Abstract
This work investigates the connection between stochastic protein dynamics and function for the enzyme cyclophilin A (CypA) in wild-type form, and three variants that feature several mutations distal from the active site. Previous biophysical studies have suggested that conformational exchange between a ‘major’ active and a ‘minor’ inactive state on millisecond timescales plays a key role in catalysis for CypA. Here this hypothesis is addressed by a variety of molecular dynamics simulation techniques. Strikingly we show that exchange between major and minor active site conformations occurs at a rate that is 5 to 6 orders of magnitude faster than previously proposed. The minor active site conformation is found to be catalytically impaired, and decreased catalytic activity of the mutants is caused by changes in Phe113 motions on a ns-μs timescale. Therefore millisecond timescale motions may not be necessary to explain allosteric effects in cyclophilins.
Similar content being viewed by others
Introduction
A major goal of modern molecular biophysics is to clarify the connection between protein motions and enzymatic catalysis1,2,3. A wide range of experimental methods, e.g. neutron scattering, X-ray crystallography, NMR, or vibrational spectroscopy have been used to characterize internal protein motions occurring from femtosecond to second timescales4,5. While there is broad consensus that protein motions are implicated in catalysis, there is much debate around the role of conformational changes occurring on a millisecond timescale, and several studies have linked changes in millisecond protein motions with changes in enzymatic function6,7,8,9. However, it remains unclear whether such motions have a causal link to catalysis, or are merely a manifestation of the inherent flexibility of proteins over a broad range of timescales.
There have been vigorous debates about the meaning of dynamics in the context of enzymatic catalysis10,11,12. In the framework of transition state theory, the reaction rate is given by Eq. 1:
where T is the temperature and R the gas constant. The pre-exponential term A(T) includes contributions from non-statistical motions such as re-crossing or tunnelling, The exponential term involves the activation free energy of the chemical step \(\Delta G^\ddagger (T)\). If transitions between reactant states are fast compared to the time scale of the chemical reaction, \(\Delta G^\ddagger \left( T \right)\) is the free energy difference between the thermally equilibrated ensembles describing the reactant and transition states13,14. Non-statistical motions described by A(T) have typically been found to make a small contribution to rate constants with respect to the exponential term that involves equilibrium fluctuations of the protein and solvent degrees of freedom15.
The current work is concerned with the connection between rates of thermally equilibrated motions, and catalysis in enzymes. Specifically, the focus is on clarifying the nature of protein motions implicated in catalysis for the well-studied enzyme cyclophilin A (CypA). CypA is a member of the cyclophilin family of peptidyl-prolyl isomerases which catalyzes the cis/trans isomerization of amide groups in proline residues16. CypA plays an essential role in protein folding and regulation, gene expression, cellular signaling and the immune system. Notably, CypA is involved in the infectious activity and the viral replication of HIV-117. Accordingly, CypA has been the subject of structure-based drug design efforts for decades18,19,20. Because of its significance as a medical target, the catalytic mechanism of CypA has been the subject of extensive studies2,3,21,22,23,24,25,26,27,28,29,30. Computational studies have shown that the speedup of the of cis/trans isomerization rate of the prolyl peptide bond is a result of preferential transition state stabilization through selective hydrogen bonding interactions in the active site of CypA26,30. Figure 1a depicts key interactions between the substrate and active site residues, whereas Fig. 1b highlights the relevant ω angle of the substrate used to track the cis/trans isomerization reaction.
The active site of cyclophilin A. a Key residues in the active site of cyclophilin A that form hydrogen bonds (cyan sticks, dashed lines) or are in contact (purple sticks) with the transition state form of the peptide Ace-AAPF-Nme (orange sticks). For clarity, the Phe side-chain of the substrate is represented as transparent sticks. The distal residues Ser99, Cys115 and Ile97 are depicted in green. b The ω torsional angle of -Ala2-Pro3- is used to track the progression of the isomerization reaction between cis and trans forms
Elegant NMR relaxation experiments by Eisenmesser et al.27 have also characterized the existence of intrinsic motions in apo CypA that couple a ‘major’ state M with a ‘minor’ conformational state m with a rate constant \(k_{M \to m}\)= 60 s−1. Fraser et al. later used ambient temperature X-ray crystallographic data to determine a high-resolution structure of this CypA state m, revealing an interconversion pathway with the ‘major’ state M that involves coupled rotations of a network of side-chains involving residues Ser99, Phe113, Met61, and Arg55. To establish the relevance of this ‘minor’ state m to catalysis, the distal residue Ser99 was mutated to Thr99 (now only referred to as ST). Further X-ray and NMR measurements on the free enzyme confirmed that the ST mutant increased the population of the m state, while decreasing the conversion rate \(k_{M \to m}\) to 1 s−1 31. Remarkably, additional NMR experiments established that this 60-fold decrease in conversion rate between M and m states in the ST mutant correlates with a ca. 70-fold decrease in bidirectional isomerization rate (\(k_{iso} = k_{cis \to trans} + k_{trans \to cis}\)) of a model substrate with respect to wild-type (WT). The effect is comparable to rate decreases observed for mutations of key active site residues such as Arg5531. More recently, two further mutants were reported in an effort to rescue the lost enzymatic activity of ST. These mutations were S99T and C115S (now only referred to as STCS), or S99T, C115S, and I97V (now only referred to as STCSIV). The two newly introduced mutants recover the enzyme activity to some extent, which correlates with an increase in \(k_{M \to m}\) values32.
