Abstract
Cell movement in vivo is typically characterized by strong confinement and heterogeneous, three-dimensional environments. Such external constraints on cell motility are known to play important roles in many vital processes e.g. during development, differentiation, and the immune response, as well as in pathologies like cancer metastasis. Here we develop a physics-driven three-dimensional computational modeling framework that describes lamellipodium-based motion of cells in arbitrarily shaped and topographically structured surroundings. We use it to investigate the primary in vitro model scenarios currently studied experimentally: motion in vertical confinement, confinement in microchannels, as well as motion on fibers and on imposed modulations of surface topography. We find that confinement, substrate curvature and topography modulate the cell’s speed, shape and actin organization and can induce changes in the direction of motion along axes defined by the constraints. Our model serves as a benchmark to systematically explore lamellipodium-based motility and its interaction with the environment.
Similar content being viewed by others
Introduction
The motility of living cells and tissue is an equally important topic for biology and non-equilibrium physics. Individual and collective cellular migration are examples of a self-organized non-equilibrium state characterized by the transduction of chemical energy delivered by the metabolism into directed mechanical motion. Recent studies demonstrated that several aspects of the seemingly complex dynamics of migrating cells and tissues can be rationalized in the framework of relatively simple physics-driven models1,2,3,4.
Due to the relative ease in observation, data analysis, and modeling, quantitative physics-driven studies of cell motion have in the past focused almost exclusively on cells crawling along flat substrates5. This has been often criticized by the biology community6, since in vivo cells never live in such an idealized world. In contrast, cells migrate in rather complex three-dimensional (3D) environments that in addition are strongly inhomogeneous and also reorganized by both forces and chemical action from the cells. For instance, both leukocytes7 and metastatic cancer cells8 move along or within blood vessels until they reach a target site, where they need to squeeze through epithelial layers, or extracellular matrix (ECM). These examples already involve a plethora of topographical and confining features, namely motion on curved substrates, in tubular and planar confinement, on fibers or fiber bundles, as well as through random 3D fiber networks.
A realistic in vivo situation, e.g., the movement of cancer cells inside living tumors, is prohibitively difficult to investigate, although progress has been achieved using novel microscopy techniques9. Consequently, it is important to study well-defined scenarios in a systematic manner, with the goal to deduce general guiding principles. In the last years, experimental studies on cells in well-defined artificial or reconstituted 3D migration assays have gained pace, see Paul et al.10 for a recent review and Maiuri et al.11 for a comparison of different confining situations. A large part of these works focuses on cells in reconstituted networks of collagen or synthetic hydrogels12,13,14. In addition, well-defined, microfabricated systems have been developed and employed to study confined cell motion, like microchannels15,16 and vertical confinement in slit-like geometries17,18. Substrates with topographical features have a longer tradition in tissue engineering19,20, and are now also employed to study motility21,22,23,24,25,26, as is the motion on micro- or nano-scale fibers27,28,29. Although a direct mapping from these model systems to the full in vivo case is in no way direct or obvious, these systems are in well-controlled conditions that allow to isolate different generic features of topography and/or confinement and their influence on cell motion.
On planar substrates, most cells move by forming lamellipodia, thin sheets containing actin filaments that polymerize and branch towards the cell membrane, thereby exerting protrusion forces30,31. A long-standing question had been whether these motile structures dominating 2D motion also exist and are relevant in 3D. There is growing evidence that this is indeed the case for fibroblasts32 and neutrophils33, at least under certain conditions. Therefore, we here develop a physics-based modeling framework that transfers the knowledge of 2D lamellipodia-based cell motion2,34,35 to 3D settings. The model is kept as simple as possible. It nevertheless is able to reproduce a plethora of the phenomena of single cell movement in confinement and on curved or topographically structured substrates found in the literature and puts them into a common framework. In fact, compared to earlier 3D models that exclusively studied flat substrates1,36,37,38 (with the exception of ref. 39 on fiber networks) the elegant use of the phase field approach40 – at the same time for the cell and for the surroundings it interacts with – allows to model motion in, in principle, arbitrarily shaped and topologically non-trivial environments.
Results
Computational model
We first briefly describe the modeling framework. The developed model is a non-trivial 3D extension of the 2D phase field approach presented in ref. 2, known to describe the subcritical onset of cellular motion41 and realistic cell shapes, with model extensions being able to describe guided motion on inhomogeneous substrates42,43 and lamellipodial waves44. The approach is conceptually simple as it has only two dynamic variables: first, the cell’s interface is described by the transition region of a dynamic 3D scalar phase field ρ(r, t) ϵ [0, 1] for a closed domain representing the cell. The phase field value ρ = 1 corresponds to the cell’s interior, ρ = 0 to the outside, and the ρ = 1/2 isosurface is identified with the cell membrane. And second, inside this domain, a 3D vector field p(r, t) describes the mean orientation of actin filaments and the degree of their parallel ordering (direction and absolute value of p, respectively). Arbitrary substrates and confinement scenarios can be modeled by specifying two additional, static phase fields: Φ(r) describing steric exclusion and Ψ(r) restricting the actin generation to regions close to the substrate. They have the same overall shape but the characteristic decay length of the latter is chosen to be larger, to allow for substantial actin inside the cell.
The cellular motility machinery is implemented via its basic physical features in the dynamics of ρ(r, t) and p(r, t): actin is generated close to the membrane (i.e., the phase field interface) and induces a protrusion force via the polymerization ratchet mechanism45, leading to an effective interface advection rate. Together with the cellular adhesion to the substrate, and assuming that actin polymerization is localized close to the substrate and predominantly tangential to the local substrate plane, the cell then spreads and flattens. Myosin motor-induced contraction then induces a symmetry breaking46 and overall cell polarity41, with a region of high p at the front identified as the lamellipodium, and the cell moves with constant speed, see Supplementary Movie 1 for a typical polarization event.
Specifically, the 3D model equations read
In Eq. (1), Dρ determines the width of the diffuse interface (and the cell’s surface tension47, see also ref. 48 for modeling more realistic membrane features). The second term implements relaxational dynamics in the phase field potential \(F[\rho ] = {\int} f(\rho )d^3{\mathbf{r}}\), where \(f(\rho ) = {\int}_0^\rho \rho \prime (1 - \rho \prime )(\delta - \rho \prime )d\rho \prime\). Since the order parameter dynamics is non-conserved (see e.g. the discussion in ref. 40), it needs to be supplemented by a volume conservation. We chose a simple implementation via a global constraint,
Here \(\frac{1}{2}\) is the stationary point, μ the constraint’s stiffness and the term in brackets is the difference between the cell’s current volume and the prescribed (initial) volume V0. The term α p⋅ ∇ρ accounts for the pushing force exerted by actin ratcheting against the cell’s boundary, with α an effective velocity (i.e., force over a friction coefficient with the substrate). Adhesion is implemented on a coarse-grained level49,50 via the term κ∇Φ ⋅ ∇ρ with an adhesion strength parameter κ, but explicit adhesive dynamics can in principle be introduced42,51. The last term describes excluded volume interaction, with a strength λ, via the potential \(\frac{\lambda }{2}{\int} \rho ^2{\mathrm{\Phi }}^2{\kern 1pt} d^3{\mathbf{r}}\). More information on the last two contributions can be found in refs. 47,50.
