Abstract
In spite of extensive investigations, the force-dependent unfolding/rupturing rate k(F) of biomolecules still remains poorly understood. A famous example is the frequently observed switch from catch-bond behaviour, where force anti-intuitively decreases k(F), to slip-bond behaviour where increasing force accelerates k(F). A common consensus in the field is that the catch-to-slip switch behaviour cannot be explained in a one-dimensional energy landscape, while this view is mainly built upon assuming that force monotonically affects k(F) along each available transition pathway. In this work, by applying Kramers kinetic rate theory to a model system where the transition starts from a single native state through a pathway involving sequential peeling of a polymer strand until reaching the transition state, we show the catch-to-slip switch behaviour can be understood in a one-dimensional energy landscape by considering the structural-elastic properties of molecules during transition. Thus, this work deepens our understanding of the force-dependent unfolding/rupturing kinetics of molecules/molecular complexes.
Similar content being viewed by others
Introduction
The force-dependent lifetime of protein domains and protein–protein complexes not only has important biological implications, but also has been an intensively investigated topic in experimental studies1,2,3,4,5 and theoretical modelling6,7,8,9,10,11. A simple phenomenological expression of force-dependent unfolding/rupturing rate proposed by Bell6, \(k(F) = k_{\mathrm{0}}e^{\frac{{F\delta _0^ \ast }}{{k_{\mathrm{B}}T}}}\), has been the most applied model to explain experiments. k0 has often been interpreted as the zero-force transition rate. \(\delta _0^ \ast\), which has the dimension of length, is often referred to as the transition distance.
Bell’s model has been proven very powerful in explaining data recorded over a wide scope of experiments where unfolding/rupturing typically occurs at high forces (>100 pN)12. k(F) fitted to experimental data by Bell’s model has often been extrapolated to forces much lower than the force range where experimental data were recorded12. However, the validity of such extrapolation is questionable since deviations from Bell’s model are often observed at forces of several to tens of piconewtons (pNs)1,2,3,4,5. Among the reported deviations, the catch-to-slip switch behaviour is particularly intriguing, which refers to a phenomenon that k(F) anti-intuitively decreases as force increases over a certain low-force range, while it switches to a more expected slip-bond behaviour at a higher force range where force speeds up k(F). Since force of several to tens of pNs is a physiologically relevant force range13,14, the catch-to-slip switch behaviour of biomolecules could play an important role in their biological functions.
The catch-to-slip switch behaviour is characterised by a non-monotonic force dependence of k(F), which cannot be explained based on a one-dimensional transition pathway if k(F) along this pathway is a monotonic function of force. As a result, non-monotonic k(F) has been mainly explained by high-dimensional phenomenological models involving multiple competitive pathways or force-dependent selection of multiple native conformations that have access to different pathways7,15,16,17,18. Using a two-pathway model, for example, the overall transition rate is described by k(F) = k1(F) + k2(F), where k1(F) and k2(F) are the force-dependent transition rates along each pathway. Even k1(F) and k2(F) could be two monotonic functions of force, their combined force dependence with one of the pathways involving a negative transition distance can result in non-monotonicity of k(F), providing an explanation to the catch-to-slip switch behaviour. On the other hand, models based on force-dependent selection of multiple native conformations that have access to different pathways are much more complex and lack analytical simplicity for general cases7,15,16. In all of those high-dimensional models7,15,16,17,18, ki(F) (the subscript i represents the ith transition pathway) along each available transition pathway is assumed to be a monotonic function. In most models, ki(F) is assumed to follow Bell’s model.
In our recent work, on the basis of Arrhenius equation with a constant prefactor, we showed that the differential force–extension curves between the transition state and the native state have a complex effect on the force dependence of unfolding/rupturing rates of molecules19. This theory can explain complex deviation from Bell’s model, including the catch-to-slip switch behaviour, highlighting the importance of the structural–elastic properties of the native and transition states of molecules, which have been ignored in most of the previous models. However, this model derived in the framework of the Arrhenius equation is independent on the underlying energy landscape; therefore, it does not provide an answer to whether high dimensionality is necessary to explain the catch-to-slip switch behaviour. In view of the structural–elastic properties of the native and transition states of molecules as critical determinants of the force-dependent unfolding/rupturing rate, we would ask a question whether the catch-to-slip switch behaviour could be understood on a one-dimensional energy landscape by taking into account the structural–elastic properties of the molecule during transition along a single pathway.
In this work, using a model system where the transition follows a single pathway involving peeling of a polymer strand till it reaches the transition state, we have obtained an expression of k(F) derived within the framework of Kramers kinetic rate theory20. Here, we show that the derived k(F) can have a complex dependence on force that is affected by the geometry of how force is applied to the molecules. It predicts catch-to-slip switch behaviour for peeling off a pre-extended flexible polymer in the native state under shearing force geometry, which explains the k(F) data obtained from titin I27 domain unfolding over a force range from 4 to 90 pN. Therefore, our result demonstrates that the catch-to-slip switch behaviour can be understood based on a one-dimensional energy landscape under certain conditions.
Results
Derive k(F) based on Kramers rate theory
Kramers investigated a one-dimensional system concerning the escaping rate of a particle from an energy well overcoming a barrier that is significantly separated from the well20. Approximating the energy landscape U(x) near the well (at xw) and the barrier (at \(x_{b}\)) by Uw(x) ≈ U(xw) + 1/2kw(x − xw)2 and Ub(x) ≈ U(xb) − 1/2kb(x − xb)2, respectively, an expression of the particle escaping rate was obtained as \(k = D\frac{{\sqrt {k_{\mathrm{w}}k_{\mathrm{b}}} }}{{2\pi k_{\mathrm{B}}T}}e^{ - \frac{{{\mathrm{\Delta }}G^ \ast }}{{k_{\mathrm{B}}T}}}\), where D is the diffusion coefficient. The escaping rate implicitly depends on the parameters related to the shape of the free-energy landscape, namely the barrier height ΔG* = U(xb) − U(xw), and the stiffness parameters kw and kb.