While this body of work suggested a link between millisecond time scale motions and catalysis in enzymes, there is currently no detailed mechanistic explanation for the decreased catalytic activity of the mutants. The present study uses a variety of extensive equilibrium and biased molecular dynamics (MD) simulations to clarify the link between catalytic activity and rates of molecular motions of CypA in wild-type and the three mutant variants. We show that the MD simulations reproduce well X-ray crystallography derived evidence for a shift in populations of major and minor active site conformations between the wild-type and mutant forms. Remarkably exchange between these active site conformations occurs at a rate that is five to six orders of magnitude faster than previously proposed. We show that the decrease in catalytic activity of the CypA mutants with respect to wild-type may be explained by changes in motions of residue Phe133 on a ns–μs timescale. Therefore millisecond time scale motions previously described in the literature may not be necessary to explain allosteric effects in cyclophilins.
Results
Major and minor conformations exchange on ns timescales
Fraser et al. have described the proposed ‘major’ and ‘minor’ states according to sets of values of χ1 (Phe113, Ser/Thr99), χ2 (Met61) and χ3 (Arg55) angles31,33. These dihedrals as well as the side-chain dihedrals χ1 of Ile97 and Cys115 were used to construct a Markov state model (MSM) to obtain quantitative information on thermodynamic and kinetic properties of the WT protein and the three experimentally studied mutants. The consistency of the MSMs was evaluated using standard protocols and by evaluating robustness of the findings with respect to a range of model parameters (See Supplementary Figures 1–3 and Supplementary Tables 1 and 2). In the case of WT the accuracy of the MSM was additionally evaluated by back-calculation of previously reported NMR observables34. The MSM yields predictions of observables that show broadly similar accuracy to that of the NMR ensembles of Chi et al.35 and Otter et al.36 (see Supplementary Figures 4). Thus the simulations were deemed sufficiently consistent with experimental data to warrant further analyses.
The X-ray structures of the key active site dihedrals in their dominantly populated states (if multiple occupancy is observed) are shown in Fig. 2a for WT and ST, Fig. 2b for WT and STCS, and Fig. 2c for WT and STCSIV mutants. The most striking feature of the ‘major’ and ‘minor’ conformations are the rotameric states of χ1 of Phe113 from the crystal structures, which in the ‘minor’ conformation is χ1 ≈ −60°. This will be referred to as the ‘out’ conformation. In contrast, the ‘major’ state χ1 ≈ 60°, takes an ‘in’ conformation. In Fig. 2d–f crystal structure occupancies for Phe113 χ1 are compared to the MSM-derived dihedral distributions comparing WT and ST, WT and STCS, and WT and STCSIV respectively. The simulations suggest that in apo WT the Phe113 ‘in’ and ‘out’ orientations are equally likely, which is consistent with the relatively similar occupancies of the two rotamers in the X-ray structure (occupancies = 0.63 and 0.37 respectively)31. In apo ST there is a significant population shift towards the ‘out’ orientation (χ1 = −60°), and the ‘in’ orientation has a marginal population (ca. 1%), see Fig. 2d. This agrees with the X-ray structure of ST where only the Phe113 ‘out’ rotamer is observed (occupancy = 1.0). This also agrees with J-coupling measurements that show the dominant Phe113 χ1 angle is ca. −60° in ST31. In the STCS and STCSIV mutants the ‘in’ rotamer is also destabilized with respect to wild-type but to a lesser extent (populations of ca. 16% and 17% respectively). Though only one ‘out’ rotamer was resolved in the X-ray structure of STCS (Fig. 2e), a major ‘out’ and a minor distorted ‘in’ rotamer (χ1 = + 31°, occupancy 0.21) are observed in the X-ray structure of STCSIV (Fig. 2f). Rotamers of other side-chain dihedrals of the key residues for all WT and mutants are found in Supplementary Figures 5 and 6.
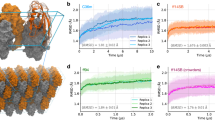
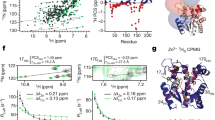
Comparison of X-ray and MSM-derived conformational preferences of Phe113. The X-ray structures of WT (blue and cyan) are compared with ST (red) (a), STCS (purple) (b) and STCSIV (dark and light green) (c)31,32. The MSM-derived probability distributions of χ1 in Phe113 for WT and ST (d), STCS (e) and STCSIV (f) are depicted as solid lines. The X-ray crystallography χ1 values are depicted for WT and mutants with their respective occupancies as dashed lines31,32
Surprisingly the Phe113 χ1 dihedral was observed to flip frequently in MD trajectories of 200 ns duration (Supplementary Figures 7–9), suggesting faster motions than determined by NMR experiments. Therefore the MSMs were used to obtain quantitative information on transition rates between ‘in’ and ‘out’ states as defined by the Phe113 χ1 rotamer. Table 1 summarises the MSM results. The exchange rates vary from 208 ± 9 μs−1 (ST) to 39 ± 3 μs−1 (STCS). Remarkably these values are five orders of magnitude faster than the exchange rates that have been determined by NMR measurements for motions involving Phe113.
