convert the signal from the optical to the electronic domain. The values \(A{}_i\) of the light intensity on the SLMʼs segments are the components of the input vector \({\mathbf{x}}\). The SLM and the optical input state realize an optical convolutional layer whose output is the hidden state vector \({\mathbf{h}}^{(1)}\). The 3DTM acts as a disordered untrained layer with output given by the hidden state vector \({\mathbf{h}}^{(2)}\). The optical output of the 3DTM is imaged on the charge-coupled device (CCD) camera (output \({\mathbf{h}}^{(3)}\)), which performs a nonlinear function by measuring the laser intensity. The digital image form the CCD is the input to an electronic pooling convolutional network35, which produces the signals denoted as \(g_1\), \(g_2\), …, \(g_n\). Each of these signals is an average over the pixels on a camera segment. The output of the system is the linear combination \(y = \mathop {\sum}\nolimits_{i = 1}^n {\beta _ig_i} \left( {\mathbf{x}} \right)\), as in the ELM6. A proper choice of the training coefficients at the readout \(\beta _1,\beta _2,...,\beta _n\) enables to implement a target function. For a single output spot, only one \(\beta_{i}\) is non-zero, for two spots we retain two \(\beta_{i}\), etc. The number \(n\) of \(\beta_{i}\) (i.e., the number of “output nodes”) is a parameter optimized to realize a given target function. All the configuration variables, as the number of SLM segments, the optical setup (magnification, numerical aperture, etc.) are the “hyperparameters” of the network. Therefore, the output signal is a function of the many variables of the ROM, as the laser intensity profile, the structure and refractive properties of the 3DTM, the optical setup, the CCD and SLM parameters. Proper training enables to extract relevant biophysical information from the device [see Supplementary Notes 1 and 2].
The optical layers are formed by the laser beam (input) and the spatial light modulator (optical convolutional layer). The 3D Tumor Model (3DTM) is the biophysical random computing reservoir. The charge-coupled device (CCD) camera acts as an electronic pooling layer and the extreme-learning machine (ELM) readout converts the signal in relevant measurements, as the tumorʼs response time and metabolism. The corresponding mathematical expressions of the ROM states are also indicated. The network can be continuously retrained to follow the dynamics of the 3DTM on slow time scales. In the bottom panels, external perturbations on the 3DTM, such as infrared (IR) laser pumping and chemotherapy, introduce further random layers and change the computing reservoir.
In the training phase, after a proper choice of the βi, we optimize the SLM weights for a target output intensity \(y\). The 3DTM is left untrained as a large-scale computing reservoir. The 3DTM mixes its input signal into many signals by deflecting in random directions the modulated input beam. We remark that, although in general the spheroid has a nonlinear optical response (mainly because of the thermal heating due to the input signal) and absorbs a portion of the input light, in our scheme the 3DTM performs a linear transformation on the input field. In sections I and II of the SI, we report the simplest model for the 3DTM in terms of Maxwell equations. We find that the spheroid acts on the input signal as a neural network layer, which weights are random complex coefficients.
Although we do not train the internal layer (which would require the individual control of thousands of cells), we perturb the 3DTM by control stimuli: (i) the IR laser pump and (ii) the addition of chemotherapy. As shown in the bottom panels of Fig. 2 (and detailed in the section II of the SI), external perturbations are mathematically equivalent to adding further random layers to the reservoir. One has an additional layer for each control input, and can turn on and off the additional internal random layers by switching on and off the corresponding perturbation. Further perturbations, such as mechanical stress, can also be adopted. Figure 2 shows that the perturbed ROM is a random network in which the signal tunnels through many random untrained layers7. In the linear regime we consider, this transformation is described by a random linear function. Alternatively, the network can be viewed as a modified single-layer perceptron in which all the weights change by coupling with an additional part of the network represented by external perturbations. In general, the fact that not all channels are retrieved (unidirectional flow) and the 3DTM can present weak nonlinearity, opens possibilities for building a “deep” network. However, studies show that reservoir computing also works with linear activation functions7 and hereafter we model our results according to this case.