Explicit dependence of Kramers rate equation on the shape of the energy landscape can be obtained by using an analytical expression of U(x), such as the linear–cubic function, \(U(x) = \frac{3}{2}{\mathrm{\Delta }}G_0^ \ast \frac{{x - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }} - 2{\mathrm{\Delta }}G_0^ \ast \left( {\frac{{x - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }}} \right)^3\). Here, \({\mathrm{\Delta }}G_0^ \ast\) and \(\delta _0^ \ast\) correspond to the energy barrier height (U(xb) − U(xw)) and the transition distance (xb − xw), respectively. Based on this energy landscape, the escaping rate becomes a function of \({\mathrm{\Delta }}G_0^ \ast\) and \(\delta _0^ \ast\), which can be derived as (see the Methods section)
Applying Kramers theory to understand protein unfolding or molecular complex rupturing, the variable x in U(x) has to be regarded as a properly defined transition coordinate. To avoid potential confusion with the molecular extension, hereafter, we use Ω to denote the transition coordinate. Ω describes the difference from the native state during transition, with Ω = 0 corresponding to the native state of the molecule. It is convenient to choose Ω such that its value increases as transition proceeds, which can be used to describe the state of the molecule during transition and to express the force-dependent energy landscape by UF(Ω) = U(Ω) + ΔΦF(Ω). Here, ΔΦF(Ω) is the force-induced change to the original energy landscape U(Ω). We use * to denote the transition state, which corresponds to the maximum point of UF(Ω). ΔΦF(Ω) can be expressed as21,22 (Supplementary Note 1 and Supplementary Figure 1)
where δz(F; Ω) = z(F; Ω) − z0(F) is the difference between the force-dependent extension of the molecule during transition (z(F; Ω)) and that in the native state (z0(F) = z(F; 0)).
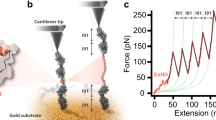
Many transitions such as force-dependent DNA strand separation follow a pathway involving sequential dissociation of bonds between a flexible polymer and the remaining structure23,24,25. For such transitions, a natural choice of the transition coordinate is n, which is the number of dissociated bonds, until reaching the transition state indicated by n*. The length of the molecule between the two force-attaching points, D(n) = L(n) + b(n), changes as the transition progresses. Here, L(n) is the contour length of the peeled polymer under force produced during transition, and b(n) is the linear distance between the two force-attaching points on the remaining folded structure. At a given point (n) during transition, under force F, the molecule has an extension of z(F; n) that is the average of its end-to-end distance projected along the force direction. In general, z(F; n) < D(n). At high forces where the entropic conformational fluctuation of the molecule is suppressed, z(F; n) approaches D(n) (Fig. 1).
Transitions through the single pathway of sequential bond separation. The figure shows the rupturing/unfolding transitions through sequential bond separation for the DNA structure (a) and the protein domain (b) under shearing force geometry. At low force Flow, the extension of the molecule in the transition state z(Flow; n) could be shorter than that of the native state z0(Flow). At high force Fhigh, the extension of the molecule in the transition state z(Fhigh; n) could be longer than that of the native state z0(Fhigh). As a result, the force-dependent extension change of the molecule between the transition state and the native state δz(F; n) (i.e., z(F; n) − z0(F)) could be a non-monotonic function of force under the shearing force pulling geometry
Any other quantities that are monotonically dependent on n can also be chosen as the transition coordinate. In order to better link to the kinetics parameters in Kramers theory (Eq. (1)), it is convenient to choose a transition coordinate that has the dimension of length. We propose to use \(\delta _{l_n} = D(n) - D(0) = L(n) + b(n) - b^{\mathrm{0}}\), the change of the molecular length during transition relative to that of the native state, as the transition coordinate. Here, D(0) = b0 is the linear distance between the two force-attaching points on the native state structure of the molecule (Fig. 1). In many cases such as DNA strand separation, \(\delta _{l_n}\) monotonically increases as n increases (Supplementary Note 2).
With this choice, \(\delta _{l_n} = 0\) corresponds to the native state, and \(\delta _{l_n} \, > \, 0\) corresponds to states during transition. The extension change of the molecule relative to the extension in the native state under force F during transition becomes a function of \(\delta _{l_n}\): \(\delta _z\left( {F;\delta _{l_n}} \right) = z\left( {F;\delta _{l_n}} \right) - z_0(F)\). Here, we clarify that since the native state is the reference point, the extension change during transition when n = 0 is always zero regardless of the value of force. For the simplest case where \(\delta _z\left( {F;\delta _{l_n}} \right)\) is proportional to \(\delta _{l_n}\), \(\delta _z\left( {F;\delta _{l_n}} \right)\) can be written as \(\delta _{l_n}\delta _{z,{\mathrm{unit}}}(F)\). Here, the dimensionless quantity δz,unit(F) is the extension change per unit molecular length change during transition. A famous example of such a simple case is the force-induced DNA/RNA strand separation transition (Supplementary Note 2).
Force-induced change to the energy landscape can be generally calculated by \({\mathrm{\Delta \Phi }}^{F}\left( {\delta _{l_n}} \right) = - \mathop {\int}\limits_0^F \delta _z\left( {f\prime ;\delta _{l_n}} \right)df\prime\) using Eq. (2). In the case when \(\delta _z\left( {F;\delta _{l_n}} \right) = \delta _{l_n}\delta _{z,{\mathrm{unit}}}(F)\), it becomes \(\Delta \Phi ^{F}(\delta _{l_n}) = - \delta _{l_n}\gamma (F)\), where \(\gamma (F) = \mathop {\int}\limits_0^F \delta _{z,{\mathrm{unit}}}(f{\prime})df{\prime}\) has the dimension of force. It can be clearly seen that, \({\mathrm{\Delta \Phi }}^{F}\left( {\delta _{l_{n^ \ast }}} \right) = - \delta _{l_{n^ \ast }}\gamma (F)\), is the force-induced barrier height change. Assuming that the transition state remains unchanged at different forces, we have \(\delta _{l_{n^ \ast }} = \delta _0^ \ast\). Force monotonically decreases/increases the original free-energy barrier height if γ(F) is a monotonically increasing/monotonically decreasing function of force. Interestingly, if γ(F) is a non-monotonic function of force, force may change the original barrier height in a non-monotonic manner.
