Abstract
A vast source of methane is found in gas hydrate deposits, which form naturally dispersed throughout ocean sediments and arctic permafrost. Methane may be obtained from hydrates by exchange with hydrocarbon byproduct carbon dioxide. It is imperative for the development of safe methane extraction and carbon dioxide sequestration to understand how methane and carbon dioxide co-occupy the same hydrate structure. Pair distribution functions (PDFs) provide atomic-scale structural insight into intermolecular interactions in methane and carbon dioxide hydrates. We present experimental neutron PDFs of methane, carbon dioxide and mixed methane-carbon dioxide hydrates at 10 K analyzed with complementing classical molecular dynamics simulations and Reverse Monte Carlo fitting. Mixed hydrate, which forms during the exchange process, is more locally disordered than methane or carbon dioxide hydrates. The behavior of mixed gas species cannot be interpolated from properties of pure compounds, and PDF measurements provide important understanding of how the guest composition impacts overall order in the hydrate structure.
Similar content being viewed by others
Introduction
Gas hydrates form in natural settings including ocean floor and sub-surface permafrost deposits when water is in the presence of a gas, such as CH4, under relatively modest pressures and low temperature conditions. These non-stoichiometric compounds are a hydrogen-bonded water framework which crystallize as a clathrate cage host structure around occluded guest molecules, primarily CH41,2,3. Gas hydrates are high-density sources of CH4 and accessible deposits contain an estimated 3000 trillion cubic meters of fuel2,4. CH4 may be produced from deposits via dissociation (warming or depressurization) or exchange with CO2. The latter method can potentially stabilize and maintain natural deposits, while sequestering the hydrocarbon byproduct4,5. As thermodynamic predictions, molecular dynamics (MD) simulations, and in situ diffraction experiments demonstrate, a gas hydrate with partial replacement of CO2 for CH4 forms and is more stable at higher temperature and lower pressure than a pure CH4 hydrate1,3. This attribute led to a field study in the Alaskan North Slope which demonstrated that CH4 can be collected from a hydrate deposit by exchange with CO2. The field study, however, did not achieve a complete exchange and resulted in a mixed hydrate deposit5. Guest–host interactions of the CH4, CO2, and mixed CH4-CO2 hydrate structures need to be thoroughly deciphered, to improve predictions of how to achieve a full CH4-CO2 exchange and to confirm the stability and safety of the altered deposit with mixed CH4-CO2 or pure CO2 hydrate.
Three clathrate structures have been observed for natural hydrates; however, sI hydrate (the cubic structure type) is the most relevant to the formation conditions of accessible CH4 hydrate and therefore our focus in structural studies1,3,4. SI hydrate has a cubic \(Pm\bar 3n\) (223) crystal structure, composed of 46 hydrogen-bonded H2O molecules, which form the host lattice. This structure is made up of eight cages: two small dodecahedral and six large tetrakaidekahedral cages, which can occlude up to one guest molecule3. The average crystal structure of sI CH4 and CO2 hydrate has been well characterized with diffraction6,7,8,9,10,11, spectroscopy12,13,14,15, MD simulations16,17,18,19,20,21,22,23, and density functional theory calculations11,24,25,26, which suggest a high degree of disorder within the crystal. This disorder is evident in the crystallographic model of sI CH4 hydrate which requires four partially occupied proton positions for each H2O molecule and many positions to represent the unbonded, freely rotating CH4 or partially constrained CO2 molecules27. The intermolecular interactions that result in structural disorder are frequently studied with pair distribution functions (PDFs) calculated from simulated atomic models18,22,23,28,29,30. Computational studies suggest that mixing guests in the hydrate structure impacts the guest–host (CH4 or CO2–H2O), guest–guest, and host–host PDFs. CH4 is found to have weaker interactions with the host, providing a more ordered lattice, whereas CO2 interacts strongly with the host and other guests, leading to disorder18,21. Complementary local structure experiments investigating the impact of CH4, CO2, and their mixture on the structure and stability of gas hydrates have not been reported to the best of our knowledge. In fact, there are only three published neutron PDF experiments involving gas hydrates which demonstrated limited PDF resolution due to instrumental capabilities31,32,33. Here we present neutron PDF data for CH4, CO2, and mixed CH4–CO2 hydrates measured at 10 K using time-of-flight neutron powder diffraction. Detailed analysis of this data was achieved using combined MD and Reverse Monte Carlo (RMC) simulations to model the neutron PDF data. Our analysis provides a path to observe the short-range interactions of CH4 and CO2 guest molecules with their host hydrate structure and how mixing guest molecules impacts stability.