The polarization dynamics has three contributions from actin turnover, see the first line of Eq. (2): a diffusion (or elastic) term, a source term proportional to the parameter β (related to the actin polymerization velocity) and discussed in more detail below, and a sink term describing degradation of actin, e.g., via depolymerization, with time scale τ being of the order of the actin turnover time (typically seconds). For the cell to be able to polarize and move, the front-back symmetry has to be broken. Two mechanisms, based on key experimental observations46 and associated to the action of myosin motors in the actin network, were already implemented in ref. 2: a term σ|p|2 added to the parameter δ in Eq. (1), that accounts for acto-myosin contraction with contraction rate σ and that can be motivated by active gel theory52. And a second term, γ(∇ρ ⋅ p)p, that models the presence of an anti-parallel acto-myosin bundle at the cell’s rear2,41 and explicitly breaks the front-back symmetry. Finally, the last term in Eq. (2) explicitly suppresses p inside the substrate.
Let us now discuss the source term of actin polarization, i.e., the second term in Eq. (2). In lamellipodia, actin nucleation is known to occur predominantly at the cell membrane, where regulators such as Wiskott-Aldrich Syndrome protein (WASP) and Arp2/3 are located30,31. This effect was modeled in ref. 2 via localizing the actin source term at the cell’s interface by using the phase field gradient ∇ρ. However, an important issue arises in modeling 3D cell movement: while the projected 2D area of spreading and moving cells has been studied extensively53, and has been explained by the prevalent forces54 (involving protrusion, adhesion, traction, and membrane bending), it is to date unknown what determines the thickness of the lamellipodium, which typically always lies in the range 100–400 nm. Although there exist some hypotheses, e.g., involving curvature-sensitive membrane-embedded regulators of actin nucleation55, the mechanisms remain speculative.
On the level of phenomenological description, we hence employed the following strategy: actin polymerization is modeled anisotropically with respect to the local tangential plane defined by the substrate. We introduce the (tensorial) projection operator onto the local tangential plane, \(\hat P = \hat I - \widehat {\mathbf{n}} \otimes \widehat {\mathbf{n}}\) with \(\hat I\) the identity and \(\widehat {\mathbf{n}} = \nabla \Phi /|\nabla \Phi |\) the normal vector to the substrate. \(\hat P\widehat {\mathbf{n}} = 0\) holds and hence the term \(\beta [(1 - \theta )\hat P(\nabla \rho ) + \theta \nabla \rho ]\) in Eq. (1) implements a polymerization rate of β in the tangential plane and of βθ normal to the substrate. The parameter θ ϵ [0, 1] hence governs the degree of this anisotropy, within the two limits of θ = 0 for polymerization exclusively in the tangential plane and θ = 1 for the isotropic case. The second main addition compared to earlier 2D models is the assumption that the actin polymerization rate decays with the distance from the substrate. This is implemented via the phase field Ψ(r), and enters the actin source term as a common factor. It is motivated by the fact that actin polymerization relies on regulatory processes close to the membrane, and has already been employed in a similar form in ref. 1.
Let us briefly discuss the relevant scales and typical parameters. Eqs. (1) and (2) are already written in rescaled units, as discussed in ref. 2. We use 1 μm and 10 s as the typical length and time scale, since then all activity-related parameters are of order one. Namely, the typical actin depolymerization rate of ≈1s−1 and maximum actin polymerization velocity ≈0.1–0.3 μms−1, i.e., ~20–60 monomers per second, amount to the used values of 1/τ1 = 0.1 and β = 1–3. We chose α = β/3 since the cell’s interface movement should be smaller then the bare polymerization velocity – not all polymerization events create propulsion, and there is friction. Typical myosin motor speeds are of order 0.1–0.2 μms−1, implying σ and γ of order one. Note that the maximum center-of-mass (c.o.m.) velocities of the modeled cells – being a model outcome – are of order one and hence correspond to the typical maximum speed of keratocytes (0.1 μms−1)45. Typical parameter values are listed in the Supplementary Table 1.
In the following we mainly discuss two distinct cell types: first, ideal “lamellipodial-type” cells, whose polymerization kinetics lies perfectly in the tangential plane defined locally by the substrate (θ = 0). This type corresponds best to the well-studied keratocytes, for which it has been shown that the actin network in the lamellipodium is very close to two-dimensional56. With sufficiently high adhesion strength to ensure the cell’s contact with the substrate, both stationary and motile states can be obtained when varying the remaining actin- (α, β) and myosin-related (γ, σ) parameters. A cell of this type moving steadily on a planar susbstrate is exemplified in Fig. 1a. The second cell type is refered to as of (partial) “wall pushing-type”, having finite θ. For this type, actin generated perpendicularly to the substrate is non-negligible, leading to stronger contact of the cell with the substrate, even in the absence of strong adhesion. Note that a somewhat similar motility mode was considered in ref. 57. Again the system is bistable with both stationary and motile states. We however found that for the same acto-myosin parameters wall-pushing cells are less motile.
Multiplicity of cell migration modes in 3D surroundings. a A 3D cell migrating on a flat substrate. Shown is the ρ = 1/2 isosurface of the cell’s phase field in green (and of the substrate’s in yellow), the distribution of the actin polarization field |p| as color-coded arrows and the direction of motion as the green arrow. b Vertical confinement, i.e. motion in a slit constrained by two close-by walls. Cells show bistable/hysteretic behavior: for same parameters but depending on the initial conditions (or history), cells can attach to only one of the walls or to both, leading to different shapes and velocities. c Cell guidance by slit width: shown are two different cells, one of pure lamellipodium-type (green) and one of wall pushing-type (red), see the main text. Upon encountering a modulation in slit width, the former cell is deflected to thinner gaps while the latter prefers wider ones. Color curves correspond to trajectories (to fit both cells in the same picture, the red cell was moving double the time than the green one). d A cell moving inside a tube (a microchannel with circular cross-section), attaining a bullet-like shape. e Both the cell’s velocity and shape strongly depend on the curvature of the substrate it crawls on. f A cell spreading on (upper image), wrapping around and finally moving along (lower image) a thin fiber. See also Supplementary Movies 1–8 for the dynamics in the different scenarios
Inspired by the plethora of recent experiments15,16,17,18,21,22,23,24,25,26,27,28,29, we applied our 3D model to study cell migration in confinement (slits and channels), its sensitivity to substrate curvature including motion on fibers, as well as motion on planar substrates with topographical features, see Fig. 1b–f for a summary.