The force-dependent energy landscape can be written as
where the linear–cubic function has been used to express the energy landscape in the absence of force, \(U\left( {\delta _{l_n}} \right) = \frac{3}{2}{\mathrm{\Delta }}G_0^ \ast \frac{{\delta _{l_n} - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }} - 2{\mathrm{\Delta }}G_0^ \ast \left( {\frac{{\delta _{l_n} - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }}} \right)^3\). Here, \(\delta _0^ \ast\) is the molecular length difference between the transition state and the native state, and \({\mathrm{\Delta }}G_0^ \ast\) is the original barrier height.
The resulting \(U^{F}(\delta _{l_n})\) is still a linear–cubic function, with \(\Delta G^ \ast (F) = {\mathrm{\Delta }}G_0^ \ast \left( {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}} \right)^{3/2}\) and \(\delta ^ \ast (F) = \delta _0^ \ast \sqrt {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}}\). Applying the Kramers rate theory, it is easy to show that (see Methods)
At large forces where the extension of the molecule at any transition point approaches the molecular length, δz,unit(F) ~1 and thus γ(F)~F. Substituting γ(F) with F in Eq. (4), the resulting expression of k(F) is identical to the Dudko–Hummer–Szabo model derived under the linear–cubic energy potential9. Therefore, the Dudko–Hummer–Szabo model can be considered as a special case of Eq. (4) under large forces where the entropic elasticity of molecules can be ignored. The differences between Eq. (4) and the Dudko–Hummer–Szabo model are discussed in the Discussion section.
Force-dependent DNA strand separation
We next apply the theory to investigate the force-dependent strand separation of double-stranded DNA (dsDNA) under unzipping (Fig. 2a) and shearing force geometry (Fig. 2b). Under the unzipping force geometry, breaking one dsDNA basepair produces two nucleotides of single-stranded DNA (ssDNA) under tension (Supplementary Figure 2). In contrast, under the shearing force geometry, breaking one basepair from one end results in the production of two nucleotides, but only one of them is under tension. In addition, it results in loss of one basepair of dsDNA under force (Supplementary Figure 3). Denoting bss and bds, the contour length per ssDNA nucleotide and dsDNA basepair, hss(F) and hds(F), the force–extension curves per unit contour length for ssDNA and dsDNA, it can be shown that δz,unit(F) = hss(F) for unzipping force geometry and \(\delta _{z,{\mathrm{unit}}}(F) = \frac{{b_{{\mathrm{ss}}}}}{{b_{{\mathrm{ss}}} - b_{{\mathrm{ds}}}}}h_{{\mathrm{ss}}}(F) - \frac{{b_{{\mathrm{ds}}}}}{{b_{{\mathrm{ss}}} - b_{{\mathrm{ds}}}}}h_{{\mathrm{ds}}}(F)\) for shearing force geometry (Supplementary Note 2).
Force-dependent DNA strand separation under different force geometries. a Schematic of the force-dependent DNA strand separation under unzipping force geometry. b Schematic of the force-dependent DNA strand separation under shearing force geometry. c The force–extension curve of dsDNA per basepair (solid line) and that of ssDNA per nucleotide (dashed line). d The curves of δz,unit(F) (black lines) and \(- \gamma (F)\delta _0^ \ast\) (red lines), calculated for shearing (solid lines) and unzipping (dashed lines) force geometries
Figure 2c shows the force–extension curves of dsDNA/ssDNA per basepair/nucleotide, calculated by an inextensible worm-like chain polymer model with the bending persistence length of 50 nm for dsDNA26 and 0.7 nm for ssDNA (typical value in 100 mM KCl)19 (Supplementary Notes 2 and 3, Supplementary Figure 4). At forces below ~5 pN, ssDNA has a shorter extension than that of dsDNA per nucleotide/basepair, while above ~5 pN, the ssDNA extension becomes longer than dsDNA extension. δz,unit(F) (Fig. 2d, black lines) and \({\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast ) = - \gamma (F)\delta _0^ \ast\) (Fig. 2d, red lines) calculated under the two different force geometries are monotonic functions of force under the unzipping force geometry, and non-monotonic functions of force under the shearing force geometry.
Assuming \({\mathrm{\Delta }}G_{\mathrm{0}}^ \ast = 20\) kBT and \(\delta _0^ \ast = 3\,{\mathrm{nm}}\), we plotted \(U^{\mathrm{F}}(\delta _{l_n})\) by Eq. (3) under the unzipping and shearing force geometries (Fig. 3a). The results reveal drastically different effects of force on the change of the energy landscape between the two distinct force geometries. Figure 3b shows k(F)/k0 predicted by Eq. (4) under the unzipping (dashed line) and shearing (solid line) force geometries. Under the unzipping force geometry, k(F) monotonically increases with force, demonstrating a slip-bond kinetics. In contrast, under the shearing force geometry, k(F) decreases as force increases at <6 pN forces, while it increases as force increases at >6 pN forces, demonstrating a catch-to-slip switching kinetics.
Force-dependent energy landscape and the DNA strand separation rates. a The figure shows the energy landscape described by Eq. (3) with \({\mathrm{\Delta }}G_0^ \ast = 20\,\,k_{\mathrm{B}}T\) and \(\delta _0^ \ast = 3\,{\mathrm{nm}}\), for F = 0 pN (black solid line), 6 pN (grey lines) and 25 pN (red lines), for unzipping (dotted lines) and shearing (dashed lines) force geometries. b Force-dependent DNA strand separation rate normalised by the zero-force rate predicted using Eq. (4) for the unzipping (dashed line) and shearing (solid line) force geometries
Although these predictions, in particular the catch-to-slip switching behaviour of DNA strand separation under the shearing force geometry, are awaiting for future experimental tests, from the theoretical point of view, this example is sufficient to demonstrate that the catch-to-slip switching behaviour can occur on a one-dimensional energy landscape.