The minor conformation is catalytically inactive
Given that the timescales of rotations of Phe113 in the four CypA variants appear much faster than previously suggested, attention turned next to substrate bound CypA simulations. Results from umbrella sampling (US) simulations were used to quantify the isomerization free energy profile for WT and the ST mutant and investigate the role of Phe113 motions in catalysis (See Supplementary Figure 10).
The isomerization free energy profiles for WT and ST mutant with the side-chain of the Phe113 in an ‘in’ and ‘out’ conformations are shown in Fig. 3a, b respectively. Ladani and Hamelberg28 have previously shown that fixed-charge classical force fields reproduce the energetics of amide bond rotation reasonably well due to relatively small changes in intramolecular polarization during this process. The calculated activation free energy for the uncatalyzed cis→trans isomerization process in water is consistent with experimental data (20.1 ± 0.1 kcal mol−1 vs ca. 19.3 kcal mol−1 for the related substrate Suc-AAPF-pNA at 283 K)37,38. The free energy profile for the substrate bound to CypA WT and ST in the ‘in’ conformation shows that the enzyme catalyzes the isomerization reaction in both directions via a transition state with a positive ω value (ca. 90–100°) equally well (Fig. 3a). There is a more significant decrease in activation free energy for trans→cis (ca. −6 kcal mol−1) than for cis→trans with less than 1 kcal mol−1 difference between WT and ST, because the cis form is more tightly bound to CypA than the trans form. According to Fig. 3b, there is no catalytic benefit from the ‘out’ conformation of the enzyme since the activation free energy of the isomerization reaction in CypA is similar to that of the substrate in water. The calculated free energy profiles for isomerization reactions in STCS and STCSIV show a similar trend (Supplementary Figure 11).
Energy profiles in the Phe113 ‘in’ and ‘out’ conformations. a Isomerization free energy profiles (in kcal mol−1) for the substrate AAPF in water (black), and bound to WT (blue) or ST mutant (red) forms of CypA with starting structures in the ‘in’ conformation. The free energies of the trans conformation were set to zero at ω = 180°. Error bars represent one standard error of the mean. The table shows the activation free energies for both directions of the isomerization reaction. (b) same as (a), but with stimulations starting in a ‘out’ configuration
Transition -state destabilization in the minor conformation
Further analysis of the US trajectories shows that for the simulations started in the ‘in’ configuration in both WT and ST the transition -state region (ω ca. 90–100°) is electrostatically stabilized by more negative Coulombic interactions between substrate and binding site atoms as shown in Fig. 4a. Figure 4b breaks down the different contribution of active site residues, showing that Arg55, Trp121, Asn102, His126, and Gln63 are important for the stabilization of the transition state ensemble via hydrogen bonding interactions as shown in Fig. 4e. In contrast, Fig. 4c shows that for simulations in the ‘out’ configuration no transition state stabilization through electrostatic interactions is observed, this is further reflected by the per-residue split of interaction energy contribution at the transition state in Fig. 4d and the lack of hydrogen bond formation in Fig. 4f. Hydrogen bonding probabilities for simulations from the ‘in’ and ‘out’ starting conformations are shown in Supplementary Figures 12–14. A similar picture holds for the STCS and STCSIV mutants (Supplementary Figure 15). Electrostatic interactions between the substrate and the solvent generally disfavour the transition state region in the ‘in’ conformation for all variants, consistent with a tightening of interactions of the active site residues with the transition state. For simulations carried out in the ‘out’ conformation no preferential electrostatic stabilization of a substrate state by the solvent is observed along the reaction coordinate, consistent with the lack of catalytic activity of CypA in this conformational state (Supplementary Figure 16)
Structural basis for the differential catalytic activity of the Phe113 in and out conformations. a Electrostatic energies between the substrate and active site residues as a function of the ω angle in WT (blue) and the ST systems (red) with simulations started from the ‘in’ configuration. b Average electrostatic energies per-active site residues at the transition state region from (a). c same as (a) with simulations started from the ‘out’ configuration. d same as (b) with simulations started in the ‘out’ conformation. Error bars denote the standard error of the mean. e and (f) typical hydrogen bonding pattern at the transition state for simulations started in the ‘in’ and ‘out’ conformation respectively
.
Preorganization explains decreased in activity of the mutants
Taken together the MSM and US data suggest a mechanistic explanation for the effect of distal mutations on the catalytic activity of cyclophilin A. In WT free form the enzyme rapidly interconverts between a catalytically active Phe113 ‘in’ form and a catalytically inactive Phe113 ‘out’ form. Because the interconversion rate between in and out forms (ca. 7 × 107 s−1) is faster than the substrate binding rate as suggested by NMR experiments (ca. 2 × 104 s−1, based on kon rate ca. 2 × 107 s−1 M−1 and substrate concentration ca. 1 mM)39 the free enzyme rapidly equilibrates between catalytically active and inactive forms before substrate binding (Fig. 5a). For the mutants, the interconversion rates between catalytically active and inactive forms are still within the μs−1 timescale, but the equilibrium is shifted towards the catalytically inactive form (Fig. 5b), thus the mutants are less pre-organized than WT and the overall catalytic activity is decreased.