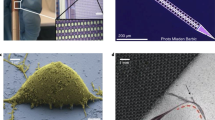
Figure 3a shows the experimental implementation of the biophotonic ROM (see “Methods” and section II of the SI). Figure 3b shows the training stage for a fixed spot image (the 3DTM is fixed by a chemical treatment). The fixed spheroids still retain some level of elasticity but the output pattern is stable over the time scales of interest (several minutes). Our goal is to train the system in a way such that the output signal computes relevant biophysical parameters of the 3DTM.
a Implementation of the biophotonic random optical-learning machine (ROM). A continuous-wave (CW) laser beam (wavelength λ = 532 nm) is phase-modulated by the spatial light modulator (SLM) in N = 72 × 72 addressable input modes that are mixed by the spheroid. Insets show a trained phase mask on the SLM and a representative confocal image of a central plane of a 3D tumor model (3DTM) labeled with diamidine-fenilindolo (in cyan, marks cell nuclei) and Phalloidin (in red, marks cells actin cytoskeleton). A charge-couple device (CCD) camera detects the transmitted light. b Intensity distribution (arb. units) in the output plane during training for an input wave that is initially randomly shaped and reaches the target shape for any spheroid by a proper fixing of the input weights. c The ROM performs different optical transformations, such as focusing on a single-point, two-points, and four-points targets. Intensities are normalized to the average transmission. Scale bars are 50 µm. P1, P2: polarizers; L1, L2, L3: lenses; OBJ1, OBJ2: microscope objectives. SH: shutter. The infrared (IR) laser is used to induce hyperthermia.
Sensing the thermal-response by the ROM
We first consider glioblastoma 3DTM that are chemically fixed (see “Methods”) and do not evolve with time, and we demonstrate morphology alterations by inducing hyperthermia. Infrared (IR) laser-induced temperature variations in brain tissue are studied in many applications, such as imaging and cancer therapy36,37. There are several open questions concerning the temperature response in cell assemblies. Temperature can affect gene expression and development. Specifically, the thermo-response may enact thermodynamic transitions whose physiological relevance is still unknown38.
To induce changes in the fixed spheroid, we use an IR pump beam that locally heats the cells (“Methods” and Supplementary Note 3). By affecting the internal structure of the reservoir through thermally activated changes in the refractive index and geometry, the IR laser modifies the output of the ROM (Fig. 4a). As the cancer spheroid changes morphology, the single-point target image moves and blurs39, and the intensity autocorrelation decreases. Similarly, a simple speckle-pattern correlation technique does not allow tracking of the structural changes since the output intensity rapidly decorrelates as the perturbation is activated (see Supplementary Fig. 8). Using recurrent training, we continuously change the SLM weights to keep constant the output (Fig. 4b). The continuous control allows us to maintain a high signal-to-noise ratio at the machine output (see Supplementary Fig. 8) and enables us to follow the network dynamics on a longer time scale.
a, b Evolution of the trained output under infrared pumping: temporal dynamics of the intensity y-profile in arbitrary units along with the single-point target for (a) static input weights and (b) recurrent feedback on. Vertical dashed lines indicate the time interval of active pumping. c Input weights under recurrent training for IR pumping with power P = 0.6 W. The color plot shows the phase value for 72 of the N modes of the spatial light modulator as a function of time. The ROM adjusts its input weights to maintain the target operation and compensate for the spheroid variations. d Tumor response time varying the pump power (dots, error bars—calculated as one standard deviation—are within the symbols) and best fit.
Recurrent training is commonly adopted in D-NN to follow signals varying on multiple time scales as in speech recognition3,40. To evidence the ability of the device to keep the signal level at the maximum, Fig. 4c shows the weights of the ROM versus time. From the time-dependent output of the ROM, we evaluate the tumor thermal-response time when varying the pump power (Fig. 4d and Supplementary Note 4). This quantity is the time scale characterizing structural rearrangement of the tumor network to a rapid temperature change, i.e., the average time taken by the 3DTM to equilibrate its structure. We are hence able to measure the response of the 3DTM at various IR pump levels, information which is extremely relevant to understand heat exchanges in tumor assemblies. It is relevant to note that the induced structural changes are irreversible, i.e., when no feedback is present and after the pump is removed, the output pattern does not return to a configuration correlated to the initial state.