Results
Fitting neutron PDF data
The interpretation of PDFs obtained from neutron scattering experiments of complex materials requires atomic-scale computational modeling. In this work, classical MD simulations of rigid molecules provide an initial structural guess, which we refine with RMC simulation to more accurately explain the PDF data. We approach this data analysis with a combination of MD relaxed models, which are optimized through RMC, which incorporates both physical constraints and experimental observation. Our goal is to provide a descriptive insight into neutron PDF data with features arising from multiple species: in this case, CO2, CH4, and water.
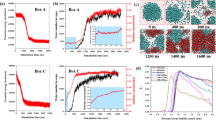
Simulated neutron PDFs are calculated from large-box (~20,000 atom) models relaxed with MD, plotted in Fig. 1a, c, e and compared with the experimental neutron PDF data. These model and data comparisons demonstrate the local disorder that is not fully described by the MD simulations of pure CH4, pure CO2, or mixed CH4–CO2 hydrate systems, as the simulated peaks are noticeably too sharp and narrow. Although the MD PDFs do predict peak locations and provide a qualitative model of the short-range order in the three hydrate systems, the peak widths and shapes do not match the neutron PDF data. They do not capture the degree of disorder, which represents differences in guest–host interactions across the compositions; however, the MD models provide a good peak location comparison with the neutron data. We implement RMC simulations to further fit this model from MD to the data with RMCProfile. This program is designed for PDF analysis of disordered crystal and amorphous structures, using chemical and physical constraints to fit a large-box model to neutron PDF data34. Figure 1a, c, e, demonstrate the improvement achieved when the relaxed MD model for each system is fit to the data with RMC modeling. A dramatic improvement in model to data fit is achieved using RMC for all cases: CH4, CO2, and mixed CH4–CO2 hydrates, with a small oscillating difference between the RMC model and the experimental data beyond 2 Å. Pair distances in the 2–10 Å region are the most interesting PDF features for these systems, where CH4–CO2 composition has the most measurable impact on guest–host and host–host interactions.
Comparison of experimental neutron PDF data with MD modeled PDF (a, c, e bottom spectra) and the model fit to the data with RMC (a, c, e top spectra) with corresponding difference patterns for CH4 (a), CH4–CO2 (c), and CO2 (e) hydrates. The overall disorder that RMC models (b, d, f top) calculate is also evident when compared with the MD structures (b, d, f bottom).
The overall lack of disorder in the MD model is the primary source of misfit in Fig. 1 for the atomic separations above 2 Å. We might suspect the rigid intramolecular potentials that were used in the MD simulations as the main culprit, but the improved fits are achieved with RMC constraints which emphasize intermolecular distance relaxations. This is seen, as the greatest RMC-data difference is below 2 Å for all three hydrate systems, where the strictest constraints were applied. We did not pursue further adjustment of the intramolecular bonds to improve this low r region or the weak oscillations in the RMC/data difference from 2 to 25 Å, because this risks overfitting and pushes the limitations of the data. Physical constraints are implemented in our RMC simulations to introduce flexible intramolecular bonds, to prevent unphysical collision between atoms and to maintain ice rules for hydrogen bonding in the hydrate host lattice. Under these constraints, the RMC simulation is driven by the goodness of fit of the model to the neutron PDF data34.
A comparison of structural snapshots from the MD and RMC simulations visualize the disorder missing in the MD relaxation. The models for all three hydrate compositions, drawn in Fig. 1b, d, f, corresponding to the MD model (bottom) and the RMC fit structure (top), show that the crystal structures are maintained while there is an increase in local disorder of atomic positions. This short-range analysis provides characterization of the local disorder in crystals which is not generally observed in characterization with Bragg diffraction. Achieving PDF analysis of these experimental data enables the investigation of the short-range interactions in these hydrates, which drives varied thermodynamic behavior across the three solid solution compositions.