Titin I27 unfolding transition
Eq. (4) can also be applied to cases where \(\delta _z(F;\delta _{l_n})\) monotonically depends on \(\delta _{l_n}\), but is not perfectly proportional to \(\delta _{l_n}\), by writing \(\delta _z\left( {F;\delta _{l_n}} \right) = \delta _{l_n}\bar \delta _{z,{\mathrm{unit}}}(F)\). Here, \(\bar \delta _{z,{\mathrm{unit}}}(F)\) is a “characteristic” extension change per unit length change, which should be calculated by \(\bar \delta _{z,{\mathrm{unit}}}(F) = \frac{{\delta _z(F;\delta _{\mathrm{0}}^ \ast )}}{{\delta _{\mathrm{0}}^ \ast }}\) to ensure that \(- \gamma (F)\delta _0^ \ast\) has a proper meaning of the force-dependent conformational free energy difference between the transition state and the native state (i.e., \({\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast )\)). However, the calculation of \(\bar \delta _{z,{\mathrm{unit}}}(F)\) depends on prior knowledge of \(\delta _{\mathrm{0}}^ \ast\), which itself is a model parameter to be determined by fitting Eq. (4) to experimental data. To solve this problem, we propose to treat the number of broken bonds in the transition state, n*, as a fitting parameter. For each testing value of n*, we calculate \(\bar \delta _{z,{\mathrm{unit}}}(F) = \frac{{\delta _z( {F;\delta _{l_{n^ \ast }}})}}{{\delta _{l_{n^ \ast }}}}\) and fit the experimental data. When the best fitting is achieved, the best-fitting value of n* along with the best-fitting values of other model parameters, including k0, \(\delta _{\mathrm{0}}^ \ast\) and \({\mathrm{\Delta }}G_{\mathrm{0}}^ \ast\) are determined. A self-consistency check should be performed by comparing the values of \(\delta _{l_{n^ \ast }}\) evaluated at the best-fitting value of n* with the best-fitting value of \(\delta _{\mathrm{0}}^ \ast\), which should be the same when ideal fitting is achieved.
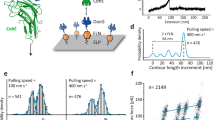
We demonstrate the application of the theory using the force-dependent unfolding of titin I27 immunoglobulin (Ig) domain as an example. We recently reported that k(F) of mechanical unfolding of titin I27 domain exhibits a catch-to-slip switching behaviour at low-force range2. Previous steered molecular dynamics (MD) simulation studies have suggested that the unfolding of I27 primarily follows a transition pathway of peeling the N-terminal strand (the A–A′ peptide), during which the residues in the A–A′ peptide are sequentially peeled off from the remaining folded core until reaching the transition state23,24,25. AFM experiments show that at forces below 100 pN, titin I27 unfolding starts from the native state with the A–A′ strand stacked with both the B and G strands27,28. According to this hypothetical transition pathway suggested by the previous steered MD simulations and AFM experiments, \(\delta _{l_{n^ \ast }} = L(n^ \ast ) + b(n^ \ast ) - b^0\), where L(n*) = n* × 0.38 nm is the contour length of the peeled peptide29, b(n*) is the length connecting the n* + 1 residue and the C-terminus of I27 and b0 ~ 4.32 nm is the length between N- and C-termini of I27 in its native state (Fig. 4a). L(n*) (Table 1, column 2), b(n*) (Table 1, column 3) and b0 can be determined from the structure of I27 (PDB ID:1TIT). Thus, \(\delta _{l_{n^ \ast }}\) of I27 can be calculated (Table 1, column 4).
Force-dependent unfolding of I27. a The figure shows the molecular length change of I27 when there are n* number of residues on the A–A′ peptide dissociated from the remaining folded core. b Best-fitting curves to experimental data of I27 reported in our previous study [2] according to Eq. (4) with \({\mathrm{\Delta }}G_0^ \ast = 5\,\,k_{\mathrm{B}}T\) (grey dotted line), \({\mathrm{\Delta }}G_0^ \ast = 20\,\, k_{\mathrm{B}}T\) (grey solid line) and \({\mathrm{\Delta }}G_0^ \ast = 200\,k_{\mathrm{B}}T\) (black dotted line) and according to the high-barrier approximation (red dashed line). The best-fitting parameters are (k0 = 0.002 s−1, A = 0.7 nm and \(\delta _{\mathrm{0}}^ \ast = 1.4\,{\mathrm{nm}}\)) for \({\mathrm{\Delta }}G_0^ \ast = 5\, k_{\mathrm{B}}T\), (k0 = 0.001 s−1, A = 0.7 nm and \(\delta _{\mathrm{0}}^ \ast = 1.3\,{\mathrm{nm}}\)) for \({\mathrm{\Delta }}G_0^ \ast = 20\,k_{\mathrm{B}}T\), (k0 = 0.001 s−1, A = 0.7 nm and \(\delta _{\mathrm{0}}^ \ast = 1.2\,{\mathrm{nm}}\)) for \({\mathrm{\Delta }}G_0^ \ast = 200\,k_{\mathrm{B}}T\) and (\(k_0^A = 0.001\,{\mathrm{s}}^{ - 1}\), A = 0.7 nm and \(\delta _{\mathrm{0}}^ \ast = 1.2\,{\mathrm{nm}}\)) for the high-barrier approximation using the formula of \(k(F) = k_0^{\mathrm{A}}e^{ - \frac{{{\mathrm{\Delta \Phi }}^{\mathrm{F}}\left( {\delta _0^ \ast } \right)}}{{k_{\mathrm{B}}T}}} = k_0^{\mathrm{A}}e^{\frac{{\gamma (F)\delta _0^ \ast }}{{k_{\mathrm{B}}T}}}\). c The black and red lines are the calculated curves of \(\bar{\delta}_{z,\text{unit}}(F)\) and \(- \gamma (F)\delta _0^ \ast\), respectively
\(\delta _{l_{n^ \ast }}\), \(\bar \delta _{z,{\mathrm{unit}}}(F)\) and γ(F) are calculated for a set of testing values of \(n^ \ast = 1,2, \cdots ,15\), where \(\bar \delta _{z,{\mathrm{unit}}}(F) = \frac{{\delta _z( {F;\delta _{l_{n^ \ast }}})}}{{\delta _{l_{n^ \ast }}}}\). The extension change at each testing value of n*, \(\delta _z( {F;\delta _{l_{n^ \ast }}})\), is calculated based on the known force–extension curves of a freely rotating rigid structure (Supplementary Figure 5) and a flexible peptide polymer with a certain bending persistence of A ∈ (0.5,1) nm5,29,30,31 (Supplementary Note 4), which is also treated as a fitting parameter. At each value n*, the I27 experimental data2 were fitted using Eq. (4) with the constraints for fitting parameters of \(\delta _{\mathrm{0}}^ \ast \, > \, 0\, {\mathrm{nm}}\), k0 > 0 s−1 and A ∈ (0.5,1) nm. A fixed value \({\mathrm{\Delta }}G_{\mathrm{0}}^ \ast = 20\, k_{\mathrm{B}}T\) was used in the fitting for reasons explained later.