Confocal microscopy analysis of laser-induced hyperthermia
To compare the measurement of the 3DTM thermal-response obtained with the ROM with a conventional optical approach, we show in Fig. 5a the confocal optical images (COI) obtained on the same sample used as computing reservoir. We investigate DAPI-stained COI measurements obtained both with and without IR irradiation, reproducing the external stimulus that the 3DTM has inside the ROM. We find that fluorescence optical imaging does not reveal any change in the nuclei morphological configuration, as also confirmed by further analysis reported in the Supplementary Notes 6 and 7. At variance with the ROM, confocal microscopy does not detect structural changes induced by hyperthermia and does not provide any information on the thermal-response time or its dependence on the IR pump. As shown in Fig. 5b, we analyze the statistical distribution of various morphological features of DAPI-stained nuclei extracted from COI and we does not find any appreciable effect of the IR pump.
a Confocal microscopy of spheroids labeled with 4’6-diamidino-2-phenylindole (DAPI) before and during irradiation with IR pulses (30 s after turning on the IR laser). Each dot in the figure is a cell, and scale bar is 30 µm. b Morphological features of cell nuclei before (“pre”, blue) and during (“post”, red) irradiation with IR pulses by image analysis with FIJI software43. Histograms show that statistically significant alterations of nuclei nearest neighbor distances (Nnd), nuclei area, aspect ratio, circularity and solidity after hyperthermia treatment are not observable.
The ROM has a very high capability to sense small tumor internal changes. In fact, the output of the biophotonics neural network contains information arriving from each elementary portion (nodes) of the sample volume. This fact explains how internal alterations that are not resolved by direct imaging are instead detected at the ROM output. Morphological variations are beyond the resolution limit of our confocal microscopy analysis (~200 nm) and, therefore, not quantifiable by this optical probe. Monitoring these thermal changes in cells would requires more complex optical methods with their own difficulties, such as optical diffraction tomography41. Previous investigations38 were able to reveal strong volume variations of cell nuclei in response to laser-induced temperature changes with an amplitude comparable to our experiments, but these observations refer to isolated nuclei in suspension in an optical stretcher38.
Conventional optical characterization of the living 3DTM
We then consider living 3DTM evolving with time. In the study of 3DTM morphodynamics, biophysical quantities as the metabolism are extremely relevant. For example, the effect of chemotherapy on the number of self-reproducing cells provides a measure of therapy efficacy. It is critical to understand the minimal amount of chemotherapy that influences tumor growth. During the tumor self-replication, the number of cells and their morphology change, and this impacts the 3DTM metabolism (denoted M hereafter). Figure 6a shows the images of a single 3DTM sample as time varies. The control sample (CTRL) has not been treated with chemotherapy, and it shows the way the tumor freely evolves and self-reproduces. Figure 6a shows a representative spheroid (CIS) treated by chemotherapy (cisplatin 80 µg/ml). Owing to the cytotoxic action of chemotherapeutics (“Methods”)42, we observe a reduced growth (Fig. 6) as the natural evolution is counter-balanced by the apoptotic effect of the cisplatin. However, despite minor differences, the small cisplatin concentration does not substantially impact the information retrieved by the optical methods. Figure 6b, c reports intensity histograms of the images in Fig. 6a (i.e., the number of pixels within the spheroid having a certain intensity level); broader distributions reveal a lower cell density and indicate that the chemotherapy limitedly hampers the tumor growth. Figure 6d compares the calculated area for CTRL and CIS versus time. The two signals are distinguishable; however, the optical microscopy characterization provides a limited accuracy in determining the effect of cisplatin.
a Bright-field images of untreated glioblastoma spheroids (CTRL) show a temporal increase in diameter compared to cisplatin-treated (CIS) samples. b, c Intensity histograms (measured as the number of pixels with a certain intensity) of spheroids images (CTRL in panel b, CIS in panel c). For both samples, histogram curves move towards darker intensities as a function of time, indicating an increase of the number of «scatterers» (i.e., cells). d Time-evolution of the area (normalized to the initial value) of the CTRL and CIS (80 μg/ml) spheroids. e Spheroids metabolism is quantified using resazurin conversion to resorufin, and it is normalized to the initial value (error bars are the standard deviation over biological replicates).
We obtain a more accurate measurement of the metabolism by resorting to fluorescence quantification of cells metabolic activity. Figure 6e reports the growth of the measured metabolism versus time (see “Methods”). The CIS sample presents a reduced metabolism due to the drug action, whereas, in the absence of chemotherapy (CTRL sample), the metabolism increases with the number of cells. We remark that the metabolism analysis in Fig. 6e requires laborious steps and labeling on several different samples (see “Methods”). As we demonstrate in the following, the same information is retrieved by using a living 3DTM in the ROM, which furnishes accurate and time-resolved information but leaves the sample unperturbed.