Proposed mechanism for allosteric inhibition of cyclophilin A function. a Catalysis in WT, with the favoured route being the ‘in’ conformations, which are similarly populated to the ‘out’ conformations. b Catalysis in the mutants still occurs through the ‘in’ conformations, which have a lower population than the ‘out’ conformations
In the case of the ST mutant and WT forms, Fraser et al.31 have reported bi-directional on-enzyme isomerization rates \((k_{cis \to trans} + k_{trans \to cis})\) by NMR spectroscopy, and found a ratio of 68 ± 13 between WT and ST. According to the model proposed in Fig. 5 and by combining the MSM-derived populations and the US-derived activation free energies, a ratio of 12 < 46 < 176 can be derived from the simulations (see Supplementary Note 2 for details). The uncertainty from the simulations is larger than that of the measurements because small variations in activation free energies contribute large change in catalytic rates. Thus the model described in Fig. 5 appears consistent with experimental data for WT and ST. No bidirectional isomerization rates have been reported for the STCS and STCSIV mutants32. However, the STCS and STCSIV mutants show populations of the catalytically active Phe113 ‘in’ conformation that are intermediate between WT and ST, which is consistent with their increased catalytic activity with respect to ST.
A defining feature of this model is that the χ1 rotamers of a number of active-site side-chains such as Gln63, Ile/Val97, Phe113, Cys/Ser115 flip in WT and mutants on ns–μs timescales. Back-calculation of Cβ–Cγ order parameters shows that this effect is captured by a decrease in S2 values upon increasing the averaging window from 10 to 100 ns (Supplementary Figure 17). Motions on these timescales are too rapid to be detected by CPMG or CEST NMR experiments that have been used extensively to study μs–ms processes in cyclophilin A3,25,31,40,41. Likewise NMR relaxation experiments cannot detect motions on this timescale as they are limited to processes occurring faster than the tumbling time τc of cyclophilin A (ca. 10 ns)42. Residual Dipolar Couplings (RDCs) can, however, provide information about dynamic orientation of inter-nuclear vectors on the supra-τc time scale43. Such experiments have been reported for backbone and methyl-RDCs in ubiquitin43,44. Therefore the model predictions can be experimentally tested with combined nuclear spin relaxation and RDC based model-free analyses coupled with a labelling scheme that resolves χ1 side-chain motions43,45.
Discussion
This work highlights the potential of detailed molecular simulation studies to guide the interpretation of biophysical measurements for the elucidation of allosteric mechanisms in proteins46. Previous work has suggested that exchange on millisecond timescales between conformational states in CypA are linked to its catalytic cycle27, leading to a proposal for a slow exchange between a ‘major’ and a ‘minor’ state of a set of side chain rotamers linking distal residue Ser99 to active-site residues27,31. The present results do not support or reject this hypothesis because the MD simulations used here do not resolve motional processes occurring on timescales slower than microseconds. However a major finding of this study is that transitions between ‘in’ and ‘out’ rotamers of Phe 113 in WT and mutants occur on a time scale of ns–μs, thus five to six orders of magnitude faster than suggested by earlier NMR relaxation dispersion measurements31. Nevertheless the simulations reproduce well the population shifts in Phe113 rotamers observed in room-temperature X-ray crystallography experiments. This suggests that the X-ray structures may have resolved motional processes occurring on a distinct timescales from the processes resolved by CPMG experiments. Indeed in reported CPMG experiments the millisecond motions of Phe113 are coupled to a large network of ca. 30 residues31, whereas the χ1 rotameric flip observed in the simulations, is a largely local motion.
Nevertheless, the simulations suggest that a local ‘in’ to ‘out’ rotation of Phe113 is sufficient to abrogate catalysis in cyclophilin A, and variations of exchange parameters on the ns–μs timescale between these two conformational states appear sufficient to explain the decreased catalytic activity of the ST, STCS, STCSIV mutants with respect to WT. Therefore it is advisable to carry out additional experiments to confirm the existence of Phe113 χ1 rotations on the ns–μs timescale before causally linking catalysis to millisecond time scale motions. On the computational side, efforts should focus on advancing MD methodologies such that millisecond timescale processes observed in experiments can be resolved in atomistic details.
The contribution of protein flexibility on the ps–ns and μs–ms timescales to enzymatic catalysis has been the focus of several computational and experimental studies3,8,10,13,15,25,27,31,32,47. Our work suggests that more efforts should be directed at resolving conformational processes on the ns–μs timescale. This has important conceptual implications for enzyme design and optimization strategies.
Methods
Systems preparation
Models for apo/substrate bound human CypA of the WT and ST were prepared for MD simulations from PDB structures 3K0N (R = 1.39 Å) and 3K0O (R = 1.55 Å) respectively. For apo STCS and STCSIV two structures were prepared from PDB structures 6BTA (R = 1.5 Å) and 5WC7 (R = 1.43 Å) and also by mutating residues in WT using Schrödinger’s Maestro47. For WT the major conformation of 3K0N (altloc A, occupancy 0.63) was retained. For STCS and STCSIV the residues with higher occupancy were chosen for initial structures. Supplementary Tables 3 and 4 summarise all simulations conducted in this study. The proteins were solvated in a rhombic dodecahedron box of TIP3P water molecules with edges extending 1 nm away from the proteins and chloride counter-ions were added to neutralise the overall net-charge. The Charmm22* forcefield48 was used to describe protein atoms in the apo simulations because previous work from Papaleo et al.33 has shown that this forcefield reproduces more accurately conformational changes in CypA. Steepest descent minimized was used for 50,000 steps followed by equilibration for 100 ps in an NVT ensemble, and 100 ps NPT ensemble, with heavy protein atoms restraint using a harmonic force constant of 1000 kJ mol−1 nm−2.