Motion in vertical confinement
We first considered the generic scenario of vertical confinement, i.e., a slit geometry, where the cell is sandwiched between two co-planar walls17 and can accommodate its shape and migration mode only in the quasi-2D plane. For the upper wall, we investigated two different types: either it has properties identical to the lower substrate, i.e., the cell can adhere, spread and form lamellipodia (“adhesive wall”). Alternatively, the upper wall is inert and just presents a steric obstacle for the cell to which it can neither adhere nor form a lamellipodium (“passive wall”). The latter can be easily modeled by putting κ = 0 there and by attributing to the wall only a steric field Φ and no actin regulatory field Ψ.
We investigated confined cell motion using the following generic preparation protocol: a top wall (either active or passive) was approached from above to a cell moving on the lower substrate gradually, in a step-by-step fashion, letting the moving cell accommodate to the new environment before approaching further. To test whether the effect of the confinement is reversible, after a strong confinement state had been reached, the upper wall was retracted in a similar step-by-step fashion. This protocol is a generalization of the classical cell compression experiment58 to moving cells and could be experimentally realized employing micromanipulation techniques, e.g. involving a magnetic upper wall.
The most interesting scenario appears for strongly driven cells, as exemplified for the lamellipodium-type cell in a symmetric slit (i.e., with adhesive top wall) by the snapshots shown in Fig. 2a corresponding to the red curves in Fig. 2b, see also Supplementary Movie 3. In this case, there is a window of gap widths exhibiting a bistability of motile modes: for the same parameters, the cell can attach to and form lamellipodia on both walls equally, see the blue cell in Fig. 2a, with the cell speed increasing with decreasing the slit width. Alternatively, the cell can attach only to one of the walls (the lower one, as the top wall was approached), see the green cell in Fig. 2a and moves with a speed that is independent of gap width and higher than for a cell attached to both walls, due to a larger lamellipodium. Upon further decreasing the gap width, finally the cell is forced to attach to the top wall when the latter is sufficiently close. Consequently, vertical confinement constitutes a geometry-induced motility hysteresis loop: the motion of a cell in a spatially inhomogeneous vertical confinement will strongly depend on the history of its path. In contrast, if the wall is passive, this behavior is absent (orange curve in Fig. 2b). Only for strong confinement the cell speeds up a little as its lamellipodium, i.e., driven area, slightly increases.
Quantification of the effects of confinement on cell motility. a Snapshot of two cells moving in a slit between two flat walls at distance d. b Analysis of the cell velocity. Two scenarios for the upper wall were considered: either its properties are identical to the lower wall, i.e. cells can adhere and form lamellipodia (“adhesive wall”, as shown in a), or it is inert (“passive wall”, only steric exclusion). For large gap widths the cell attaches only to one wall and is not influenced by the confinement. For small gap widths d and adhesive walls (red and blue curves), cells attach to both walls, with the speed increasing with increase of confinement. Interestingly, in the case of strongly driven cells and adhesive top wall, there is a hysteresis loop (red curves). For the passive wall, this behavior is absent (orange curve). Less-driven cells can be stopped when approaching the upper wall (blue curve). Parameters: α = 1.5, β = 4.5, γ = 0.4, σ = 0.6 for all cells. Red and orange curves correspond to lamellipodium-type cells with θ = 0, κ = 8; blue curve to a wall-pushing cell (effectively less driven) with θ = 0.2, κ = 1; other parameters see Supplementary Table 1
Wall-pushing cells show a similar overall behavior in vertical confinement as lamellipodium-type cells. For the same driving parameters as the respective lamellipodium-type cells, they are however effectively less driven and hence less motile. Instead of exhibiting hysteresis, sufficiently weakly driven cells rather get stopped when starting to feel the upper wall, see the blue curve in Fig. 2b. Upon further decrease in gap width, they can be induced to migrate again. Apparently, for intermediate gap widths, the upper wall represents a perturbation inducing a shape change towards the third dimension (i.e., the thickness of the cell) that interferes with the in-plane dimension of the cell and renders it less motile. In turn, sufficiently thin slits squeeze the cell, and they become more motile than on a single flat substrate: they can move for lower values of the actin-associated driving since the strong confinement enlarges the region of adhesion, the lamellipodium and hence force generation. This can be further exemplified by studying cells on a substrate below their threshold of motion. Slowly approaching a second wall sets them into motion (see Supplementary Fig. 1).
We also studied briefly the possibility of guiding cells by spatially modulated slit widths, as shown in Fig. 1c. There, two different cells, one of lamellipodium-type (green) and one of wall-pushing type (red; both weakly driven), were steadily moving and adhering to both plates, before encountering a smooth variation in the distance of the walls, orthogonal to the direction of motion. Interestingly, the lamellipodium-type cell is deflected to thinner gaps while the wall-pushing one prefers wider gaps. The former is easily understandable: the part of the cell in the narrower region broadens the active front while the one in the wider region reduces, hence the cell turns. The latter is more subtle: it appears that the wall-pushing cell is somewhat pinned by the gap gradient and that the broader lamellipodium on the thinner side runs around this pinning point, leading effectively to a turn towards the broader gap width, see Supplementary Movie 4 for details.
Effects of substrate curvature and cylindrical confinement
We next quantified the motion of cells on curved substrates exhibiting a single curvature direction (i.e., cylindrical symmetry), including confinement in a tube and motion on a fiber. We later also briefly discuss effects of Gaussian curvature. The signed substrate curvature c = 1/R (with R the signed radius of curvature) is defined as sketched in Fig. 3a: a cell moving on the outside of a cylinder surface along the main axis corresponds to negative and a cell moving inside a cylindrical tube to positive curvature.
Quantification of the effects of substrate curvature on cell motility. a Definition of signed substrate curvature. b Snapshots of cell shapes moving on substrates of various curvatures. c Cell velocity of four different cells as a function of the substrate curvature (1/R with R the radius of curvature). The vertical black dashed line marks the flat substrate. Lamellipodium- and wall-pushing cells were studied, for two different driving strengths each. For all cell types, migration is stalled by strongly negative curvatures (cells driven more strongly continue to move on more strongly negative curvatures). Close to this stalling curvature, cells elongate in the direction of motion. The cell speed increases with the signed curvature, passing by the flat state. The black straight line is a geometric scaling approximation, V(R) = V0[1 + h/(2R)], explaining the approximately linear behavior for small curvatures (see text). In the case of lamellipodium-type cells, at a certain positive curvature, a competition between the perturbed crescent-shape and a bullet-like shape with cylindrical symmetry sets in, leading to complex dynamics, namely shape and position oscillations inside the tube. Finally the bullet shape stabilizes for even thinner tubes. Wall-pushing (and hence less driven) cells become typically stopped in this region of competition, before they attain similar bullet shapes at higher substrate curvatures
Figure 3b displays snapshots of the respective cell shapes, clearly showing that they are severely perturbed by the imposed curvature. This is also reflected in the cell’s center-of-mass velocity along the cylinder axis, as quantified in Fig. 3c: Inside a tube (right part), cells move faster than on the flat substrate. In contrast, on the outer cylinder surface they move slower and finally get stopped (“stalled”) for a critical cylinder radius that depends on the driving force, i.e., on the acto-myosin parameters.