The best fitting is achieved at \(n^ \ast = 8,9, \cdots ,13,14\) with a similar residual sum of squares (Table 1, the last column). In addition, similar values of best-fitting \(\delta _{\mathrm{0}}^ \ast\) of 1.2–1.3 nm are obtained at all these candidate values of n*. However, the best consistency between \(\delta _{\mathrm{0}}^ \ast\) and \(\delta _{l_{n^ \ast }}\) is achieved only at n* of 10–12 (Table 1, comparing between column 4 and column 7). Therefore, these results suggest that the transition state of I27 corresponds to a structure with 10–12 residues in the A–A′ strand peeled away from the remaining folded core, which is consistent with previous predictions based on MD simulations and single-molecule force spectroscopy experiments that suggest 12–13 peeled residues in the transition state of I272,23,24,25. At the values of n* = 10–12, the best-fitting value of the bending persistence length for a peptide polymer is A = 0.6–0.7 nm, which is close to the values determined in previous experimental measurements using a lock-in force spectroscopy technique and magnetic tweezers5,29,30,31.
At any values of \({\mathrm{\Delta }}G_0^ \ast \, > \, 5\) kBT, the fitting always converges to a narrow range of k0 \(\in\) (0.0013, 0.0020) s−1, A ∈ (0.68, 0.69) nm and \(\delta _0^ \ast \in (1.23,1.40)\,{\mathrm{nm}}\) (Fig. 4b, using n* = 12 for example), indicating that the fitting is insensitive to the values of \({\mathrm{\Delta }}G_0^ \ast \gg 5\) kBT. In addition, the fitting result is similar to that fitted with \(k(F) = k_0^{\mathrm{A}}e^{ - \frac{{{\mathrm{\Delta \Phi }}^{\mathrm{F}}(\delta _0^ \ast )}}{{k_{\mathrm{B}}T}}} = k_0^{\mathrm{A}}e^{\frac{{\gamma (F)\delta _0^ \ast }}{{k_{\mathrm{B}}T}}}\) (Fig. 4b), which is only valid when \(|\gamma (F)\delta _{\mathrm{0}}^ \ast | \ll {\mathrm{\Delta }}G_{\mathrm{0}}^ \ast\) (see Methods). Noting that \({\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast ) = - \gamma (F)\delta _{\mathrm{0}}^ \ast\), the agreement between the two fittings suggests that \(|{\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast )| \ll \Delta G_0^ \ast\), (i.e., the force-induced change of the barrier height is a small perturbation to the original barrier height). Over the force range of the experimental data, the maximal value of \(|{\mathrm{\Delta \Phi }}^{\mathrm{F}}(\delta _0^ \ast )|\) is ~6 kBT (Fig. 4c). These results imply that \({\mathrm{\Delta }}G_0^ \ast\) should be significantly larger than 6 kBT for I27 unfolding.
Figure 4c shows \(\bar \delta _{z,{\mathrm{unit}}}(F)\) (black line) and \(- \gamma (F)\delta _{\mathrm{0}}^ \ast\) (red line) calculated at n* = 12 with the fitting parameters of A = 0.7 nm and \(\delta _{\mathrm{0}}^ \ast = 1.3\,{\mathrm{nm}}\). As force increases through ~21 pN, \(\bar \delta _{z,{\mathrm{unit}}}(F)\) switches from negative to positive values. As a result, \(- \gamma (F)\delta _{\mathrm{0}}^ \ast\) is also a non-monotonic function that switches from an increasing function to a decreasing function as F increases through ~21 pN. The complex force-dependent extension changes during transition and the resulting non-monotonic \(- \gamma (F)\delta _{\mathrm{0}}^ \ast\) result in the observed catch-to-slip behaviour of I27.
Discussion
In summary, we have discussed the force-dependent two-state unfolding/rupturing rates of molecules/molecular complexes over a one-dimensional energy landscape using a model system where the transition is a process of peeling of a flexible polymer strand from the remaining folded core until reaching the transition state. The peeling is assumed to follow a path involving sequential bond dissociation between the polymer and the remaining folded core, which ensures that the energy landscape can be described by a one-dimensional transition coordinate (i.e., the number of dissociated bonds or the change in the molecular length). By modelling the energy landscape at zero force with a linear–cubic function, we derived a new expression of k(F) for mechanical unfolding/rupturing of biomolecules. Careful analysis of this expression of k(F) reveals a number of important aspects of force-dependent unfolding/rupturing rate of molecules/molecular complexes, which were previously not acknowledged.
The most important finding is that the catch-to-slip switch behaviour can occur on a one-dimensional energy landscape, which is in sharp contrast to the current consensus that such behaviour can only be understood based on a multi-dimensional energy landscape7,15,16,17,18. In all previous models, a monotonic function of ki(F) is assumed in each available transition pathway. This is the reason why the catch-to-slip switch behaviour, which implies a non-monotonic dependence on force, cannot be explained in a one-dimensional energy landscape. Two types of multi-dimensional models have been proposed: (1) single-state multi-pathway models where the transitions starting from the same native state can follow different pathways, and (2) multistate models where the transitions can start from different “native” states, each following a single pathway leading to unfolding/rupturing. Among the native states in the second type of models, reversible transitions are allowed and the rates of the reversible transitions are assumed to be force-dependent.
In single-state multi-pathway models, the catch-to-slip switch behaviour can be explained. However, it requires that ki(F) of one pathway is a monotonically deceasing function of force and at least in one pathway ki(F) is a monotonically increasing function of force. This can be clearly seen using a two-pathway model where k(F) = k1(F) + k2(F). The catch-to-slip switch behaviour implies the existence of a minimum of k(F), which in turn implies the existence of solution to the following function: \(k\prime (F) = k_1^\prime (F) + k_2^\prime (F) = 0\), where′ indicates a derivative of F. Clearly, \(k_1^\prime (F)\) and \(k_2^\prime (F)\) must have opposite signs. In multistate models, the catch-to-slip switch behaviour can be explained without assuming a monotonically decreasing function for any of the transition rates along each pathway. However, it requires a force-dependent switch from a faster transition path starting from a “native” state to a slower transition path starting from another “native” state as force increases.