Models of CypA WT and other mutants in complex with the Ace-AAPF-Nme substrate were prepared. The amber99sb forcefield was used for the complex simulations because Doshi and co-workers have reported optimised ω angle parameters for amides to simulate cis/trans isomerisation reactions49. The crystal structure of the CypA-cis AAPF peptide complex (PDB ID: 1RMH)50 was used to obtain a suitable orientation for the substrate in the active site of WT and other mutants. PDB structure 1RMH was aligned to the structure of WT and all mutants, and the N-terminal and C-terminal ends of the proteins and substrate were capped using Schrödinger’s Maestro47. In order to generate starting structures of ‘in’ and ‘out’ CypA-substrate complexes, MD simulations of CypA-substrate complexes (cis-conformation) were performed for 10 ns. For ST, STCS and STCSIV mutants the χ1 values of Phe113 were measured to monitor transitions between ‘out’ and ‘in’ rotamers. The last snapshot structure of ‘in’ and ‘out’ complexes structures were used as input US calculations. For WT complexes, only the ‘in’ rotamer was observed in a 10-ns MD simulation. Thus, US simulations of χ1 (Phe113) were performed serially to generate the ‘out’ (χ1 ≈ −60°) rotamer starting from the ‘in’ rotamer (χ1 ≈ 60°) using the software PLUMED251. Also, in order to retain the substrate in the active site, the distance between the proline ring of substrate and the phenyl rings of Phe113 and Phe60 were restrained using a force constant of 300 kJ mol−1 rad−2. Each US simulation was performed for 5 ns. The bias parameters and the restrained variables for the US of χ1 (Phe113) are summarised in Supplementary Table 7.
apo WT and mutant MD simulations
Eighty independent 200 ns MD trajectories of the apo WT, ST, STCS, and STCSIV proteins (20 each) were generated using Gromacs 5.052. For apo STCS and STCSIV the MD simulations were split between both structures prepared independently. A 2 fs time step was used, and the first 5 ns discarded for equilibration. Temperature was maintained at 300 K with a stochastic Berendsen thermostat53. The Parrinello-Rahman barostat was used for pressure coupling at 1 bar54. The Particle Mesh Ewald scheme was used for long-range electrostatic interactions with a Fourier grid spacing of 0.16 nm, and fourth-order cubic interpolation55. Short-range van der Waals and electrostatic interaction were cutoff at 1 nm. The LINCS algorithm was used to constrain all bonds56.
Markov state models
All MSM analysis was carried out with the software package pyemma version 2.3.257. The focus was on the side-chain motion of binding site residues. Details on which dihedral angles were used for TICA58 is given in Supplementary Table 8. A more detailed description of the MSM in particular with respect to best model selection is given in the SI. Clustering was done using all trajectory data from the WT and mutant trajectories using a set of 24 input coordinates, with selecting dominant coordinates using a 90% variance in TICA for the subsequent k-means clustering. Two hundred clusters were used to discretize the trajectory. With the same cluster assignment for all trajectories, MSM transition matrices were estimated, using the Bayesian MSM option, and choosing lagtimes 0.6 ns for WT, S99T, C115S, and I97V. Means and errors of observables (e.g. populations and MFPT) were estimated from the Bayesian MSM using the provided functions in pyEMMA. Membership assignments were based on the MSM microstate dihedral probabilities of being in the ‘in’ or ‘out’ state respectively. The microstate definition used for the MSMs is the same across the WT and all mutants. The MFPTs are estimated between the manually grouped two states depending on whether the Phe113 rotamer is ‘in’ or ‘out’ in the microstate. MSM validation and further details on the MSM can be found in the SI and in particular Supplementary Figures 1–3, Supplementary Tables 1–2 and Supplementary Note 159. MSM analyses were restricted to apo-enzymes because experimental data on the major to minor conformational exchange for the four variants is available for the apo forms only27,31,32.
US simulations
Series of US simulations60,61,62 of the ‘in’ and ‘out’ conformers were performed to compute free energy profiles along ω26,28,63,64. For substrate in solution, the initial structure of US was in a trans conformation taken from 10-ns equilibration run, while all protein-substrate complexes were in a cis conformation. For both of ‘in’ and ‘out’ US calculations, a standard harmonic potential was used to bias the ω angle towards a series of target values ωk spanning the interval [−180°,180°]. The force constants of the biasing potential and the spacing between ωk values were adjusted by trial and error in order to obtain a good overlap between probability distributions of neighbouring ωk values (Supplementary Tables 5 and 6 and Supplementary Figure 10). For ‘out’ US calculation the distances between the proline ring of substrate and the phenyl rings of Phe113 and Phe60 were restrained using a flat-bottom harmonic restraint with force constants of 200 kJ mol−1 rad−2 and 300 kJ mol−1 rad−2, respectively. Simulations were performed serially initially for 7 ns, with the starting conformation for a given target angle ωk taken from the preceding run performed at the neighbouring ωk+Δω value. Each US was then extended to 20 ns. A total of 22 (substrate in solution) or 24 (substrate bound to protein) umbrellas were used. In order to estimate uncertainties of free energy profiles six repeats of the entire procedure were performed for ‘in’ and ‘out’ US. All simulations were carried out using a PLUMED2 patched version of Gromacs 5.0 with simulation parameters identical to the previously described apo MD simulation protocols unless otherwise mentioned. The weighted histogram analysis method (WHAM) was used to produce a free energy profile from the pool of US simulations65.