Characterization of the living 3DTM by the ROM
To demonstrate the tracking of the morphodynamics by the ROM, we realize the optical network with living 3DTM for the CTRL and CIS samples. We observe that, in the presence of recurrent training, the machine output follows the 3DTM dynamics (Fig. 7a, b) and needs to be continuously retrained to maintain the maximum intensity in the output target. The ROM can track cellular processes in the 3DTM beyond the pure unconstrained growth, but also with the presence of cisplatin. The output signal increases on large time scales (Fig. 7c). This dynamics results from the 3DTM growth and it is an outcome of an enhanced metabolism, as confirmed by monitoring the spheroid with conventional methods (Fig. 6d, e)43.
a, b Intensity profile (arb.units) in the y-direction versus time. The ROM is trained to focus light in a single point and operates under recurrent training to track the living 3DTM (CTRL, panel a) also in the presence of chemotherapeutic (CIS, panel b). c Persistence time TP measured by the ROM for freely evolving (CTRL) and chemotherapy-administered (CIS) spheroids. d Metabolism M versus the measured persistence time. The two sets of data correspond to the CTRL and CIS samples, which collapse on a single unique master curve showing the direct link between TP and M (normalized to the initial value for comparison). Errors are one standard deviation over biological replicates; in d they are within the symbols.
Previous work has shown that the output of a random optical process in an evolving environment corresponds to the so-called persistence time TP of transmitted speckle44. In our case, the persistence time is determined by the time scale of the 3DTM internal changes (cell positions and refractive properties). TP scales linearly with the efficiency of the target image formation44 (see Supplementary Notes 4 and 5). We use this quantity as a precursor for detecting tissue structural and metabolic changes. The measured TP is reported in Fig. 7c. The trend of TP for CTRL and CIS is remarkably distinct (at variance with the area signals in Fig. 6d), which demonstrates the high sensitivity of the ROM. By analyzing the TP in Fig. 7c, we obtain evidence of the fact that the output of the machine is a direct indication of the metabolism. Figure 7d shows TP as a function of the metabolism (measured by the conventional method in Fig. 6e). It is notable that the two metabolism signals in Fig. 6e collapse in a single master curve when plotting them in terms of TP. The two samples have the same metabolism at different instants (the CTRL evolve freely, while chemotherapy hampers the CIS; hence the CIS needs longer times to growth), but the ROM gives the same output when the two metabolisms are comparable. The continuously trained network gives a direct measure of metabolism.
These results demonstrate that the ROM can directly provide biophysical information of the 3DTM. The recurrent training of the network enables to follow non-invasively the time-evolution of the relevant biophysical observables. The computation performed by the ROM indicates if and how fast the tumor is changing in response to an external stimulus. The result can be applied to discriminate the effect of different chemotherapy treatments by separating treatments with different concentrations, or effectiveness, even though without distinguishing the specific features of the therapy. The ROM is a noninvasive tracking method since it does not require any mechanical or chemical modification of the 3DTM sample. In contrast, fluorescence microscopy in Fig. 5 is not label-free and metabolism measurements in Fig. 6e requires cell treatment (see “Methods”).
Once established that the ROM can compute the spheroid metabolism, we can show that it provides a time-dependent characterization and it is much more sensitive than a standard optical microscopy characterization as that reported in Fig. 6a. By comparing the signals in Fig. 6d (conventional optical microscopy) and 7d (trained ROM), one sees that the ROM can distinguish by at least one order of magnitude the dynamics of the CIS and CTRL in the initial time interval when the imaging method do not reveal statistically relevant changes. The effectiveness of the ROM outcome does not depend on the 3DTM size. On the contrary, in microscopy, to detect area variations for spheroids with larger sizes it is necessary to increase the microscope field of view, thus losing resolution. The living ROM hence furnishes real-time scalable information on the tumor dynamics with a sensitivity considerable larger than obtainable by monitoring structural parameters with a confocal microscopy approach.
Discussion
Nowadays various methods are being developed to characterize tumor spheroids, including immunohistochemistry, confocal laser scanning microscopy, and two-photon microscopy28. All these techniques have various advantages, but also many drawbacks. The most significant limitations of the pathological analysis of tumor morphology—commonly adopted for biopsies from patients—28are the artifacts of the cutting process, the variability in staining and the batch effects and the lack of information about dynamics. Live imaging is being improved to follow cell dynamics, but dyes, probes, and phototoxic routines modify native cell behavior34. Our random optical-learning machine furnish—after proper training—direct time-resolved structural information, which is real-time, label-free, and extremely sensitive, as it depends on thousands of scattering events and interference processes occurring in the entire volume of the 3DTM.