Overall, cell speed increases with signed substrate curvature, passing by the flat state. The linear dependence of the migration speed V on the substrate curvature 1/R can be anticipated from the following geometric arguments. Let us assume that the migration speed V0 on the flat substrate depends linearly on the width of the active lamellipodium front W, since the propulsion force depends on the total number of pushing actin filaments. The height h of the cell does not change upon slightly curving the substrate, as verified numerically. Assuming further that the length of the cell (in the direction of motion) remains approximately constant, volume conservation then implies that the cell’s cross-sectional area (normal to the direction of motion) does not change. Let us approximate for simplicity the cell’s cross-section by a rectangle, S = Wh. On a substrate with positive radius of curvature R, i.e., for a cell inside a channel, this rectangle transforms into a cylindrical shell segment of cross-section \(\tilde S = \tilde W/R[R^2/2 - (R - h)^2/2] = \tilde Wh[1 - h/(2R)]\). Using \(S = \tilde S\), and solving for \(\tilde W\) for \(R \gg 1\) we obtain \(\bar W = W[1 + h/(2R)]\) and hence the migration speed can be given as
This relation is in excellent agreement with the simulations, as exemplified in Fig. 3c for the weakly driven lamellipodium-type cell by the black line, using the measured velocity and height of the cell on the flat substrate.
For sufficiently high positive curvature, corresponding to thin capillaries, cells adhere and form lamellipodia all around. This corresponds to the cylindrical confinement inside a microchannel, restricting the motion to quasi-1D as studied e.g. in refs. 15,16. In the case of lamellipodium-type cells, at a certain positive radius of curvature a competition starts between the perturbed crescent-shape states, as observed for lower curvature, and bullet-like shapes with cylindrical symmetry. This competition induces more complex dynamics, as discussed below. Finally, for even thinner tubes, the bullet-shape wins and the cell’s speed is increasing again with signed curvature. Wall-pushing cells also switch from crescent-like to bullet shape, close to the transition, however, they typically become arrested.
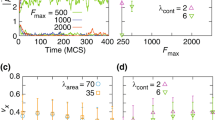
In the regime of complex dynamics marked in Fig. 3c, the cell apparently cannot decide which shape to adopt – the perturbed crescent or the bullet. Consequently, the cells do not move on average, but as they remain active they induce shape and position oscillations inside the tube. In Fig. 4a the center of mass position in z-direction together with a measure for the axial length of the cell were tracked. The former oscillates, corresponding to alternating crescent shapes attached to the top and the bottom of the channel. In between these extremes, however, the two wings of the crescent cells become unstable and fuse to a more bullet shape, giving rise to double period oscillations of the axial extension of the cell. This can be understood from the shapes during half a period shown in Fig. 4b. The detailed dynamics is sensitive to the specific initial conditions. The fact that the weaker driven wall-pushing cell did not display oscillations hints at the existence of a threshold value of activity for the phenomenon. For longer times even more complex oscillations can emerge, involving corkscrew-type deformations, see Fig. 4c.
Complex dynamics inside narrow channels. a Quantification of the complex dynamics of lamellipodium-type cells by tracking the z-coordinate of the center of mass position (black) and the axial extension of the cell (along the channel; red). Both display oscillations, corresponding to the shape changes displayed in b, due to the competition between a deformed crescent and a bullet-like shape, resulting in no overall net motion. c A more complex corkscrew-type of motion in this parameter regime, developing from the one shown in b for longer times. See also Supplementary Movie 7
Motion on fibers
Coming back to negative signed curvatures, we also studied cell motion on fibers, as experimentally studied for instance in refs. 27,28. As already shown in Fig. 3, cells can move on fibers with radii larger than their unperturbed size. They can even wrap around sufficiently thin fibers, see Fig. 1f, and move if perturbed by adding a small initial polarization. Interestingly, we found a generic alignment effect: cells initially polarized obliquely to the fiber axis tend to align their direction of motion with the axis. To obtain quantitative insight, we studied the alignment time τ as a function of the fiber radius: cells were first allowed to spread on fibers of different radii and then motion was initiated with a substantial off-axis component (typically 50% of the final axial velocity). Figure 5 displays results for wall pushing-type cells. The alignment rate (inverse alignment time; obtained by exponential fits to the off-axis velocity component) is roughly linear in fiber curvature for the currently accessible range of curvatures – note that a strict validation down to c → 0 would need excessively large simulation boxes. For the highest curvature, c = 1/R = −0.06, the found value of τ ~ 800 roughly corresponds to the time the cell needs to move 20–30 times its own size.
Alignment of cells crawling on fibers. Cells were allowed to spread on cylindrical fibers with different radii and then initially polarized obliquely to the fiber axis (left inset). While circling around the fiber, the cells gradually aligned their motion to the axis. The main plot shows the rate of alignment, i.e., the inverse alignment time, as a function of fiber curvature. The right inset shows a late stage, with almost complete alignment
Effects of Gaussian curvature
We next studied the effect of a finite Gaussian curvature \(c_G = \frac{1}{{R_1}} \cdot \frac{1}{{R_2}}\), i.e., of substrates displaying two non-zero principal curvatures 1/R1, 1/R2. The simplest case is a sphere (having R1 = R2 and cG > 0), whereon cells are found to circle around with constant velocity. A more interesting case is the “sphere-with-skirt” structure recently proposed in ref. 26, which we studied by depositing cells on the top of the structure (cG > 0, Fig. 6a), on the skirt (cG < 0, Fig. 6b), as well as on the transition region (Fig. 6c). Experimentally it was found that cells can not spread on the top, an effect attributed to stress fibers, which are stiff fibers constituted of acto-myosin. Instead they were found to spread preferentially in the region where cG changes from positive to negative and to circle around the structure. The first effect is not captured by our model, due to the neglect of modeling the stiff stress fibers, i.e., cells deposited on top spread as shown in Fig. 6a and prefer to stay there upon perturbations. However, cells indeed prefer the region where cG changes and then preferentially move along the azimuthal direction, see Fig. 6c. If deposited on the region with cG < 0, cells typically are repelled and move away from the structure as shown in Fig. 6b. Nevertheless, if sufficiently polarized and heading towards the structure, they can be “captured” by and then circle around it.