Host–host interactions
Once a good fit to the neutron PDF is achieved, we isolate contributions from atom types by calculating radial distribution functions (RDFs) from simulations of the fit models. RDFs are calculated from MD simulations and the RMC fits for analysis of individual species pair distances in the pure CH4, CO2, and mixed CH4–CO2 hydrate structures to discern which molecular interactions define the three hydrate systems. RDFs calculated from MD simulation versus RMC fits to experimental data highlight the features that are not captured in the MD simulation. Specifically, host–host, guest–host, and guest–guest intermolecular interactions are compared for the two guest and cage types for both CH4 and CO2 hydrates, and mixed CH4–CO2 hydrate via the RDFs calculated from RMC fits. The total neutron PDFs in Fig. 1a, c, e exhibit more disorder in the experimental models than the MD models, so the species-specific RDFs can indicate what interactions lead to that disorder.
Host–host intermolecular interactions are most clearly described by the Owater–Owater RDFs, which are used to measure distortion in the host lattice. RDFs from the fits to neutron data and MD simulations for the pure CH4, pure CO2, or CH4–CO2 mixed hydrate samples which compare the impact of guest composition on the hydrate host lattice are shown in Fig. 2a. Structures of large and small cages selected from the large-box models are shown in Fig. 2b, c to visualize the MD and the experimental results. The RDF peaks in Fig. 2a, which measure Owater–Owater pair distances for the D2O molecules that form hydrate cages, are more distorted in the experiment than MD models predict and are generally wider, shorter, and not normally distributed. Analysis of the experimental data shows that the first nearest neighbor Owater–Owater peak, at ~2.8 Å, is much broader than MD simulations indicate. Instead of normally, tightly distributed distances, the O pairs distribute from 2 to 3.25 Å with an average of 2.7 Å and a defined high-r shoulder at 3.2 Å. The mixed CH4–CO2 hydrate also has a low r shoulder due to some shortening of pentagonal face edges. This detail, in combination with the smeared peak shape as r increases, indicates that the hydrate lattice is the most disordered for a guest composition of mixed CH4–CO2. We can visualize a large and small cage extracted from experimental fits in Fig. 2c to show that mixed CH4–CO2 hydrate cages are the most distorted. As these compositions follow experimental occupancy, some CH4 and the mixed CH4–CO2 hydrate cages are vacant. The cages drawn in Fig. 2b, c were chosen as ones where the large cage is occupied by a CO2 and the small cage is occupied by a CH4. The imbalance of guest molecule shape and interaction potential in the mixed hydrate system, which arises from the cage filling where CO2 may be surrounded by CH4 or a vacancy, clearly leads to higher level of disorder in the structure.
a RDFs calculated from MD simulations (dashed lines) and RMC fits to neutron data (solid) for the Owater–Owater pair distances for observing the impact of guest molecule on cage distortion. b A large and small cage pair before RMC and c after RMC fit to data showing distortion in the cages as observed with neutron PDF data. The distortion of the cages is the most extreme for the mixed guest hydrate (middle row).
Guest–host Interactions
A range of cage filling results have been predicted and observed for hydrates synthesized with CH4 or CO2 as the feed gas under varying pressure–temperature conditions and synthesis procedures. At 10 K, despite the larger size of the CO2 molecule, hydrate with CO2 as a guest has a smaller lattice parameter and CO2 will preferentially fill the large cage over CH43,7. Local structure analysis of the guest–host RDFs in Fig. 3 lead to an understanding of how the guest molecules in different compositions interact with the host and affect cage filling, formation, stability, and decomposition behavior3,7,18. RDFs corresponding to the CO2 molecule center (the central C) for pure CO2 and mixed CH4–CO2 hydrates are plotted in the top row of Fig. 3 for the large cage (a) and small cage (b). The first peak for the Owater–CCO2 pair distances is centered around ~4.5 Å for both compositions, but the mixed hydrate has a distinct split and is wider. The CO2 guest in the mixed hydrate is located at