We demonstrated an hybrid biological/photonic computational hardware. The use of biological matter in photonic devices is not only relevant for understanding fundamental processes in tumor dynamics but allows us to explore evolving cellular assemblies as new architectures for computing. May we realize computing devices with biological systems? May we use biophotonic random neural networks to study fundamental processes in tumor evolution, and assessing the therapy efficacy? Our demonstration is an essential step in answering these and related questions. Although, as a sensing method, our approach does not allow distinguishing external stimulus, it furnishes a quantitative response on the time scale of the ongoing process that may be challenging to obtain non-invasively with other techniques. On the other hand, tracking by the ROM can be further developed to differentiate various applied perturbations by coupling with additional probes to increase the dimensionality of the output signal. For instance, through an additional broadband input source and performing also spectral measurements during recurrent training we could discriminate between two different processes inducing the same metabolism.
Moreover, leveraging on the universality of random neural networks45, one can imagine developing specially- trained ROM for properties as spatial heterogeneities in the tumor dynamics by using multipoint correlations in the output signals or resolving processes in different time scales. The use of multimode fibers for coupling light with the sample would allow to miniaturize the device and to employ it in conditions more close to in vivo situations. The ROM and its many possible developments open several possibilities for new computational systems, novel diagnostic equipment, and - more in general—for the application of photonic and artificial intelligence to the study of living complex systems.
Methods
Tumor spheroids growth
Three-dimensional tumor models have been prepared as reported in ref. 29 using human glioblastoma cell line U-87 MG (ATCC® HTB-14™) cultured in Dulbecco’s Modified Eagle Medium with 10% fetal bovine serum at 37 °C, 5% CO2. For forming the spheroids, 2500 cells have been seeded into ultra-low-adherence round-bottomed 96-well plates and immediately centrifuged (300 × g, 2 min) obtaining a suspended loose cell aggregate. After 4 days, tight spheroidal cell aggregates had formed. Depending on the experiments, spheroids have been analyzed in two possible states, namely fixed or living state. Fixed spheroids have been prepared with formaldehyde (3.7%) and glutaraldehyde (2.5%) mixture for 30 min and then washed with phosphate-buffered saline (PBS). Living spheroids have not been processed to monitor real-time response to therapy.
Chemotherapy treatment
Spheroids treated with different concentrations (10–80 μg/ml) of Cisplatin (Accord Healthcare) were imaged with a multi-well plate reader (Cytation 3, Biotek) at controlled temperature (37 °C) over time to analyze spheroid growth from bright-field images. Image analysis has been performed using INSIDIA ImageJ Macro29.
Optical experimental setup
Light from a continuous-wave (CW) laser source with wavelength λ = 532 nm is expanded and made to impinge on a twisted nematic liquid crystal reflective modulator (SLM, Holoeye LC-R 720). The active area of the SLM is divided into N = 72 × 72 independent blocks (100 pixels per block) forming the set of controlled input modes (input layer). Using a suitable combination of incident and analyzed polarizations (polarizers P1 and P2), the SLM is set into a phase-mostly modulation mode with <10% residual intensity modulation. Phase-modulated light is imaged through a 1:3 demagnifying telescope (lens L1 and L2) on the back-focal plane of a long working distance objective (OBJ1, 50x, NA = 0.55), so that the phase of each block of the SLM matches a wavevector at the entrance of the sample. The 3 mW beam is focused inside the cancer spheroid, and the intensity transmitted in a plane 1.5 mm apart is imaged on a CCD camera by a collecting objective (OBJ2, 20x, NA = 0.25). During the experiments, the tumor spheroid is embedded in a physiological water solution and kept at room temperature. The morphological structure of the quenched tumor spheroid, ~450 µm sized, has been fixed during growth, which makes the biological sample optically stationary over the measurement time (several minutes, see Supplementary Fig. 8). A CW near-infrared laser source (λ = 1064 nm, tunable from 0.1 to 3 W maximum power) is used as a pump beam to induce cancer morphodynamics (laser-induced hyperthermia) by means of mechanical deformations mediated by the optical absorption of water molecules. Thermo-optical effects are in fact the principal source of local variations of the index of refraction profile on the micrometric scale in the visible range, dominating over other photo-induced effects. The entire specimen, consisting of the 3DTM embedded in ~100 mm3 physiological solution, undergoes temperature variations in the interval 1–20 °C ( ± 3 °C, measured with a custom-made system employing a IR temperature sensor) above room temperature when the pump power is varied. The structural changes induced by the IR laser are irreversible except that for low pump powers (<0.2 W). Once the pumping conditions are fixed, the temporal features of the dynamics are repeatable.