Effect of Gaussian curvature on cell spreading and motility. A ‘sphere-with-skirt’ surface was studied as recently suggested26. a On the top, where cG > 0, a cell can spread but will not move if perturbed. b On the ‘skirt’, where cG < 0, cells are repelled, i.e., polarize and move away from the structure. c In fact, as in experiments, cells prefer the region in between, where they can easily spread and circle around the structure
To sum up the curvature effects, they can all be understood along the line of arguments that led to Eq. (4). Namely, with the signed substrate curvature the cell-substrate interaction increases and concomitantly also the cell’s actin polarization field is enhanced. In other words, cells spread more easily on positive curvatures (the inside of capillaries) than on negatively curved substrates (the outside of fibers), and hence the direction of motion aligns in the direction of the maximum signed curvature.
Specifically, for cells moving on the outside of fibers, the maximum curvature is directed along the principle axis (as the non-zero curvature along the circumference is negative) and hence motile cells align along that axis. For cells inside a capillary, on the other hand, the maximum curvature is along the circumference and we indeed find a preference of cells to circle around the cylinder wall. This may also provide insight into the complex dynamics regime, occurring for intermediate curvatures as shown in Figs. 3c and 4. There, two active fronts are circling against each other and if they “collide” head-on, shape oscillations are observed as shown in Fig. 4. Note that the level of driving is important again, because the (comparatively) weaker driven, wall-pushing type cells simply stop, not exhibiting shape oscillations. This is to some extent analogous to the behavior in vertical confinement, see Fig. 2b, where the two motile branches for strongly driven cells exhibit bistability while weaker driven cells are stopped. Finally, in the “sphere-with-skirt” geometry, the Gaussian curvature varies from positive (spherical cap) to negative (skirt). The skirt has one positive curvature, the other principal curvature (with respect to the symmetry axis) is negative and increases to zero with the distance from the spherical cap. Hence, there is an increase of mean curvature in the direction of the widening skirt (pointing away from the structure), explaining why cells generally move away from such structures.
Guiding cells by substrate topography
The 3D modeling framework also allows to study cell motion on essentially flat substrates exhibiting simple topographical features, complementing experimental and theoretical analyses of cell guidance on microprinted adhesive patterns42,59,60. Although there are many possible surface features, we focused here on a type that has been extensively studied experimentally21,22,23,24,25, namely periodic grooves/ridges on the substrate’s surface. Experimentally, such substrates are supposed to model the sub-cellular topography of extracellular matrix fibers, often serving as directional cues for moving cells61.
The surface pattern was created by allowing a square wave in the substrate’s phase field to equilibrate for a few time steps before fixing the resulting smoothened pattern. The amplitude and the wavelength of the pattern were varied, ranging from small to similar compared to the height and planar extension of the cell, respectively. Cells were initialized at a 45° angle to the periodic grooves and then characterized by their attained direction of motion as shown in Fig. 7a.
Cell guidance by topographical features. a A diagram demonstrating cell guidance by topographical substrate features for wall pushing-type cells. Marked is the type of alignment as a function of the amplitude and the wavelength of periodic ridges on the substrate’s surface. Cells were initially polarized at 45° angle to the preferred direction defined by the pattern. Long wavelength patterns (red circles) lead to a rapid alignment of cell motion along the ridges, as shown in b, while the cell deforms to a seal-like shape. In contrast, small wavelengths fail to align cells (black squares). In between, at intermediate wavelengths (blue triangles), cells slowly align and move perpendicularly to the ridges as shown in c. See also Supplementary Movie 9
We found that long wavelength patterns (but still of sub-cellular size) led to perfect alignment of motion of both lamellipodium- and wall pushing-type cells along the grooves, see the red circles in Fig. 7a. In addition, cells elongated on the pattern in the direction of motion, see Fig. 7b. The alignment was found to be rather insensitive to the (sub-cellular) pattern amplitude, and it was typically fast, i.e., the cell only needed to move few times its size. In contrast, patterns of smaller wavelengths led to practically no alignment for both cell types and all studied amplitudes (black squares), and also the shapes were less affected. Interestingly, in between there is a wavelength region (blue triangles) where wall pushing-type cells align, albeit very slowly, perpendicularly to the grooves as shown in Fig. 7c. Concluding, in general, cells prefer to align along grooves comparable to their size. They are “attracted” by the grooves and rather “dislike” the barriers.
Discussion
The framework developed and analyzed here is a generic physical model for lamellipodium-based motion. It lacks, e.g., specific regulation mechanisms as well as the cell nucleus (which can influence motion in confinement8) and adhesion is taken into account only on the level of substrate friction. Moreover, experimentally observed migration behaviors depend on the specific cell type and even the phenotype. Direct comparisons to experiments must hence be taken with caution. It is nevertheless constructive to confront the general trends found in the generic framework to available experimental evidence.
Cells confined between two co-planar walls have been investigated in refs. 17,18. However, the focus was on the transitions from mesenchymal (i.e. adhesive, lamellipodial) to amoeboidal (non-adhesive) motion. The transitions were induced by reducing the adhesiveness in a confined surrounding, favoring the latter motility mode. A detailed study of lamellipodium-based motion in varying confinement at constant adhesiveness is lacking to date. The non-trivial results obtained already in our simple physical model, namely bistability and guidance (see Figs. 1c and 2) strongly suggest that it is worthwhile to carry out such a study.
Cells confined in microchannels have been investigated in refs. 15,16, albeit with a focus on invasiveness and integrin-myosin interplay, respectively. In both experiments, the channel cross-section was rectangular due to microfabrication restrictions. Nevertheless, cells adhered and formed lamellipodia all around the channel16, as in our model. Concerning cell speeds, a velocity increase by 30% in the channel compared to the flat substrate was found for wild type CHO cells16. The increase was much more pronounced when α4/paxillin binding was negatively regulated, resulting in amoeboidal motion which is, however, beyond the scope of our model. Amoeboidal motion was probably also induced by the confinement in ref. 15 studying pancreatic cancer cells, where a velocity increase by almost a factor of three was found. The generic model suggests a typical velocity increase by 40–50%, see Fig. 3c. This increase is a consequence of the confinement-induced shape change and the concomitant reorganization of the active lamellipodial front. Consequently, the experimentally observed increase could, at least in part, be traced back to the physical interaction between the environment and the cell and its internal actin structure.
Cell motion on fibers of various sizes has been experimentally studied for instance in refs. 27,28,29. The early study in ref. 27 investigated various cell types on fibronectin covered glass slides compared to thin fibronectin fibers deposited on glass slides. Little influence on cell speed was found, but the persistence of motion was strongly increased by the presence of the fibers. This is confirmed by very recent experiments29 investigating the alignment of fibroblasts (albeit without tracking the motion explicitly) on highly and weakly curved cylinders. There it was found that cells align more efficiently on cylinders with radii of the order of their own size compared to larger radii. These results are in agreement with the alignment effect found in the model and shown in Fig. 5. Collective cell migration on fibers has been also studied28. Although different from single cell motion, an arrest of motion for submicron fibers was observed. Since at this scale the front consists of a single cell, this arrest may be related to the one observed in the model (see Fig. 3c) for small fiber radii.