more distinct positions than in the pure CO2 hydrate, where the molecule is primarily centrally located in the cage. Overall, though, the guest RDF peaks in the large cage show that the local guest–host structure of CO2 in mixed hydrate is more disordered. This will impact stability upon heating and suggests that host–CO2 interactions in mixed hydrate are more favorable than in pure CO2 hydrate. In the small cage (Fig. 3b), the position of the molecule center is essentially the same in both compositions. The CH4 molecule in CH4 and mixed hydrate systems occupies a tighter distribution of orientations than CO2. The Owater–CCH4 RDFs in the middle row (Fig. 3c, d) are defined by sharper, narrower peaks than in the CO2 hydrate cases. Though the CH4 positions are slightly less ordered in the mixed hydrate composition, the CH4 guest has a lower interaction potential with the other molecules in the system, is not a polar molecule, and essentially occupies cage centers. The quadrupolar CO2 occupies a more distributed position, especially in the large cage, leading to different interaction surfaces with its immediate host lattice D2O neighbors. Finally, as CO2 is a linear molecule, the Owater–OCO2 RDFs in the bottom row of Fig. 3e, f show that CO2 in both cages approaches the D2O molecules at distances just above 2 Å, close enough to form weak hydrogen bonds which result in the distortions observed in Fig. 2 and structural stabilization. Guest–host distances across the RDFs of both compositions show that largest distortion of the local hydrate cage structure occurs in the mixed hydrate system. In this composition, CO2 may be neighbored by another CO2, CH4, or a vacancy. We see that a change in surrounding environment allows the molecule to distribute across a broader range of positions and orientations in the cages, causing OCO2 to interact more closely with enclathrating D2O molecules.
RDFs to investigate guest–host interactions in three hydrate compositions. Owater–CCO2 RDFs for the large cage (a) and small cage (b) CO2 centers and Owater–CCH4 RDFs for the large cage (c) and small cage (d) CH4 centers indicate that CO2 molecules are not as ordered in their positions as CH4. Orientations of the linear CO2 in Owater–OCO2 RDFs in the large cage (e) and small cage (f) show that the CO2 in mixed hydrate orients in more positions towards the cage.
We can visualize and better understand the RDFs with three-dimensional density distributions calculated from MD trajectory frames and RMC fits. Orientation rotations were performed on the large cage guest molecules in order to accurately describe how they occupy the cage space with respect to the two offset hexagonal faces. Consistent with former experimental and computational results, the CH4 molecule in both the MD and experimental distributions in Fig. 4a–f, m–r, respectively, orients with a spherical distribution in both the large and small cages7,18. Previously, the CO2 molecule was shown to orient in an oblate shape parallel to the hexagonal face of the large hydrate cage, along the longer axis7,10,24,35. Neutron diffraction experiments showed this orientation in both pure CO2 and mixed CH4–CO2 hydrate, but MD simulations showed that this only held for distributions in the pure CO2 hydrate. MD simulations in Fig. 4h, i show CO2 occupying an additional orientation towards the hexagonal face across the short axis of the cage, elaborating on previous diffraction studies where this orientation seems to be averaged out in the long-range crystallographic analysis. Neutron PDF analysis allows for direct experimental evidence of this local structure for the mixed hydrate. Figure 4t, u, w, x provide density distributions of the CO2 produced from models of the experimental data, which indicate that in mixed CH4–CO2 hydrate the guest in the large cage does occupy additional orientations toward the hexagonal face that do not occur in CO2 hydrate. Experimental orientations are not as defined as MD predicts, as might be expected from the trend of higher disorder in experimental data seen throughout the model comparisons in previous sections. We do see, however, a low density of orientations pointing directly into the hexagonal face and greater distribution of OCO2 positions angling toward that face than in the pure CO2 composition where a narrower oblate distribution is occupied.