Electronic equipment
The morphodynamic processes we probe, from hyperthermia to chemotherapeutics effects, occur on a time scale ranging from 102 to 105 s, where ultra-fast optoelectronic feedback is not necessary. In our setting, the overall iteration time is ~0.2 s, this indicating that dynamic biological processes in the living tumor spheroids do not considerably affect light scattering on smaller time scales. We employ a high-speed SLM with a maximum refresh frame rate of 180 Hz (response time up to 3 ms) connected to a PC via DVI interface and programmed using MATLAB. The iteration time is thus set by the CCD frame rate (maximum 60 fps) at exposure times of tens of milliseconds. However, it can be lowered to a few milliseconds by different technologies for light modulation and data transfer and processing.
Training method
The generic input state is a combination of the N optical modes with arbitrary phases from 0 to 2π and gives a low-intensity speckle-pattern as output (Fig. 3). Training of the optical random neural network is achieved by a feed-forward algorithm that minimizes a cost function expressed in terms of a target local intensity distribution in the transmission plane (output layer). Specifically, at each iteration of the algorithm, we randomly select a cluster of input modes and adjust the corresponding phases with a π/5 resolution on the SLM; the change is stored only if the cost function decreases, that is, if the intensity detected on specific groups of the CCD pixel (target) increases. As the algorithm converges, random clusters of decreasing size are selected to avoid trapping into local computational minima. After the training stage (in Fig. 3b we limit to ~103 iterations), the input layer encodes the proper distribution of modes (input weights) that mixed by the deep multicellular layers performs the specific operation. These photonic input-output functions are extremely sensitive to changes in the reservoir structure (the multicellular network of the 3DTM) and can be exploited for monitoring and sensing its morphology. When the ROM operates with feedback on, this training algorithm runs while the spheroid changes morphology over time, adapting the input phases on the SLM to maintain the output target. The Supplementary Note 5 reports further information on the training algorithm. The persistence time in Fig. 7 for both CTRL and CIS is normalized to the initial value for the sake of comparison. The initial value of TP for CTRL is 52 s, and for CIS is 28 s.
Confocal microscopy imaging and analysis
To verify if light treatment impaired spheroid structure, the hyperthermia experiment was performed on fixed spheroids stained with 4',6-diamidine-2-fenilindolo (DAPI, Sigma Aldrich) under a Confocal Microscope (Nikon A1 MP). z-stacks of each spheroid have been acquired (Δy = 1 µm) and analyzed with FIJI software43. A set of nuclei morphology describers have been retrieved from confocal images, i.e., nearest neighbor distance, area, circularity (4π × Area/Perimeter2), the aspect ratio of the nucleus fitted ellipse (major axis/minor axis) ranges from 0 (infinitely elongated polygon) to 1 (perfect circle). The aspect ratio of the particle’s fitted ellipse (Major Axis/Minor Axis), and solidity (Area/convex area). Labeling of actin cytoskeleton has been obtained by Alexa Fluor 647 phalloidin (Life Technologies), according to the manufacturer protocol. In Fig. 6a, optical microscopy is performed without any fluorescent dye to avoid photobleaching on long times. Histograms in Fig. 6b, c have been calculated by using ImageJ, function Analyze > Histogram. Pixel intensity in 8-bit images goes from 0 (dark pixels inside spheroids) to 255 (white pixels of transmitted light of background).
Statistical analysis
Each experiment has been repeated on biological replicates and errors in Figs. 6 and 7 are given by three replicates. Each microscopy images is measured over three technical replicates to average on intensity fluctuations. In conventional optical characterization methods, data statistics were analyzed by calculating the t-test between two groups, and one-way analysis of variance (ANOVA) for multiple groups with post-hoc Bonferroni test (GraphPad Prism v7, GraphPad Software Inc.). Unless otherwise noted, all results were expressed as the mean ± s.d. A value of p < 0.05 was considered statistically significant. Power analysis was not conducted to determine sample size, and investigators were not blinded.
Metabolism measurement
CellTiter-Blue® Cell Viability Assay (Promega) was used to assess cell viability of spheroids after chemotherapeutic treatment. Twenty microliters of a reagent to each 100 μl of the medium in the 96-well containing spheroids was used. A blank well (medium and drug alone) was prepared to subtract the background. This assay allows prolonged incubation time without affecting cell viability; we optimized the incubation time to 3 h to allow deep penetration of the reagent in the spheroid core. Experiments have been done in triplicate. This assay is based on the ability of living cells to convert a dye (resazurin) into a fluorescent product (resorufin) emitting at 590 nm. Nonviable cells rapidly lose metabolic capacity and thus do not generate the signal. Resazurin can penetrate cells, where it becomes reduced to the resorufin, as a result of the action of several different enzymes. The fluorescent resorufin dye can then diffuse out of cells and into the surrounding medium. The metabolism in Figs. 6 and 7 is normalized to the initial values for both CTRL and CIS for the sake of comparison. The initial fluorescence values are for CTRL is 42, and for the CIS is 34 in arbitrary units.