The effect of Gaussian curvature, see Fig. 6, was already critically discussed above in view of the recent experiments of ref. 26. For a better reproduction of the experimentally observed behavior, the model has to be augmented by actin stress fibers and possibly the cell’s nucleus.
Cell motion on periodic grooves or ridges on the surface of planar substrates has been extensively studied experimentally21,22,23,24,25. The alignment along the grooves as obtained in our model for patterns with long but still sub-cellular wavelengths (see red circles in Fig. 7a) has been found in all these experiments. In addition, the elongation of cells along the direction of motion (see Fig. 7b) was found in refs. 24,25 for both T cells and Dictyostelium cells. Nevertheless, the perpendicular motion could also be observed implying less elongated cells25, as recovered by the model (Fig. 7c). Ref. 25 even determined a “contact guidance efficiency” that first increased with the pattern wavelength before decreasing again when ridges become too distant for a single cell to sense. Although the model displays fast alignment (within a few cell sizes traveled) for long but sub-cellular wavelengths, which is the optimum range obtained in ref. 25, the time scale is quite sensitive to the actin parameters (especially θ). Hence, on the modeling side, the model should be tuned to a specific cell type to become quantitative. On the experimental side, it would be very interesting to include systematic variations of the pattern’s amplitude, to finally obtain a full picture of the contact guidance mechanism and its determining factors.
Let us also briefly compare our modeling framework to previous 3D cell migration modeling efforts. Previously, a two-fluid framework37 had been developed that allowed to reproduce shapes reminiscent of both fibroblasts and keratocytes36. Starting from a disc-shaped cell one had, however, to assume a certain fraction of the cell’s circumference to be (and always remain) protrusive. A more recent approach1 used active gel theory, but also had to assume a predefined leading edge. Hence while both models are more detailed concerning the internal cellular dynamics, the cell symmetry was broken (and hence cell polarity induced) by construction. Contrary, in our model, the equations of motion are completely symmetric and symmetry breaking emerges spontaneously in a self-organized way, as also recently confirmed by the occurrence of shape waves in a 2D version of the model44. Recently, a finite-element framework similar to ref. 36, but including adhesion, was used to model pseudopod-based migration38. All these works only investigated motion on flat substrates, but possibly would in principle allow for a detailed analysis of motion in confinement, curved substrates, and topographical features, as performed here, which would be interesting to critically assess our findings. Concerning motion in 3D fiber matrices, there are already a few modeling studies on ameboidal motion. In ref. 62, a hybrid agent-based/finite-element model was developed that includes pseudopod formation and even blebbing (via variable actin-plasma membrane linkage), while ref. 39 used a phase field approach using multiple internal density fields and especially an actin activator located but diffusing on the cell’s surface. These works are clearly complementary to our work, as they focus on activated pseudopods instead of well-defined lamellipodia. It would be very interesting to study within our model cell motion on an ensemble of few fibers, to compare the two distinct motility modes in this biologically relevant setting.
To conclude, we have demonstrated that the developed three-dimensional computational model is capable to describe lamellipodium-driven crawling motion of cells in – in principle – arbitrarily shaped surroundings and for a variety of cell types. Additionally, in contrast to earlier 3D models1,36,37,38, the approach does not need to pre-define protrusive regions in the cell, as it inherently displays bistability between immobile vs. moving states2. We investigated in depth several well-defined and experimentally studied scenarios of geometrical perturbation during cellular motion: such as a systematic variation of the substrate’s curvature (from cells on thin fibers to the movement inside a capillary), vertical confinement between two plates, as well as motion on topographically structured substrates. The discovered modulations in cell behavior and the resulting guiding principles of motile cells are due to purely physical effects (confinement, curvature) and their interplay with the self-organized shape and acto-myosin machinery. This study will therefore be fruitful in helping to differentiate such simpler effects from more complex cellular response due to biochemical cues and regulation.
The model was intentionally kept simple. We have already shown for the two-dimensional case that the modeling approach is modular63, hence explicit adhesion dynamics and substrate deformation42,43, global feedback on actin due to membrane tension48, as well as regulation (e.g., via Rho GTPases64,65) or additional components involved in the motility66 can be added without too much complication. Our approach may pave the way to study other 3D modes of motion beyond lamellipodia in the future, which are often triggered by confinement8,67. For instance, a confinement-induced transition of slow mesenchymal cells to a fast ameboidal motion has been found in the slit geometry17. Modes of 3D cellular motion distinct from lamellipodia also comprise blebbing68, osmotic force69 (i.e., flow of water through the cell), as well as pseudo-/lobopodial motion32,39,62 or the formation of invadopodia70. The latter is especially interesting as they allow cells to modify their surroundings and hence feedback on the confinement8,61 not only by deforming, but also by chemically degrading the extracellular matrix71. Another route is related to the cell nucleus affecting migration, especially when the confinement or radius of curvature becomes comparable to the nucleus size. The nucleus can be included into the computational framework via an additional passive phase-field, as done in 2D in ref. 72. In fact, including a deformable nucleus for motion in complex 3D environments would be a worthwhile endeavor for future investigations. Ultimately, motion inside 3D networks should become accessible73, similar as developed in ref. 39 for ameboidal motion employing pseudopods. Also for lamellipodial motion, this situation is arguably the most abundant and generic scenario for 3D motility. There, the mesh size vs. the size and deformability of the nucleus present new limiting steps on motion8, including feedback between motility and the genetic program of cells.
Methods
Numerical method and model validation
The Eqs. (1) and (2) are solved on a cubic, typically 2563, grid with an operator split Fourier pseudo-spectral code on graphical processing units (GPUs) using the CUDA programming language. The static substrate fields, Φ and Ψ are predefined beforehand. The simulation data were rendered and displayed using ParaView, a 3D data analysis and visualization application.
To validate the model, we studied the spreading dynamics, see Supplementary Fig. 2, and the onset of motion on flat substrates, see Supplementary Fig. 3. The latter reproduces the 2D behavior described in ref. 2, see also Fig. 1a). More details on the model implementation and testing can be found in Supplementary Note 1.
Data availability
Data that support the findings of this study are available upon reasonable request from the corresponding author.
Code availability
The code used to generate the data shown here is available upon reasonable request from the corresponding author.
References
Tjhung, E., Tiribocchi, A., Marenduzzo, D. & Cates, M. E. A minimal physical model captures the shapes of crawling cells. Nat. Commun. 6, 5420 (2015).