Discussion
Guest–host interactions in CH4, CO2, and mixed CH4–CO2 hydrates, as observed by experimental neutron PDF analysis, are dependent on guest type and cage filling. The mixed hydrate local structure is more disordered and distorted than the pure CH4 and CO2 hydrates. Although these solid solution compositions follow a rule of mixtures for crystallographic properties including lattice parameter, the mixed hydrate is the outlier when the local structure is inspected. Large-box models were fit to neutron PDF data for CH4, CO2, and mixed CH4–CO2 hydrate powders, to produce RDFs and density distributions for local structure analysis. CO2, when surrounded by CH4 and vacant cages, has the freedom to move into a broader distribution of orientations in the hydrate cages. This is an important detail to consider when attempting a CH4–CO2 exchange, as a mixed hydrate will form in the intermediate steps. RDFs show that the local structure of the occluded species in the mixed hydrate clearly does not fall between that in the pure CH4 and CO2 hydrates. Guest molecule interactions with the host lattice cannot be expected to behave as an intermediate of pure CH4 and CO2 hydrate, even though predicted pressure and temperature of formation from gas and liquid components do. The altered CO2–host local structure and interactions may create a free energy well that requires an interruption or altered temperature/pressure input to overcome as the disordered positions are more closely interacting with the host molecules. Our analysis approach provides a path forward to analyze neutron PDF experiments of an in situ CH4–CO2 exchange. Developing these studies allows for investigation into possible remedies to achieve a full CH4–CO2 exchange, such as a helper molecule to overcome the energy barrier5. Characterization of guest–host structure and interactions leads to understanding of the long-term stability of an altered natural hydrate deposit. This local characterization at 10 K shows that framework distortion in mixed CH4–CO2 hydrate, stabilized by CO2, results in a disordered structure that inhibits further CH4 removal. We demonstrate that the CH4–CO2 guest composition impacts the guest–host interactions in hydrates during static temperature measurements at 10 K, implying that these interactions are important in formation and decomposition. Further studies should apply these techniques during those reactions if a complete structure–property relationship is to be determined. These experimental and analysis methods have successfully presented observable differences in the hydrate compositions with CH4 and CO2 at 10 K. The development of these experiments and analysis at cold temperatures, where the complexity of molecular thermal motion of is minimal, makes investigations such as this possible at the higher working temperatures of a CH4–CO2 exchange.
Methods
Sample synthesis
CH4, CO2, and mixed CH4–CO2 hydrate powders were synthesized for the neutron total scattering experiments. One milligram of Pseudomonas syringae 31a, branded as Snomax, was mixed with 10 mL D2O for each sample. Snomax is an ice nucleating protein and has been demonstrated to decrease the formation pressure required for gas hydrates35. A 300 mL pressure reactor vessel containing the solution and steel milling media was evacuated and then pressurized at room temperature with 600 psi of gas. The three samples were pure CO2 and CH4 hydrates, and mixed CH4–CO2 hydrates, where the CH4/CO2 feed gas fraction was 0.5/0.5. For each composition after pressurization, the vessel was placed in a freezer where the temperature was dropped to 277 K and tumbled periodically for ~7 days. The freezer temperature was dropped to 253 K for at least 1 day to quench the sample after the pressure in the vessel equilibrated. Hydrate powders were collected under a nitrogen atmosphere and stored in vanadium cans in liquid nitrogen until the neutron experiments.
Measuring neutron PDFs
Neutron PDF data were collected from CH4, CO2, and mixed CH4–CO2 hydrate powder samples on the Nanaoscale-Ordered Materials Diffractometer at the Spallation Neutron Source at Oak Ridge National Laboratory. Samples were measured at 10 K in a 50 mm He cryostat. Vanadium cans containing each sample were transferred from the liquid nitrogen storage dry shipper to the cryostat set to 90 K. The sample was then held in the cryostat at 90 K, to allow any liquid nitrogen to boil off before temperature was dropped to 2 K and increased to 10 K. Data were collected under low He pressure at 55 mbar, the working pressure for the cryostat. Neutron PDF data were collected in reciprocal space and Fourier transformed to obtain the real space function. The neutron S(Q)s from 0.6-35 Å−1 were converted in PyStoG36 to the real space function using
with
and the intermolecular bond distances (greater than ~2.5 Å) are emphasized by weighting the function as
which is the function that was used to fit all neutron PDF data37,38.
Measuring a neutron PDF is achieved by collecting data through a wide range of Q to observe Bragg reflections and diffuse scattering, providing a powder diffraction pattern for crystal structure analysis. Rietveld refinements of the Bragg data to confirm the hydrate structure, obtain average composition and lattice parameters, and amount of ice (present as a minor secondary phase) were performed using GSAS-EXPGUI39,40. The details are outlined in Supplementary Table 1 and Supplementary Fig. 1, and are in agreement with the results for the same three compositions synthesized under the same conditions measured with high-resolution neutron powder diffraction at 10 K7.