Data availability
The data that support the plots and other findings of this paper are available from the corresponding author upon reasonable request.
Code availability
The codes are available from the corresponding author upon reasonable request.
References
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Russell, S. J. & Norvig, P. Artificial Intelligence: A Modern Approach (Prentice Hall, 1995).
Schmidhuber, J. Deep learning in neural networks: an overview. Neural Netw. 61, 85–117 (2015).
Verstraeten, D., Schrauwen, B., D’Haene, M. & Stroobandt, D. An experimental unification of reservoir computing methods. Neural Netw. 20, 391–403 (2007).
Lukoševičius, M. & Jaeger, H. Reservoir computing approaches to recurrent neural network training. Computer Sci. Rev. 3, 127–149 (2009).
Huang, G.-B., Wang, D. H. & Lan, Y. Extreme learning machines: a survey. Int. J. Mach. Learn. Cyber 2, 107 (2011).
Gallicchio, C., Micheli, A. & Pedrelli, L. Design of deep echo state networks. Neural Netw. 108, 33–47 (2018).
Huisman, S. R., Huisman, T. J., Wolterink, T. A. W., Mosk, A. P. & Pinkse, P. W. H. Programmable multiport optical circuits in opaque scattering materials. Opt. Express 23, 3102–3116 (2015).
Matthès, M. W., del Hougne, P., de Rosny, J., Lerosey, G. & Popoff, S. M. Optical complex media as universal reconfigurable linear operators. Optica 6, 465–472 (2019).
Leedumrongwatthanakun, S. et al. Programmable linear quantum networks with a multimode fiber. Nat. Photon. 14, 139–142 (2020).
Marcucci, G., Pierangeli, D., Pinkse, P., Malik, M. & Conti, C. Programming multi-level quantum gates in disordered computing reservoirs via machine learning. Opt. Express 28, 14018–14027 (2020).
Dong, J., Gigan, S., Krzakala, F. & Wainrib, G. Scaling up Echo-State Networks with multiple light scattering. 2018 IEEE Statistical Signal Processing Workshop (SSP) 448–452 (2018).
Saade, A. et al. Random projections through multiple optical scattering: approximating kernels at the speed of light. 2016 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) 6215–6219 (2016).
Miller, D. A. B. Waves, modes, communications, and optics: a tutorial. Adv. Opt. Photon. 11, 679–825 (2019).
Duport, F., Schneider, B., Smerieri, A., Haelterman, M. & Massar, S. All-optical reservoir computing. Opt. Express 20, 22783–22795 (2012).
Lin, X. et al. All-optical machine learning using diffractive deep neural networks. Science 361, 1004–1008 (2018).
Shen, Y. et al. Deep learning with coherent nanophotonic circuits. Nat. Photon. 11, 441–446 (2017).
Larger, L. et al. High-speed photonic reservoir computing using a time-delay-based architecture: million words per second classification. Phys. Rev. X 7, 011015 (2017).
Zhao, P. et al. Universal linear optical operations on discrete phase-coherent spatial modes with a fixed and non-cascaded setup. J. Opt. 21, 104003 (2019).
Ghofraniha, N. et al. Experimental evidence of replica symmetry breaking in random lasers. Nat. Commun. 6, 6058 (2015).
Pierangeli, D. et al. Observation of replica symmetry breaking in disordered nonlinear wave propagation. Nat. Commun. 8, 01612 (2017).
Inagaki, T. et al. A coherent Ising machine for 2000-node optimization problems. Science 354, 603 (2016).
Marandi, A. et al. Network of time-multiplexed optical parametric oscillators as a coherent Ising machine. Nat. Photon. 8, 937–942 (2014).
Pierangeli, D., Marcucci, G. & Conti, C. Large-Scale Photonic Ising Machine by Spatial Light Modulation. Phys. Rev. Lett. 122, 213902 (2019).
Prabhu, M. et al. Accelerating recurrent Ising machines in photonic integrated circuits. Optica 7, 551 (2020).