Other trajectory analyses
Average proton–proton distances were derived as \(r_{ij}^{\mathrm{avg}} = \left\langle {r_{ij}^{ - 6}} \right\rangle ^{{\textstyle{{ - 1} \over 6}}}\) from snapshots sampled from the MSM of WT for comparison with NOEs and eNOEs-derived distance intervals66. 3J(HN, Hα), 3J(HN, \(C^\prime\)), and 3J(HN, Cβ) were also computed using Karplus equations and backbone dihedral angle values <ϕ> and <ψ> sampled from the MSM67.
Interaction energies between binding site residues (Arg55, Ile57, Phe60, Met61, Gln63, Asn102, Gln111, Phe113, Trp121, Leu122 and His126) and all atoms of the substrate were analysed with the Gromacs g_energy module, using snapshots from the US simulations. The probability distribution of distances between key residues and substrate atoms during the simulations were computed using the MDAnalysis library69.
Data availability
All input files and scripts used for the preparation, execution and analysis of the MD, MSM and US calculations are freely available at https://github.com/michellab/CypAcatalysis_input and are also provided as Supplementary Data 1–3. Other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Warshel, A. Dynamics of enzymatic reactions. Proc. Natl Acad. Sci. USA 81, 444–448 (1984).
McGowan, L. C. & Hamelberg, D. Conformational plasticity of an enzyme during catalysis: Intricate coupling between cyclophilin a dynamics and substrate turnover. Biophys. J. 104, 216–226 (2013).
Eisenmesser, E. Z., Bosco, D. A., Akke, M. & Kern, D. Enzyme dynamics during catalysis. Science 295, 1520–1523 (2002).
Rader, S. D. & Agard, Da Conformational substates in enzyme mechanism: the 120 K structure of alpha-lytic protease at 1.5 A resolution. Protein Sci. 6, 1375–1386 (1997).
Bu, Z. et al. A view of dynamics changes in the molten globulenative folding step by quasielastic neutron scattering. J. Mol. Biol. 301, 525–536 (2000).
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Wolf-Watz, M. et al. Linkage between dynamics and catalysis in a thermophilic-mesophilic enzyme pair. Nat. Struct. Mol. Biol. 11, 945–949 (2004).
Palmer, A. G., Kroenke, C. D. & Loria, J. P. NMR methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzym. 339, 204–238 (2001).
Schlegel, J., Armstrong, G. S., Redzic, J. S., Zhang, F. & Eisenmesser, E. Z. Characterizing and controlling the inherent dynamics of cyclophilin-A. Protein Sci. 18, 811–824 (2009).
Pisliakov, A. V., Cao, J., Kamerlin, S. C. L. & Warshel, A. Enzyme millisecond conformational dynamics do not catalyze the chemical step. Proc. Natl Acad. Sci. USA 106, 17359–17364 (2009).
Kamerlin, S. C. L. & Warshel, A. At the Dawn of the 21 st Century: Is Dynamics the Missing Link. Proteins Struct. Funct. Bioinforma. 78, 1339–1375 (2010).
Warshel, A. & Bora, R. P. Perspective: defining and quantifying the role of dynamics in enzyme catalysis. J. Chem. Phys. 144, 180901 (2016).
Kohen, A. Role of dynamics in enzyme catalysis: substantial versus semantic controversies. Acc. Chem. Res. 48, 466–473 (2015).
Wolfenden, R. Transition state analogues for enzyme catalysis. Nature 223, 704–705 (1969).
Delgado, M. et al. Convergence of theory and experiment on the role of preorganization, quantum tunneling, and enzyme motions into flavoenzyme-catalyzed hydride transfer. ACS Catal. 7, 3190–3198 (2017).
Go, S. F., Marahiel, M. A. & Gothel, S. F. Peptidyl-prolyl cis-trans isomerases, a superfamily of. Cell. Mol. Life Sci. 55, 423–436 (1999).
Li, J. et al. Discovery of dual inhibitors targeting both HIV-1 capsid and human cyclophilin A to inhibit the assembly and uncoating of the viral capsid. Bioorg. Med. Chem. 17, 3177–3188 (2009).
Dornan, J., Taylor, P. & Walkinshaw, M. Structures of Immunophilins and their Ligand Complexes. Curr. Top. Med. Chem. 3, 1392–1409 (2003).
Yang, Y. et al. Structure-based discovery of a family of synthetic cyclophilin inhibitors showing a cyclosporin-A phenotype in Caenorhabditis elegans. Biochem. Biophys. Res. Commun. 363, 1013–1019 (2007).
