Abstract
Energy migration between the identical chromophores is a necessary process in both natural and artificial photosynthesis. The distance and orientation dependence of energy migration have not been experimentally investigated in detail. Here we propose a method to investigate energy migration. Two fluorophores are introduced into one strand of a DNA duplex with a quencher placed opposite one of fluorophores. This design enables asymmetrization of identical fluorophores and allows one fluorophore to behave as an acceptor. The emission intensities and lifetimes decrease depending on the efficiency of energy migration. Distance and orientation dependence are successfully quantified, and the excitation energy migration efficiencies measured are in excellent agreement with those calculated based on Förster theory. We also demonstrate that multi-step energy migration among four fluorophores can be estimated from the theory. These results may provide a basis for design and preparation of efficient light-harvesting photonic devices and chemical probes.
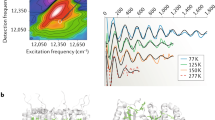
Similar content being viewed by others
Introduction
Excitation energy migration (EM), also called excitation energy hopping or homo Förster resonance energy transfer (homo-FRET), is energy transfer among identical chromophores. This type of transfer is a key process in natural photosynthesis since excitation energy can be transported without losing photon energy. In natural light-harvesting complexes, absorbed energy migrates first among a number of identical chlorophyll molecules and subsequently transfers to reaction centre. Natural light-harvesting complexes achieve efficient solar energy conversion by tuning energy migration efficiency1. Energy migration is also important in artificial light-harvesting systems, chemical sensing, photon energy conversion, and analyses of biomolecular probes2,3,4. Although it has been shown that EM efficiencies can be controlled to an extent through positioning of covalent or non-covalent bonds, precise control of distance and orientation of chromophores over a wide range still remains challenging.
DNA spontaneously forms a well-defined right-handed double helix, in which four nucleobases are aligned according to the sequence. Arrays of non-natural molecules with pre-determined size, distance, and even sequence can be easily prepared by incorporating unnatural monomer into DNA through covalent bonding5,6. DNA has been widely used as a platform to prepare chromophoric arrays7,8,9,10,11, and recently, photonic arrays and circuits based on DNA nano-structures have been reported12,13,14,15. In these structures, EM and hetero FRET play crucial roles in efficient energy harvesting and transport16,17,18,19,20,21,22,23,24,25,26. Several research groups, including ours, have utilized DNA scaffolds to quantify orientation and distance dependence of hetero FRET from a donor to an acceptor27,28,29,30,31,32,33. Distance and orientation dependences of energy migration between identical chromophores have not been elucidated. In homo-FRET, energy migration occurs among the identical chromophores, and each chromophore can function both as a donor and an acceptor. Such symmetry among the chromophores makes it impossible to separate donor and acceptor emission. Time-resolved anisotropy measurement has been used to analyse EM; however, the analyses are model-dependent, and kinetic parameters cannot be determined directly34.
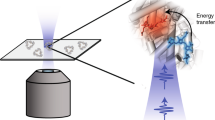
Herein, we describe a system for analysis of EM using a DNA scaffold in which the symmetry between the identical fluorophores is broken. Our design is depicted in Fig. 1a. Two identical fluorophores are introduced into the DNA strands. The distance between the two is controlled by changing the number of intervening base pairs. A quencher is located opposite one of two fluorophores in the DNA duplex, which makes that particular fluorophore non-emissive so that the quenched fluorophore functions not as a donor but as an acceptor. Since absorbed energy on this fluorophore is no longer transferred, EM efficiency can be quantified by measuring the decrease of emission intensity and lifetime as hetero FRET. In this paper, we elucidated distance and orientation dependence of energy migration by using this system. Multi-step energy migration among four fluorophores is also analysed. To our knowledge, this is the first study to experimentally reveal the distance and orientation dependence of energy migration among identical dyes.
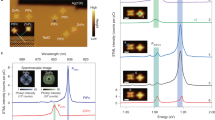
Design of duplex systems used in this study. a Schematic illustration of EQ-E duplex and chemical structures of E and Q residues. b Absorption and emission spectra of perylene and anthraquinone. c Illustration of EQ-E, E, EQ and Q-E duplexes. These duplexes can be prepared by changing the combination of the three strands
Results
Sequence design
Perylene and anthraquinone (Fig. 1a) were selected as the fluorophore and the quencher, respectively, for these experiments for three reasons. First, anthraquinone quenches perylene emission very efficiently as we reported previously35. Therefore, back energy transfer from the quenched fluorophore to the emissive fluorophore can be neglected. Second, anthraquinone absorption has almost no overlap with perylene emission (Fig. 1b) so that hetero energy transfer from perylene to anthraquinone, which disturbs the quantitative analyses of energy migration, does not occur. Third, these molecules have planar structures, and therefore stack with natural base pairs. A D-threoninol linker was used to introduce these chromophore as it facilitates intercalation of these molecules into DNA duplex36.
The DNA duplex used to investigate orientation and distance dependence of EM is schematically illustrated in Fig. 1c, and the actual sequences are listed in Supplementary Table 1. We introduced one perylene residue into a 32-mer DNA (strands E1-E4). One perylene or one anthraquinone residue was introduced into strands complementary to the 32-mer (strands cE1s-cE3s or cQE1-cQE4, respectively). By hybridizing three strands, nicked duplexes with two perylenes, one of which is located in opposite anthraquinone, were prepared (duplex EQ-E). The number of base pairs between perylene and anthraquinone were varied from 2 to 13 (see Supplementary Table 1 for details). Control duplexes with one perylene residue or one perylene-anthraquinone pair (duplexes E and EQ, respectively) were prepared by using native strands (NE, cNE or cNEs).