Horstmeyer, R., Ruan, H. & Yang, C. Guidestar-assisted wavefront-shaping methods for focusing light into biological tissue. Nat. Photon. 9, 563 (2015).
Tamura, M. et al. Morphology-based optical separation of subpopulations from a heterogeneous murine breast cancer cell line. PLoS ONE 12, e0179372 (2017).
Yuan, Y. et al. Quantitative image analysis of cellular heterogeneity in breast tumors complements genomic profiling. Sci. Transl. Med. 4, 157ra143–157ra143 (2012).
Moriconi, C. et al. INSIDIA: a FIJI macro delivering high-throughput and high-content spheroid invasion analysis. Biotechnol. J. 12, 1700140 (2017).
Mosk, A. P., Lagendijk, A., Lerosey, G. & Fink, M. Controlling waves in space and time for imaging and focusing in complex media. Nat. Photon. 6, 283 (2012).
Rotter, S. & Gigan, S. Light fields in complex media: mesoscopic scattering meets wave control. Rev. Mod. Phys. 89, 015005 (2017).
Tait, A. N. et al. Neuromorphic photonic networks using silicon photonic weight banks. Sci. Rep. 7, 7430 (2017).
Marcucci, G., Pierangeli, D. & Conti, C. Theory of neuromorphic computing by waves: machine learning by rogue waves, dispersive shocks, and solitons. Phys. Rev. Lett. 125, 093901 (2020).
Kasprowicz, R., Suman, R. & O’Toole, P. Characterising live cell behaviour: traditional label-free and quantitative phase imaging approaches. Int. J. Biochem. Cell Biol. 84, 89–95 (2017).
Mittal, S. A survey of FPGA-based accelerators for convolutional neural networks. Neural Comput. Appl. 32, 1109–1139 (2020).
Nourhashemi, M., Mahmoudzadeh, M. & Wallois, F. Thermal impact of near-infrared laser in advanced noninvasive optical brain imaging. Neurophotonics 3, 015001 (2016).
Scholkmann, F. et al. A review on continuous wave functional near-infrared spectroscopy and imaging instrumentation and methodology. Neuroimage 85, 6 (2014).
Chan, C. J., Li, W., Cojoc, G. & Guck, J. Volume transitions of isolated cell nuclei induced by rapid temperature increase. Biophys. J. 112, 1063–1076 (2017).
Fleming, A., Conti, C. & Di Falco, A. Perturbation of transmission matrices in nonlinear random media. Annalen der Physik 19, 1900091 (2019).
Pascanu, R., Gulcehre, C., Cho, K. & Bengio, Y. How to construct deep recurrent neural networks. In Proceedings of the Second International Conference on Learning Representations (ICLR, 2014).
Schürmann, M. et al. Three-dimensional correlative single-cell imaging utilizing fluorescence and refractive index tomography. J. Biophotonics 11, e201700145 (2018).
Jiang, P. et al. Novel anti-glioblastoma agents and therapeutic combinations identified from a collection of FDA approved drugs. J. Transl. Med. 12, 13 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Vellekoop, I. M. & Mosk, A. P. Phase control algorithms for focusing light through turbid media. Opt. Commun. 281, 3071–3080 (2008).
Antonik, P., Marsal, N., Brunner, D. & Rontani, D. Human action recognition with a large-scale brain-inspired photonic computer. Nat. Mach. Intell. 1, 530–537 (2019).
Acknowledgements
We acknowledge support from the PRIN PELM (20177PSCKT). We thank MD Deen Islam for the assistance in the laboratory.
Author information
Authors and Affiliations
Contributions
C.C. and M.P.: conceptualization and supervision, ideation of experiments, methodology, project administration. D.P.: optical setup realization, investigation by experiments, and data analysis. V.P.: resources and investigation by sample fabrication, imaging, and biological assays, validation. G.M. and C.C.: theoretical model. C.C., G.M., and M.D.S.: methodology and theoretical analysis. C.M. and G.P.: resources by providing samples and expertize on tumor models. All the authors contributed to the writing of the original draft, review, and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pierangeli, D., Palmieri, V., Marcucci, G. et al. Living optical random neural network with three dimensional tumor spheroids for cancer morphodynamics. Commun Phys 3, 160 (2020). https://doi.org/10.1038/s42005-020-00428-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-020-00428-9
This article is cited by
-
Carboxylated graphene quantum dots-mediated photothermal therapy enhances drug-membrane permeability, ROS production, and the immune system recruitment on 3D glioblastoma models
Cancer Nanotechnology (2023)
-
Extreme transport of light in spheroids of tumor cells
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.