Georgiou, C. et al. Pushing the limits of detection of weak binding using fragment based drug discovery: identification of new cyclophilin binders. J. Mol. Biol. 429, 2556–2570 (2017).
Nagaraju, M., McGowan, L. C. & Hamelberg, D. Cyclophilin a inhibition: Targeting transition-state-bound enzyme conformations for structure-based drug design. J. Chem. Inf. Model. 53, 403–410 (2013).
Kofron, J. L., Kuzmic, P., Kishore, V., Colón-Bonilla, E. & Rich, D. H. Determination of kinetic constants for peptidyl prolyl cis-trans isomerases by an improved spectrophotometric assay. Biochemistry 30, 6127–6134 (1991).
Doshi, U., Holliday, M. J., Eisenmesser, E. Z. & Hamelberg, D. Dynamical network of residue-residue contacts reveals coupled allosteric effects in recognition, catalysis, and mutation. Proc. Natl Acad. Sci. USA 113, 4735–4740 (2016).
Camilloni, C. et al. Cyclophilin A catalyzes proline isomerization by an electrostatic handle mechanism. Proc Natl Acad. Sci. USA 111, 10203–10208 (2014).
Holliday, M. J., Camilloni, C., Armstrong, G. S., Vendruscolo, M. & Eisenmesser, E. Z. Networks of dynamic allostery regulate enzyme function. Structure 25, 276–286 (2017).
Hamelberg, D. & McCammon, J. A. Mechanistic insight into the role of transition-state stabilization in cyclophilin A. J. Am. Chem. Soc. 131, 147–152 (2009).
Eisenmesser, E. Z. et al. Intrinsic dynamics of an enzyme underlies catalysis. Nature 438, 117–121 (2005).
Ladani, S. T. & Hamelberg, D. Entropic and surprisingly small intramolecular polarization effects in the mechanism of cyclophilin A. J. Phys. Chem. B 116, 10771–10778 (2012).
Trzesniak, D. & van Gunsteren, W. F. Catalytic mechanism of cyclophilin as observed in molecular dynamics simulations: pathway prediction and reconciliation of X-ray crystallographic and NMR solution data. Protein Sci. 15, 2544–2551 (2006).
Ladani, S. T. & Hamelberg, D. Intricacies of interactions, dynamics and solvent effects in enzyme catalysis: a computational perspective on cyclophilin A. Mol. Simul. 40, 765–776 (2014).
Fraser, J. S. et al. Hidden alternative structures of proline isomerase essential for catalysis. Nature 462, 669–673 (2009).
Otten, R. et al. Rescue of conformational dynamics in enzyme catalysis by directed evolution. Nat. Commun. 9, 1314 (2018).
Papaleo, E., Sutto, L., Gervasio, F. L. & Lindorff-Larsen, K. Conformational changes and free energies in a proline isomerase. J. Chem. Theory Comput. 10, 4169–4174 (2014).
Vögeli, B., Kazemi, S., Güntert, P. & Riek, R. Spatial elucidation of motion in proteins by ensemble-based structure calculation using exact NOEs. Nat. Struct. Mol. Biol. 19, 1053–1058 (2012).
Chi, C. N. et al. A Structural ensemble for the enzyme cyclophilin reveals an orchestrated mode of action at atomic resolution. Angew. Chem. - Int. Ed. 54, 11657–11661 (2015).
Ottiger, M., Zerbe, O., Güntert, P. & Wüthrich, K. The NMR solution conformation of unligated human cyclophilin A. J. Mol. Biol. 272, 64–81 (1997).
Harrison, R. K. & Stein, R. L. Mechanistic studies of enzymic and nonenzymic prolyl cis-trans isomerization. J. Am. Chem. Soc. 114, 3464–3471 (1992).
Dugave, C. & Demange, L. Cis-trans isomerization of organic molecules and biomolecules: implications and applications. Chem. Rev. 103, 2475–2532 (2003).
Kern, D., Kern, G., Scherer, G., Fischer, G. & Drakenberg, T. Kinetic analysis of cyclophilin-catalyzed prolyl cis/trans isomerization by dynamic NMR spectroscopy. Biochemistry 34, 13594–13602 (1995).
Holliday, M. J., Armstrong, G. S. & Eisenmesser, E. Z. Determination of the full catalytic cycle among multiple cyclophilin family members and limitations on the application of CPMG-RD in reversible catalytic systems. Biochemistry 54, 5815–5827 (2015).
Holliday, M. J. et al. Structure and dynamics of GeoCyp: a thermophilic cyclophilin with a novel substrate binding mechanism that functions efficiently at low temperatures. Biochemistry 54, 3207–3217 (2015).
Gu, Y., Li, D. W. & Brüschweiler, R. NMR order parameter determination from long molecular dynamics trajectories for objective comparison with experiment. J. Chem. Theory Comput. 10, 2599–2607 (2014).
Lakomek, N. A. et al. Self-consistent residual dipolar coupling based model-free analysis for the robust determination of nanosecond to microsecond protein dynamics. J. Biomol. NMR 41, 139–155 (2008).