Our model is a single-state single-pathway (i.e., one-dimensional) model; however, it can explain the catch-to-slip switch behaviour. The key mechanism underlying the success of our model is that the unfolding/rupturing rate k(F) is a non-monotonic function, which is a natural result from the structural–elastic properties of the molecules during transition. A molecule in any given structural state undergoes conformational thermal fluctuations, which are affected by force applied to the molecule. Such fluctuations result in a force–extension relation of the molecule in a given state, with the extension in general shorter than the molecule length at that state. Depending on the structural–elastic nature of the molecule and the pulling force geometry applied to the molecule, we show that with the generation of a flexible polymer under shearing force geometry, the extension change during transition could be negative at lower forces and positive at higher forces. This in turn results in a switch of k(F) from a decreasing function to an increasing function as force increases, i.e., the catch-to-slip switch behaviour.
Another important point we want to stress is that all one-dimensional unfolding/rupturing models can be more generally interpreted as effective projections from a higher-dimensional free-energy landscape, as elegantly discussed in previous study10. In our work, the number of separated bonds (n) is chosen as the transition coordinate. It is proper when the molecular extension at any value of n reaches equilibrium, which allows us to use equilibrium force–extension curves to calculate the force-dependent free-energy change during transition. If this condition is unsatisfied, the transition coordinate should be described by both the number of separated bonds (n) and the molecular extension (x), which results in a two-dimensional free-energy landscape.
Under conditions where γ(F) can be approximated by γ(F)~F, the expression of Eq. (4) is identical to the Dudko–Hummer–Szabo model derived under the linear–cubic energy potential9. Therefore, the Dudko–Hummer–Szabo model is asymptotic to Eq. (4) only under conditions, such as a large applied force, where the entropic elasticity of biomolecules can be ignored. In the Dudko–Hummer–Szabo model, the molecular extension is chosen as the transition coordinate. However, due to the entropic conformational fluctuation at low forces, the molecular extension becomes a function of force and is improper to be used as the transition coordinate. In contrast, our expression is derived based on the force-independent molecular length change during transition; therefore, it can serve as a proper transition coordinate, regardless of the force applied to the molecule. As a result, in spite of the structural similarity between Eq. (4) and the Dudko–Hummer–Szabo model, the underlying physics and application scope are significantly different between the two models. For instance, k(F) predicted by the Dudko–Hummer–Szabo model is a monotonically increasing \(\left( {\delta _0^ \ast \, > \, 0} \right)\) or decreasing \(\left( {\delta _0^ \ast \, < \, 0} \right)\) function; therefore, it cannot explain the catch-to-slip switch behaviour typically observed at a low-force range.
In our previous work19, by analysing the force-dependent change of the energy barrier height \({\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast )\) (i.e., the additional change to the free-energy difference between the transition state and the native state), we obtained an expression of the force-dependent rate \(k(F) = k_0^{A}e^{ - \frac{{{\mathrm{\Delta \Phi }}^{F}\left( {\delta _0^ \ast } \right)}}{{k_{\mathrm{B}}T}}}\) derived based on the Arrhenius equation with a constant prefactor, where \({\mathrm{\Delta \Phi }}^{F}(\delta _0^ \ast )\) was calculated based on the structural–elastic properties of the molecule between the transition and the native states. We showed that the expression can explain catch-to-slip switch behaviour under a certain pulling force geometry, highlighting the importance of the structural–elastic properties of a molecule as crucial determinants of the force-dependent transition rate. However, as the expression was derived independent from the energy landscape, it does not provide an answer concerning whether the catch-to-slip switch behaviour could be understood on a one-dimensional energy landscape. The question concerning whether the catch-to-slip switch behaviour could be allowed in a one-dimensional energy landscape has been answered by the work described in this paper.
We emphasise that the model described in this paper is to demonstrate that it is possible to have catch-to-slip switch behaviour on a one-dimensional energy landscape, which overturns the widely accepted belief that the catch-to-slip switch behaviour can only be interpreted on a high-dimensional energy landscape. In addition, it is also possible to apply Eq. (4) to fit experimental data to obtain information on the barrier height and transition distance of the underlying energy landscape. For such applications, several requirements need to be met: (1) prior knowledge of the transition pathway is known, (2) the energy landscape can be described by a one-dimensional sequential bond-breaking process and (3) the energy landscape can be approximated using a linear–cubic function. This is the case of force-dependent strand separation of DNA and RNA duplexes and mechanical unfolding of some protein domains, such as the titin I27 domain.
Like any other models derived based on a preassumed energy landscape, one should be cautious to apply the model to explain experimental data since the shape of the energy landscape underlying the experiments could be significantly different from that assumed in the model derivation. Fortunately, in many experiments, the force-dependent change of the barrier height at a low-force regime is much smaller than the original barrier height. Under such conditions, the force-dependent transition rate can be approximated by the Arrhenius equation with a constant prefactor, \(k(F) = k_0^{\mathrm{A}}e^{ - \frac{{{\mathrm{\Delta \Phi }}^{\mathrm{F}}\left( {\delta _0^ \ast } \right)}}{{k_{\mathrm{B}}T}}}\), which only depends on the force-induced change of barrier height and is insensitive to the details of the transition pathways as well as dimensionality. As shown in our previous study, under this condition for typical unfolding/rupturing transitions, at forces greater than 5 pN, k(F) has a simple asymptotic expression: \(k(F) = \tilde k_{\mathrm{0}}e^{\beta \left( {\sigma F + \alpha F^2/2 - \eta F^{1/2}} \right)}\), which contains three structure–elasticity-dependent model parameters: σ = L(n*) + b(n*) − b0 − (kBT/γ* − kBT/γ0), α = b(n*)/γ* −\(b^0 / \gamma^0\) and \(\eta = L(n^ \ast )\sqrt {k_{\mathrm{B}}T/A}\). Here, γ0 and γ* are the stretching rigidity of the folded structure in the native state and that of the folded core in the transition state of the molecule, respectively; A is the persistence length of the flexible polymer peeled off in the transition state19.