MD simulations and PDF analysis
Large-box models of ~20,000 atoms (depending on composition) were produced by building 5 × 5 × 5 unit cell models of the host lattice following the ideal proton configuration for sI hydrates determined by Takeuchi et al.27. The large and small cages were filled with CH4, CO2, or a vacancy according to the experimentally determined occupancies, outlined in Supplementary Table 27. Two sets of models were built with this method for the three hydrate compositions using experimental lattice parameters. One set of initial large boxes were equilibrated with the LAMMPS (large-scale atomic/molecular massively parallel simulator) simulations package in the NPT (constant atoms, pressure, and temperature) and then simulated in the NVT (constant atoms, volume, and temperature) ensemble for 100 ps with a 1 fs timestep and 100 fs damping constant41. RDFs and density distributions were calculated from the NVT trajectories to compare with the experimental results. Fixed rigid models were used in the MD simulations for the H2O, CH4, and CO2 molecules. The TIP4P potential model with a long-range Coulombic solver was used for the water molecules, while the CH4 and CO2 partial charges and Lennard Jones parameters were taken from Tse et al.30 (CH4) and Duan and Zhang42 (CO2). Finally, all intermolecular interactions for H2O, CH4, and CO2 were determined with Lorentz–Berthelot combination rules22. The second set of boxes were relaxed in NVT for 100 ps at the lattice parameter and composition determined with Rietveld refinements to produce initial models for RMC fits.
Large-box models were fit to the data using RMCProfile34. RMC model constraints, summarized in Supplementary Table 3, result in an acceptance rate of ~10% once equilibration is reached. RDFs and density distributions were calculated from the equilibrated MD trajectories and sets of frames from RMC fits using custom codes written in FORTRAN. RMC fits for the CH4, CO2, and mixed CH4–CO2 hydrate models were each simulated for an additional 25 × 106 generated moves from 15 unique initial configurations and the resulting structures were sampled to produce a collection of ~200 frames of accepted atom configurations for each structure. We mimicked MD simulation trajectories from equilibrated RMC frames and calculated partial RDFs for individual species pair distances in the three structures. These RDFs are a probability density function, which describe the likelihood of a particle being at a distance r from another particle and are calculated as:
where \(g\left( r \right)\)has the limits as \(r \to \infty\), \(g \to 1\)37.
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
Code availability
The codes used in this study are primarily cited. RMCProfile version 6.7.4.3 was used to fit the neutron data and is available at RMCProfile.org. PyStog version 0.1.3 is available at https://pystog.readthedocs.io/en/latest/. The code to calculate the radial distribution function from data fits and simulations is found at http://utkstair.org/clausius/docs/mse614/text/examples.html. MD simulations were performed with LAMMPS, documented at https://lammps.sandia.gov/doc/Manual.html.
References
Koh, C. A. & Sloan, E. D. Natural gas hydrates: recent advances and challenges in energy and environmental applications. AIChE J. 53, 1636–1643 (2007).
Koh, C. A., Sloan, E. D., Sum, A. K. & Wu, D. T. Fundamentals and applications of gas hydrates. Annu. Rev. Chem. Biomol. Eng. 2, 237–257 (2011).
Sloan, E. D. & Koh, C. A. Clathrate Hydrates of Natural Gases (CRC Press, 2007).
Chong, Z. R., Yang, S. H. B., Babu, P., Linga, P. & Li, X.-S. Review of natural gas hydrates as an energy resource: prospects and challenges. Appl. Energy 162, 1633–1652 (2016).
Boswell, R. et al. The Iġnik Sikumi Field Experiment, Alaska North Slope: design, operations, and implications for CO2–CH4 exchange in gas hydrate reservoirs. Energy Fuels 37, 140–153 (2016).
Circone, S. et al. CO2 hydrate: synthesis, composition, structure, dissociation behavior, and a comparison to structure I CH4 hydrate. J. Phys. Chem. B 107, 5529–5539 (2003).
Everett, S. M. et al. Insights into the structure of mixed CO2/CH4 in gas hydrates. Am. Mineral. 100, 1203–1028 (2015).
Everett, S. M. et al. Kinetics of methane hydrate decomposition studied via in situ low temperature X-ray powder diffraction. J. Phys. Chem. A 117, 3593–3598 (2013).