Farès, C. et al. Accessing ns-μs side chain dynamics in ubiquitin with methyl RDCs. J. Biomol. NMR 45, 23–44 (2009).
Kasinath, V., Valentine, K. G. & Wand, A. J. A 13C labeling strategy reveals a range of aromatic side chain motion in calmodulin. J. Am. Chem. Soc. 135, 9560–9563 (2013).
Michel, J. Current and emerging opportunities for molecular simulations in structure-based drug design. Phys. Chem. Chem. Phys. 16, 4465–4477 (2014).
Schrödinger Release 2017-4: Maestro, Schrödinger, LLC, New York, NY, 2017.
Piana, S., Lindorff-Larsen, K. & Shaw, D. E. How robust are protein folding simulations with respect to force field parameterization? Biophys. J. 100, L47–L49 (2011).
Doshi, U. & Hamelberg, D. Reoptimization of the AMBER force field parameters for peptide bond (Omega) torsions using accelerated molecular dynamics. J. Phys. Chem. B 113, 16590–16595 (2009).
Zhao, Y. & Ke, H. Crystal structure implies that cyclophilin predominantly catalyzes the Trans to Cis isomerization. Biochemistry 35, 7356–7361 (1996).
Tribello, G. A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED 2: New feathers for an old bird. Comput. Phys. Commun. 185, 604–613 (2014).
Abraham, M. J. et al. Gromacs: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Scherer, M. K. et al. PyEMMA 2: A Software Package for Estimation, Validation, and Analysis of Markov Models. J. Chem. Theory Comput. 11, 5525–5542 (2015).
Schwantes, C. R. & Pande, V. S. Improvements in Markov State Model construction reveal many non-native interactions in the folding of NTL9. J. Chem. Theory Comput. 9, 2000–2009 (2013).
Prinz, J. H. et al. Markov models of molecular kinetics: Generation and validation. J. Chem. Phys. 134, 174105 (2011).
Torrie, G. M. & Valleau, J. P. Nonphysical sampling distributions in Monte Carlo free-energy estimation: Umbrella sampling. J. Comput. Phys. 23, 187–199 (1977).
Hooft, R. W. W., van Eijck, B. P. & Kroon, J. An adaptive umbrella sampling procedure in conformational analysis using molecular dynamics and its application to glycol. J. Chem. Phys. 97, 6690–6694 (1992).
Northrup, S. H., Pear, M. R., Lee, C. Y., McCammon, J. A. & Karplus, M. Dynamical theory of activated processes in globular proteins. Proc. Natl Acad. Sci. 79, 4035–4039 (1982).
Agarwal, P. K. Cis/trans isomerization in HIV-1 capsid protein catalyzed by cyclophilin A: Insights from computational and theoretical studies. Proteins Struct. Funct. Genet. 56, 449–463 (2004).
Hamelberg, D., Shen, T. & McCammon, J. A. Phosphorylation effects on cis/trans isomerization and the backbone conformation of serine-proline motifs: Accelerated molecular dynamics analysis. J. Am. Chem. Soc. 127, 1969–1974 (2005).
Kumar, S., Rosenberg, J. M., Bouzida, D., Swendsen, R. H. & Kollman, P. A. THE weighted histogram analysis method for free energy calculations on biomolecules. I. The method. J. Comput. Chem. 13, 1011–1021 (1992).
Zagrovic, B. & Van Gunsteren, W. F. Comparing atomistic simulation data with the NMR experiment: how much can NOEs actually tell us? Proteins Struct. Funct. Genet. 63, 210–218 (2006).
Salvador, P. Dependencies of J-Couplings upon Dihedral Angles on proteins. Annu. Rep. NMR Spectrosc. 81, 185–227 (2014).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Naveen Michaud-Agrawal, ElizabethJ., Dening, ThomasB. & Woolf, O. B. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 31, 2967–2970 (2010).
Acknowledgements
Gratitude is expressed to Fernanda Duarte for thoughtful discussions about this work. J.M. is supported by a University Research Fellowship from the Royal Society. The research leading to these results has received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007–2013)/ERC grant agreement No. 336289. P.W. is supported by The Development and Promotion of Science and Technology Talents Project (DPST) Scholarship, Royal Thai Government. This project made use of time on ARCHER granted via the UK High-End Computing Consortium for Biomolecular Simulation, HECBioSim (http://hecbiosim.ac.uk), supported by EPSRC (grant no. EP/L000253/1).
Author information
Authors and Affiliations
Contributions
P.W., A.S.J.S.M.: Carried out experiments, analysed data, wrote manuscript. M.D.W.: Analysed data, wrote manuscript. J.M.: Designed study, analysed data, wrote manuscript. The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wapeesittipan, P., Mey, A.S.J.S., Walkinshaw, M.D. et al. Allosteric effects in cyclophilin mutants may be explained by changes in nano-microsecond time scale motions. Commun Chem 2, 41 (2019). https://doi.org/10.1038/s42004-019-0136-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-019-0136-1
This article is cited by
-
Deconstructing allostery by computational assessment of the binding determinants of allosteric PTP1B modulators
Communications Chemistry (2023)
-
Engineering and application of a biosensor with focused ligand specificity
Nature Communications (2020)
-
Temperature-jump solution X-ray scattering reveals distinct motions in a dynamic enzyme
Nature Chemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.