Our analysis for force-dependent strand separation of a short DNA duplex predicts that the force dependence of the strand separation rate strongly depends on the pulling force geometry. Under unzipping force geometry, k(F) monotonically increases with force (i.e., a slip bond), while under shearing force geometry, k(F) exhibits a non-monotonic, catch-to-slip switching behaviour. These predictions warrant future experimental validation.
Methods
Derivation of Eq. (1)
Eq. (1) is derived based on the linear–cubic function, \(U(x) = \frac{3}{2}{\mathrm{\Delta }}G_0^ \ast \frac{{x - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }} - 2{\mathrm{\Delta }}G_0^ \ast \left( {\frac{{x - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }}} \right)^3\), where \({\mathrm{\Delta }}G_0^ \ast\) and \(\delta _0^ \ast\) are two parameters. The linear–cubic potential has a well and a barrier at the position of xw = 0 and \(x_{\mathrm{b}} = \delta _0^ \ast\), respectively. It can be easily shown that the energy barrier height, ΔG* = U(xb) − U(xw), is \(\Delta G_0^ \ast\), and the transition distance, δ* = xb − xw, is \(\delta _0^ \ast\). Approximating the energy landscape U(x) near the well (xw = 0) and the barrier \(\left( {x_{\mathrm{b}} = \delta _0^ \ast } \right)\) by Uw(x) ≈ U(xw) + 1/2kw(x − xw)2 and Ub(x) ≈ U(xb) − 1/2kb(x − xb)2, kw and kb can be obtained in terms of \({\mathrm{\Delta }}G_0^ \ast\) and \(\delta _0^ \ast\): \(k_{\mathrm{w}} = k_{\mathrm{b}} = 6{\mathrm{\Delta }}G_0^ \ast /\delta _0^{ \ast 2}\). Substituting the expressions of ΔG*, kw and kb into the Kramers equation \(k = D\frac{{\sqrt {k_{\mathrm{w}}k_{\mathrm{b}}} }}{{2\pi k_{\mathrm{B}}T}}e^{ - \frac{{{\mathrm{\Delta }}G^ \ast }}{{k_{\mathrm{B}}T}}}\), we can obtain Eq. (1) as \(k_0 = \frac{{3D}}{{\pi \delta _0^{ \ast 2}}}\frac{{{\mathrm{\Delta }}G_0^ \ast }}{{k_{\mathrm{B}}T}}e^{ - \frac{{{\mathrm{\Delta }}G_0^ \ast }}{{k_{\mathrm{B}}T}}}\).
Derivation of Eq. (4)
Force can change the shape of the free-energy landscape of molecules; therefore, the energy barrier height \({\mathrm{\Delta }}G_0^ \ast\) and the transition distance \(\delta _0^ \ast\) in Eq. (1) can be functions of force. As a result, the force-dependent transition rate can be expressed as
according to Eq. (1). Eq. (4) is derived based on the force-dependent free-energy landscape, which is a linear–cubic function, \(U^{F}\left( {\delta _{l_n}} \right) = \frac{3}{2}{\mathrm{\Delta }}G_0^ \ast \frac{{\delta _{l_n} - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }} - 2{\mathrm{\Delta }}G_0^ \ast \left( {\frac{{\delta _{l_n} - 1/2\delta _0^ \ast }}{{\delta _0^ \ast }}} \right)^3 - \delta _{l_n}\gamma (F)\). The linear–cubic potential has a well and a barrier at the position of \(\delta _{l_n,{\mathrm{w}}} = \frac{{\delta _0^ \ast }}{2} - \frac{{\delta _0^ \ast }}{2}\sqrt {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}}\) and \(\delta _{l_n,{\mathrm{b}}} = \frac{{\delta _0^ \ast }}{2} + \frac{{\delta _0^ \ast }}{2}\sqrt {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}}\), respectively. It can be easily shown that the energy barrier height, \({\mathrm{\Delta }}G^ \ast (F) = U^{\mathrm{F}}\left( {\delta _{l_n,{\mathrm{b}}}} \right) - U^{\mathrm{F}}\left( {\delta _{l_n,{\mathrm{w}}}} \right)\), is \({\mathrm{\Delta }}G_0^ \ast \left( {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}} \right)^{3/2}\), and the transition distance, \(\delta ^ \ast (F) = \delta _{l_n,{\mathrm{b}}} - \delta _{l_n,{\mathrm{w}}}\), is \(\delta _0^ \ast \sqrt {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}}\). Substituting the expressions of ΔG*(F) and δ*(F) into Eq. (5), we can obtain
Combined with Eq. (1) for the zero-force transition rate, it can be shown that the transition rate under force F becomes
High-barrier approximation of Eq. (4)
In the case when the force-dependent change of barrier height is much smaller than the original barrier height, it suggests that \(|\gamma (F)\delta _{\mathrm{0}}^ \ast | \ll \Delta G_{\mathrm{0}}^ \ast\), or equivalently \(\left| {\frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}} \right|\sim 1\). It is obvious that \(\sqrt {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}} \sim 1\) and \(\left( {1 - \left( {1 - \frac{{2\gamma (F)\delta _0^ \ast }}{{3{\mathrm{\Delta }}G_0^ \ast }}} \right)^{3/2}} \right)\sim \frac{{\gamma (F)\delta _0^ \ast }}{{{\mathrm{\Delta }}G_0^ \ast }}\). Eq. (4) is approximated by \(k(F) = k_0^{\mathrm{A}}e^{\frac{{\gamma (F)\delta _0^ \ast }}{{k_{\mathrm{B}}T}}} = k_0^{\mathrm{A}}e^{ - \frac{{{\mathrm{\Delta \Phi }}^{\mathrm{F}}(\delta _0^ \ast )}}{{k_{\mathrm{B}}T}}}\), which is in the form of the Arrhenius equation.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its supplementary information files.
References
Marshall, B. T. et al. Direct observation of catch bonds involving cell-adhesion molecules. Nature 423, 190–193 (2003).
Yuan, G. et al. Elasticity of the transition state leading to an unexpected mechanical stabilization of titin immunoglob ulin domains. Angew. Chem. Int. Ed. 129, 5582–5585 (2017).