Hansen, T. C., Falenty, A. & Kuhs, W. F. Lattice constants and expansivities of gas hydrates from 10 K up to the stability limit. J. Chem. Phys. 144, 054301 (2016).
Ikeda, T. et al. Neutron diffraction study of carbon dioxide clathrate hydrate. J. Phys. Chem. Solids 60, 1527–1529 (1999).
Izquierdo-Ruiz, F., Otero-de-la-Roza, A., Contreras-García, J., Prieto-Ballesteros, O. & Recio, J. Effects of the CO2 guest molecule on the sI clathrate hydrate structure. Materials 9, 777 (2016).
Casco, M. E. et al. Methane hydrate formation in confined nanospace can surpass nature. Nat. Commun. 6, 6432 (2015).
Qin, J. & Kuhs, W. F. Quantitative analysis of gas hydrates using Raman spectroscopy. AIChE J. 59, 2155–2167 (2013).
Ripmeester, J. A. & Ratcliffe, C. I. Low-temperature cross-polarization/magic angle spinning carbon-13 NMR of solid methane hydrates: structure, cage occupancy, and hydration number. J. Phys. Chem. 92, 337–339 (1988).
Tse, J. S., Ratcliffe, C. I., Powell, B. M., Sears, V. F. & Handa, P. Y. Rotational and translational motions of trapped methane. Incoherent inelastic neutron scattering of methane hydrate†. J. Phys. Chem. A 101, 4491–4495 (1997).
Anderson, B. J., Tester, J. W. & Trout, B. L. Accurate potentials for argon−water and methane−water interactions via ab initio methods and their application to clathrate hydrates. J. Phys. Chem. B 108, 18705–18715 (2004).
Chialvo, A. A., Houssa, M. & Cummings, P. T. Molecular dynamics study of the structure and thermophysical properties of model sI clathrate hydrates. J. Phys. Chem. B 106, 442–451 (2002).
Cladek, B. R. et al. Guest-host interactions in mixed CH4–CO2 hydrates: insights from molecular dynamic simulations. J. Phys. Chem. C. 122, 19575–19583 (2018).
Cladek, B. R. et al. Molecular rotational dynamics in mixed CH4–CO2 hydrates: insights from molecular dynamics simulations. J. Phys. Chem. C. 123, 26251–26262 (2019).
Conde, M. M. et al. Can gas hydrate structures be described using classical simulations? J. Chem. Phys. 132, 114503 (2010).
Costandy, J., Michalis, V. K., Tsimpanogiannis, I. N., Stubos, A. K. & Economou, I. G. Lattice constants of pure methane and carbon dioxide hydrates at low temperatures. Implementing quantum corrections to classical molecular dynamics studies. J. Chem. Phys. 144, 124512 (2016).
Jiang, H. & Jordan, K. D. Comparison of the properties of xenon, methane, and carbon dioxide hydrates from equilibrium and nonequilibrium molecular dynamics simulations†. J. Phys. Chem. C. 114, 5555–5564 (2010).
Zhu, J. et al. Encapsulation kinetics and dynamics of carbon monoxide in clathrate hydrate. Nat. Commun. 5, 1428 (2014).
Cox, S. J., Towler, M. D., Alfè, D. & Michaelides, A. Benchmarking the performance of density functional theory and point charge force fields in their description of sI methane hydrate against diffusion Monte Carlo. J. Chem. Phys. 140, 174703 (2014).
Pérez-Rodríguez, M. et al. Computational study of the interplay between intermolecular interactions and CO2 orientations in type I hydrates. Phys. Chem. Chem. Phys. 19, 3384–3393 (2016).
Vidal-Vidal, Á., Pérez-Rodríguez, M., Torré, J. P. & Piñeiro, M. M. DFT calculation of the potential energy landscape topology and Raman spectra of type I CH4 and CO2 hydrates. Phys. Chem. Chem. Phys. 17, 6963–6975 (2015).
Takeuchi, F. et al. Water proton configurations in structures I, II, and H clathrate hydrate unit cells. J. Chem. Phys. 138, 124504 (2013).
Buch, V. et al. Clathrate hydrates with hydrogen -bonding guests. Phys. Chem. Chem. Phys. 11, 10245–10265 (2009).
Jia, J., Liang, Y., Tsuji, T., Murata, S. & Matsuoka, T. Elasticity and stability of clathrate hydrate: role of guest molecule motions. Sci. Rep. 7, 1290 (2017).