Ziebert, F., Swaminathan, S. & Aranson, I. S. Model for self-polarization and motility of keratocyte fragments. J. R. Soc. Interface 9, 1084–1092 (2012).
Saw, T. et al. Topological defects in epithelia govern cell death and extrusion. Nature 544, 212–216 (2017).
Palmieri, B., Bresler, Y., Wirtz, D. & Grant, M. Multiple scale model for cell migration in monolayers: elastic mismatch between cells enhances motility. Sci. Rep. 5, 11745 (2015).
Mogilner, A. & Keren, K. The shape of motile cells. Curr. Biol. 19, R762–R771 (2009).
Schindler, M. et al. Living in three dimensions. Cell Biochem. Biophys. 45, 215–227 (2006).
Nourshargh, S., Hordijk, P. L. & Sixt, M. Breaching multiple barriers: leukocyte motility through venular walls and the interstitium. Nat. Rev. Mol. Cell Biol. 11, 366–378 (2010).
Paul, C. D., Mistriotis, P. & Konstantopoulos, K. Cancer cell motility: lessons from migration in confined spaces. Nat. Rev. Cancer 17, 131–140 (2017).
Sahai, E. Illuminating the metastatic process. Nat. Rev. Cancer 7, 737–749 (2007).
Paul, C. D., Hung, W.-C., Wirtz, D. & Konstantopoulos, K. Engineered models of confined cell migration. Annu. Rev. Biomed. Eng. 18, 159–180 (2016).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Hooper, S., Marshall, J. F. & Sahai, E. Tumor cell migration in three dimensions. Methods Enzymol. 406, 625–643 (2006).
Van Goethem, E., Poincloux, R., Gauffre, F., Maridonneau-Parini, I. & Le Cabec, V. Matrix architecture dictates three-dimensional migration modes of human macrophages: differential involvement of proteases and podosome-like structures. J. Immunol. 184, 1049–1061 (2010).
Bordeleau, F., Tang, L. N. & Reinhart-King, C. A. Topographical guidance of 3d tumor cell migration at an interface of collagen densities. Phys. Biol. 10, 065004 (2013).
Rolli, C. G., Seufferlein, T., Kemkemer, R. & Spatz, J. P. Impact of tumor cell cytoskeleton organization on invasiveness and migration: a microchannel-based approach. PLoS ONE 5, e8726 (2010).
Hung, W.-C. et al. Distinct signaling mechanisms regulate migration in unconfined versus confined spaces. J. Cell Biol. 202, 807–824 (2013).
Liu, Y.-J. et al. Confinement and low adhesion induce fast amoeboid migration of slow mesenchymal cells. Cell 160, 659–672 (2015).
Renkawitz, J. et al. Adaptive force transmission in amoeboid cell migration. Nat. Cell Biol. 11, 1438–1443 (2009).
Curtis, A. & Wilkinson, C. Topographical control of cells. Biomaterials 18, 1573–1583 (1997).
Lim, J. Y. & Donahue, H. J. Cell sensing and response to micro- and nanostructured surfaces produced by chemical and topographic patterning. Tissue Eng. 13, 1879–1891 (2007).
Teixeira, A. I., Abrams, G. A., Bertics, P. J., Murphy, C. J. & Nealey, P. F. Epithelial contact guidance on well-defined micro- and nanostructured substrates. J. Cell Sci. 116, 1881–1892 (2003).
Kim, D.-H. et al. Mechanosensitivity of fibroblast cell shape and movement to anisotropic substratum topography gradients. Biomaterials 30, 5433–5444 (2009).
Hamilton, D. W., Oates, C. J., Hasanzadeh, A. & Mittler, S. Migration of periodontal ligament fibroblasts on nanometric topographical patterns: influence of filopodia and focal adhesions on contact guidance. PLoS ONE 5, e15129 (2010).
Kwon, K. W. et al. Nanotopography-guided migration of t cells. J. Immunol. 189, 2266–2273 (2012).
Driscoll, M. K., Sun, X., Guven, C., Fourkas, J. T. & Losert, W. Cellular contact guidance through dynamic sensing of nanotopography. ACS nano 8, 3546–3555 (2014).
Bade, N. D., Xu, T., Kamien, R. D., Assoian, R. K. & Stebe, K. J. Gaussian curvature directs stress fiber orientation and cell migration. Biophys. J. 114, 1467–1476 (2018).
Wojciak-Stothard, B., Denyer, M., Mishra, M. & Brown, R. A. Adhesion, orientation, and movement of cells cultured on ultrathin fibronectin fibers. In Vitro Cell Dev. Biol. Anim. 33, 110–117 (1997).
Yevick, H. G., Duclos, G., Bonnet, I. & Silberzan, P. Architecture and migration of an epithelium on a cylindrical wire. Proc. Natl Acad. Sci. USA 112, 5944–5949 (2015).
Bade, N. D., Kamien, R. D., Assoian, R. K. & Stebe, K. J. Curvature and rho activation differentially control the alignment of cells and stress fibers. Sci. Adv. 3, e1700150 (2017).
Carlier, M. F. & Pantaloni, D. Control of actin assembly dynamics in cell motility. J. Biol. Chem. 282, 23005–23009 (2007).
Pollard, T. D. & Cooper, J. A. Actin, a central player in cell shape and movement. Science 326, 1208–1212 (2009).
Petrie, R. J., Gavara, N., Chadwick, R. S. & Yamada, K. M. Nonpolarized signaling reveals two distinct modes of 3d cell migration. J. Cell Biol. 197, 439–455 (2012).
Fritz-Laylin, L. K. et al. Actin-based protrusions of migrating neutrophils are intrinsically lamellar and facilitate direction changes. eLife 6, e26990 (2017).
Rubinstein, B., Fournier, M. F., Jacobson, K., Verkhovsky, A. B. & Mogilner, A. Actin-myosin viscoelastic flow in the keratocyte lamellipod. Biophys. J. 97, 1853–1863 (2009).
Shao, D., Rappel, W. J. & Levine, H. Computational model for cell morphodynamics. Phys. Rev. Lett. 105, 108104 (2010).
Herant, M. & Dembo, M. Form and function in cell motility: from fibroblasts to keratocytes. Biophys. J. 98, 1408–1417 (2010).
Herant, M. & Dembo, M. Cytopede: a three-dimensional tool for modeling cell motility on a flat surface. J. Comput. Biol. 17, 1639–1677 (2010).
Allena, R. Cell migration with multiple pseudopodia: temporal and spatial sensing models. Bull. Math. Biol. 75, 288–316 (2013).
Moure, A. & Gomez, H. Phase-field model of cellular migration: three-dimensional simulations in fibrous networks. Comput. Methods Appl. Mech. Eng. 320, 162–197 (2017).