Jagannathan, B., Elms, P. J., Bustamante, C. & Marqusee, S. Direct observation of a force induced switch in the anisotropic mechanical unfolding pathway of a protein. Proc. Natl Acad. Sci. USA 109, 17820–17825 (2012).
Rakshit, S., Zhang, Y., Manibog, K., Shafraz, O. & Sivasankar, S. Ideal, catch, and slip bonds in cadherin adhesion. Proc. Natl Acad. Sci. USA 109, 18815–18820 (2012).
Chen, H. et al. Dynamics of equilibrium folding and unfolding transitions of titin immunoglobulin domain under constant forces. J. Am. Chem. Soc. 137, 3540–3546 (2015).
Bell, G. I. Models for the specific adhesion of cells to cells. Science 200, 618–627 (1978).
Evans, E., Leung, A., Heinrich, V. & Zhu, C. Mechanical switching and coupling between two dissociation pathways in a p-selectin adhesion bond. Proc. Natl Acad. Sci. USA 101, 11281–11286 (2004).
Hummer, G. & Szabo, A. Kinetics from nonequilibrium single-molecule pulling experiments. Biophys. J. 85, 5–15 (2003).
Dudko, O. K., Hummer, G. & Szabo, A. Intrinsic rates and activation free energies from single-molecule pulling experiments. Phys. Rev. Lett. 96, 108101 (2006).
Zhuravlev, P. I. et al. Force-dependent switch in protein unfolding pathways and transition-state move-ments. Proc. Natl Acad. Sci. USA 113, E715–E724 (2016).
Rebane, A. A., Ma, L. & Zhang, Y. Structure-based derivation of protein folding intermediates and energies from optical tweezers. Biophys. J. 110, 441–454 (2016).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Roca-Cusachs, P., Conte, V. & Trepat, X. Quantifying forces in cell biology. Nat. Cell Biol. 19, 742 (2017).
Huse, M. Mechanical forces in the immune system. Nat. Rev. Immunol. 17, 679 (2017).
Barsegov, V. & Thirumalai, D. Dynamics of unbinding of cell adhesion molecules: transition from catch to slip bonds. Proc. Natl Acad. Sci. USA 102, 1835–1839 (2005).
Pierse, C. A. & Dudko, O. K. Distinguishing signatures of multipathway confor mational transitions. Phys. Rev. Lett. 118, 088101 (2017).
Pereverzev, Y. V., Prezhdo, O. V., Forero, M., Sokurenko, E. V. & Wendy, E. The two-pathway model for the catch-slip transition in biological adhesion. Biophys. J. 89, 1446–1454 (2005).
Bartolo, D., Derényi, I. & Ajdari, A. Dynamic response of adhesion complexes: beyond the single-path picture. Phys. Rev. E 65, 051910 (2002).
Guo, S. et al. Structural-elastic determination of the force-dependent transition rate of biomolecules. Chem. Sci. 9, 5871–5882 (2018).
Kramers, H. A. Brownian motion in a field of force and the diffusion model of chemical reactions. Physica 7, 284–304 (1940).
Cocco, S., Yan, J., Léger, J.-F., Chatenay, D. & Marko, J. F. Over stretching and force-driven strand separation of double-helix dna. Phys. Rev. E 70, 011910 (2004).
Rouzina, I. & Bloomfield, V. A. Force-induced melting of the DNA double helix 1. Thermodynamic analysis. Biophys. J. 80, 882–893 (2001).
Lu, H., Isralewitz, B., Krammer, A., Vogel, V. & Schulten, K. Unfolding of titin immunoglob ulin domains by steered molecular dynamics simulation. Biophys. J. 75, 662–671 (1998).
Lu, H. & Schulten, K. Steered molecular dynamics simulation of conformational changes of immunoglobulin domain i27 interprete atomic force microscopy observations. Chem. Phys. 247, 141–153 (1999).
Best, R. B. et al. Mechanical unfolding of a titin Ig domain: structure of transition state revealed by combining atomic force microscopy, protein engineering and molecular dynamics simulations. J. Mol. Biol. 330, 867–877 (2003).
Marko, J. F. & Siggia, E. D. Stretching DNA. Macromolecules 28, 8759–8770 (1995).
Marszalek, P. E. et al. Mechanical unfolding intermediates in titin modules. Nature 402, 100 (1999).
Williams, M. et al. Hidden complexity in the mechanical properties of titin. Nature 422, 446 (2003).
Winardhi, R. S., Tang, Q., Chen, J., Yao, M. & Yan, J. Probing small molecule binding to unfolded polyprotein based on its elasticity and refolding. Biophys. J. 111, 2349–2357 (2016).
Schlierf, M., Berkemeier, F. & Rief, M. Direct observation of active protein folding using lock-in force spectroscopy. Biophys. J. 93, 3989–3998 (2007).
Rief, M., Pascual, J., Saraste, M. & Gaub, H. E. Single molecule force spectroscopy of spectrin repeats: low unfolding forces in helix bundles1. J. Mol. Biol. 286, 553–561 (1999).
Acknowledgements
This work is supported by the Singapore Ministry of Education Academic Research Fund Tier 3 (MOE2016-T3-1-002); the National Research Foundation (NRF), Prime Minister’s Office, Singapore under its NRF Investigatorship Programme (NRF Investigatorship Award No. NRF-NRFI2016-03); and the National Research Foundation, Prime Minister’s Office, Singapore and the Ministry of Education under the Research Centres of Excellence programme.
Author information
Authors and Affiliations
Contributions
J.Y. developed the main theory. S.G. predicted the DNA separation rate under unzipping and shearing force geometries. S.G. and A.K.E. analysed the experimental data of titin I27. J.Y. and S.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, S., Efremov, A.K. & Yan, J. Understanding the catch-bond kinetics of biomolecules on a one-dimensional energy landscape. Commun Chem 2, 30 (2019). https://doi.org/10.1038/s42004-019-0131-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-019-0131-6
This article is cited by
-
Catch bond models may explain how force amplifies TCR signaling and antigen discrimination
Nature Communications (2023)
-
Modulating mechanical stability of heterodimerization between engineered orthogonal helical domains
Nature Communications (2020)
-
A mechano-chemo-biological model for bone remodeling with a new mechano-chemo-transduction approach
Biomechanics and Modeling in Mechanobiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.