Tse, J. S., Klein, M. L. & McDonald, I. R. Computer simulation studies of the structure I clathrate hydrates of methane, tetrafluoromethane, cyclopropane, and ethylene oxide. J. Chem. Phys. 81, 6146–6153 (1984).
Koh, C. A. et al. Mechanisms of gas hydrate formation and inhibition. Fluid Phase Equilibria 194, 143–151 (2002).
Thompson, H. et al. Methane hydrate formation and decomposition: structural studies via neutron diffraction and empirical potential structure refinement. J. Chem. Phys. 124, 164508 (2006).
Tulk, C. A., Klug, D. D., Molaison, J. J., dos Santos, A. M. & Pradhan, N. Structure and stability of an amorphous water–methane mixture produced by cold compression of methane hydrate. Phys. Rev. B 86, 054110 (2012).
Tucker, M. G., Keen, D. A., Dove, M. T., Goodwin, A. L. & Hui, Q. RMCProfile: reverse Monte Carlo for polycrystalline materials. J. Phys. Condens. Matter 19, 335218 (2007).
McCallum, S. D., Riestenberg, D. E., Zatsepina, O. Y. & Phelps, T. J. Effect of pressure vessel size on the formation of gas hydrates. J. Pet. Sci. Eng. 56, 54–64 (2007).
McDonnell, M. PyStog, https://pystog.readthedocs.io/en/latest/ (2018).
Egami, T., Kolko, J., Billinge, S. J. L. & Inc, E. I. Underneath the Bragg Peaks: Structural Analysis of Complex Materials (Elsevier, 2003).
Keen, D. A. A comparison of various commonly used correlation functions for describing total scattering. J. Appl. Crystallogr. 34, 172–177 (2001).
Larson, A. C. & Von Dreel, R. B. General structure analysis system (GSAS) Los Alamos National Laboratory Report LAUR 86-748 (2000).
Toby, B. H. EXPGUI, a graphical user interface for GSAS. J. Appl. Crystallogr. 34, 210–213 (2001).
Plimpton, S. Fast parallel algorithms for short-range molecular dynamics. J. Comput. Phys. 117, 1–19 (1995).
Duan, Z. & Zhang, Z. Equation of state of the H2O, CO2, and H2O –CO2 systems up to 10 GPa and 2573.15 K: molecular dynamics simulations with ab initio potential surface. Geochim. Cosmichim. Acta 70, 2311–2324 (2006).
Acknowledgements
B.R.C. received partial support from the University of Tennessee Chancellor’s Fellowship program and the Center for Materials Processing, a Tennessee Higher Education Commission (THEC) Center of Excellence. Part of this research was sponsored by the Office of Science Graduate Student Research (SCGSR) program, administered by the Oak Ridge Institute for Science and Education for DOE. Research at the Spallation Neutron Source (SNS), a US Department of Energy (DOE) Office of Science User Facility operated by Oak Ridge National Laboratory (ORNL), was sponsored by the DOE Office of Basic Energy Sciences. The ICE-MAN software suite used for this work was funded by the ORNL Laboratory Directed Research and Development program. We thank Dayton Kizzire from University of Tennessee, Knoxville, for assisting with the neutron total scattering data collection at the SNS and Jared Floyd for assistance in sample synthesis.
Author information
Authors and Affiliations
Contributions
B.R.C. prepared the samples, performed the MD simulations, RMC fits, and data analysis. B.R.C., M.T.M., and M.G.T. performed the neutron data reduction. All authors participated in the neutron experiments at the SNS. D.J.K. wrote the codes to calculate RDFs and three-dimensional density distributions. Sample synthesis and neutron powder diffraction experiments were initiated by B.R.C. and C.J.R. This work is based on previous studies by S.M.E. and C.J.R. B.R.C. wrote the initial manuscript draft and C.J.R., D.J.K., and M.G.T. contributed to early editing. All authors contributed to, revised, and approved of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cladek, B.R., Everett, S.M., McDonnell, M.T. et al. Local structure and distortions of mixed methane-carbon dioxide hydrates. Commun Chem 4, 6 (2021). https://doi.org/10.1038/s42004-020-00441-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-020-00441-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.