Ziebert, F. & Aranson, I. S. Computational approaches to substrate-based cell motility. npj Comput. Mater. 2, 16019 (2016).
Verkhovsky, A. B., Svitkina, T. M. & Borisy, G. G. Self-polarization and directional motility of cytoplasm. Curr. Biol. 9, 11–20 (1999).
Ziebert, F. & Aranson, I. S. Effects of adhesion dynamics and substrate compliance on the shape and motility of crawling cells. PLoS ONE 8, e64511 (2013).
Löber, J., Ziebert, F. & Aranson, I. S. Modeling crawling cell movement on soft engineered substrates. Soft Matter 10, 1365–1373 (2014).
Reeves, C., Winkler, B., Ziebert, F. & Aranson, I. S. Rotating lamellipodium waves in polarizing cells. Commun. Phys. 1, 73 (2018).
Mogilner, A. Mathematics of cell motility: have we got its number? J. Math. Biol. 58, 105–134 (2009).
Yam, P. T. et al. Actin-myosin network reorganisation breaks symmetry at the cell rear to spontaneously initiate polarized cell motility. J. Cell Biol. 178, 1207–1221 (2007).
Ziebert, F., Löber, J. & Aranson, I. S. in Physical Models of Cell motility, (ed. I. S. Aranson) p. 1–67 (Springer, Switzerland) (2016).
Winkler, B., Aranson, I. S. & Ziebert, F. Membrane tension feedback on shape and motility of eukaryotic cells. Phys. D 318–319, 26–33 (2016).
Nonomura, M. Study on multicellular systems using a phase field model. PLoS ONE 7, e33501 (2012).
Löber, J., Ziebert, F. & Aranson, I. S. Collisions of deformable cells lead to collective migration. Sci. Rep. 5, 9172 (2015).
Shao, D., Levine, H. & Rappel, W.-J. Coupling actin flow, adhesion, and morphology in a computational cell motility model. Proc. Natl Acad. Sci. USA 109, 6851–6856 (2012).
Jülicher, F., Kruse, K., Prost, J. & Joanny, J. F. Active behavior of the cytoskeleton. Phys. Rep. 449, 3–28 (2007).
Keren, K. et al. Mechanism of shape determination in motile cells. Nature 453, 475–480 (2008).
Schwarz, U. S. & Safran, S. A. Physics of adherent cells. Rev. Mod. Phys. 85, 1327 (2013).
Schmeiser, C. & Winkler, C. The flatness of lamellipodia explained by the interaction between actin dynamics and membrane deformation. J. Theor. Biol. 380, 144–155 (2015).
Small, J. V. Pushing with actin: from cells to pathogens. Biochem. Soc. Trans. 43, 84–91 (2015).
Hawkins, R. J. et al. Pushing off the walls: a mechanism of cell motility in confinement. Phys. Rev. Lett. 102, 058103 (2009).
Thoumine, O. & Ott, A. Time scale dependent viscoelastic and contractile regimes in fibroblasts probed by microplate manipulation. J. Cell Sci. 110, 2109–2116 (1997).
Csucs, G., Quirin, K. & Danuser, G. Locomotion of fish epidermal keratocytes on spatially selective adhesion patterns. Cell Motil. Cytoskeleton 64, 856–867 (2007).
Mizuhara, M. S., Berlyand, L. & Aranson, I. S. Minimal model of directed cell motility on patterned substrates. Phys. Rev. E 96, 052408 (2017).
Provenzano, P. P. et al. Collagen reorganization at the tumor-stromal interface facilitates local invasion. BMC Med. 4, 38 (2006).
Tozluoglu, M. et al. Matrix geometry determines optimal cancer cell migration strategy and modulates response to interventions. Nat. Cell Biol. 15, 751–762 (2013).
Ziebert, F. & Aranson, I. S. Modular approach for modeling cell motility. Eur. Phys. J. Spec. Top. 223, 1265–1277 (2014).
Marée, A. F., Grieneisen, V. A. & Edelstein-Keshet, L. How cells integrate complex stimuli: the effect of feedback from phosphoinositides and cell shape on cell polarization and motility. PLoS Comput. Biol. 8, e1002402 (2012).
Marth, W. & Voigt, A. Signaling networks and cell motility: a computational approach using a phase field description. J. Math. Biol. 69, 91–112 (2014).
Wolgemuth, C. W., Stajic, J. & Mogilner, A. Redundant mechanisms for stable cell locomotion revealed by minimal models. Biophys. J. 101, 545–553 (2011).
Lämmermann, T. & Sixt, M. Mechanical modes of amoeboid cell migration. Curr. Opin. Cell Biol. 21, 636–644 (2009).
Charras, G. & Paluch, E. Blebs lead the way: how to migrate without lamellipodia. Nat. Rev. Mol. Cell Biol. 9, 730–736 (2008).
Stroka, K. M. et al. Water permeation drives tumor cell migration in confined microenvironments. Cell 157, 611–623 (2014).
Buccione, R., Orth, J. D. & McNiven, M. A. Foot and mouth: podosomes, invadopodia and circular dorsal ruffles. Nat. Rev. Mol. Cell Biol. 5, 647–657 (2004).
Zaman, M. H. et al. Migration of tumor cells in 3d matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc. Natl Acad. Sci. USA 103, 10889–10894 (2006).
Camley, B. A. et al. Polarity mechanisms such as contact inhibition of locomotion regulate persistent rotational motion of mammalian cells on micropatterns. Proc. Natl Acad. Sci. USA 111, 14770–14775 (2014).
Zhu, J. & Mogilner, A. Comparison of cell migration mechanical strategies in three-dimensional matrices: a computational study. Interface Focus 6, 20160040 (2016).
Acknowledgements
B.W. and F.Z. acknowledge funding from the German Science Foundation (DFG) via project ZI 1232/2-2. The research of I.S.A. was supported by the NSF PHY-1707900.
Author information
Authors and Affiliations
Contributions
B.W., I.S.A., and F.Z. were involved in developing the model. B.W. developed the numerical code and analyzed the data. F.Z. designed and supervised the research. All authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Winkler, B., Aranson, I.S. & Ziebert, F. Confinement and substrate topography control cell migration in a 3D computational model. Commun Phys 2, 82 (2019). https://doi.org/10.1038/s42005-019-0185-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-019-0185-x
This article is cited by
-
In silico analysis shows that dynamic changes in curvature guide cell migration over long distances
Biomechanics and Modeling in Mechanobiology (2024)
-
Control of protein-based pattern formation via guiding cues
Nature Reviews Physics (2022)
-
Spontaneous polarization and cell guidance on asymmetric nanotopography
Communications Physics (2022)
-
Phase field model for cell spreading dynamics
Journal of Mathematical Biology (2022)
-
Investigating the nature of active forces in tissues reveals how contractile cells can form extensile monolayers
Nature Materials (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.