We previously showed that incorporation of planar molecules via D-threoninol does not distort the double helical structure36,37 and that dyes are stacked with a base pair at 5′ side when they are introduced into the base-pairing position37. Accordingly, in our design, anthraquinone is located between two perylenes in the EQ-E duplex, and the distance between anthraquinone and perylene is the same in EQ-E and Q-E duplexes.
Quantitative analysis of energy migration between two perylenes
We first measured melting temperatures (Tms) of duplexes and confirmed that duplexes were stable below 20 °C (Supplementary Table 2). Incorporation of perylene and/or anthraquinone residues resulted in duplexes more stable than the appropriate control duplexes, indicating that there are stacking interactions of perylene and anthraquinone with neighbouring natural base pairs. Emission spectra of Q-E duplexes were measured to assess anthraquinone to perylene emission. We measured steady-state emission spectra of Q-E duplexes that contain one perylene and one anthraquinone separated by 2 to 9 base pairs (Fig. 2a and Supplementary Fig. 1). Emission intensity of the Q-E duplex with a 2-base-pair separation was much lower than that of the control E duplex, whereas quenching was not detectable when residues were separated by 3 or more base pairs. Anthraquinone quenches the perylene fluorescence through electron transfer38. Thus, separation of E and Q by more than two base pairs does not reduce perylene emission. These results led us to conclude that quenching by anthraquinone can be neglected when the distance between perylene and anthraquinone is more than 2 base pairs.
Analyses of energy migration through steady-state or time-resolved fluorescence measurements. a Effects of quenching of perylene emission by an anthraquinone located 2 to 5 base pairs away. b Fluorescence emission spectra of EQ-E, E and EQ duplexes with 5-base-pair separation. c Fluorescence lifetime measurements of EQ-E and E duplexes with 5-base-pair separation. d Emission intensities and lifetimes of EQ-E duplexes
Next, we compared emission spectra of E, EQ and EQ-E duplexes. In the EQ-E duplexes the perylene and the anthraquinone are separated by from 2 to 13 base pairs. The E duplex had a strong emission, whereas virtually no emission was observed with the EQ duplex, demonstrating the efficient quenching of perylene by anthraquinone. The duplexes in the EQ-E series had much lower emission than the E duplex. A representative result for the duplex with a 5-base-pair separation between fluorophores is shown in Fig. 2b. This quenching clearly demonstrated the occurrence of energy migration between two perylenes. Similar quenching was observed with all the other EQ-E duplexes (Supplementary Figs. 2 and 3), and their emission intensities are summarized in Fig. 2d. Generally, emission intensity increased as the number of base pairs increased; however, maxima of emission intensity were observed when 8 and 13 base pairs separated the fluorophores. Similar a 5-base-pair cycle of emission intensity was observed with hetero FRET from pyrene to perylene that corresponds to a half turn of the B-form DNA duplex32,39. Our result indicated that energy migration between identical chromophores also exhibits orientation dependence.
We also performed time-resolved fluorescence measurements and representative results of the duplex with 5-base-pair spacing are shown in Fig. 2c (see Supplementary Fig. 4 for decay curves of other duplexes). The EQ-E duplex showed much faster decay than the E duplex, indicative of quenching due to energy migration. We also monitored decay of EQ duplexes; however, lifetimes could not be determined as decay was too rapid. Lifetimes of EQ-E duplexes are listed in Table 1 (lifetimes of control duplexes are listed in Supplementary Table 3). Although two lifetimes were observed when fluorophores were 3 to 6 base pairs apart, the longer lifetimes are attributed to emission from perylene in residual single strands.
We used the shorter lifetimes for calculation of energy migration efficiencies. We calculated EM efficiencies from the decrease of emission intensities and lifetimes (ΦT1 and ΦT2 in Table 1, respectively). ΦT1 and ΦT2 generally decreased as the distance between two perylenes increased. The lifetimes exhibited similar trends as emission intensities with the longest lifetimes observed at intervals of about 5 base pairs, also demonstrating the orientation dependence of energy migration. The difference between ΦT1 and ΦT2 at short distances is relatively large; ΦT2 values are above 0.95 with 3 to 5 base pairs between fluorophores, whereas ΦT1 values are about 0.7. We hypothesized that these differences are due to the emission from excess perylene strands. Although perylene strands (Em and cEns in Fig. 1c) of the same concentration were used for measurements, emission from perylene in the excess single strands lowered the apparent ΦT1. In contrast, lifetime measurements can discriminate emission of excess single strands from that of EQ-E duplex.
Comparison with Förster theory
These experimentally determined efficiencies were compared with values calculated from Förster theory. Distance, orientation factor, and spectral overlap were determined and used for the calculation of migration efficiency. We first investigated the effect of anthraquinone on perylene absorption (Supplementary Fig. 5). Slight hypsochromism was observed, however, its effect on spectral overlap was <1%. Therefore, we concluded that optical property of quenched perylene is virtually identical to that of the unquenched one. Molecular modelling of EQ-E duplexes indicated that perylene and anthraquinone are stacked between base pairs without disturbing double helical structure (Fig. 3a). The angle between perylene and the neighbouring base pairs were estimated from curve fitting and molecular modelling (Fig. 3b). We used a cylinder model where the distance increased by 3.3 Å per base pair and the angle by 33° per base pair (Fig. 3c). The increment of distance due to anthraquinone was assumed to be the same as for a natural base pair (3.3 Å). The theoretically calculated EM efficiencies are shown in Fig. 4. The theoretical curve showed the same tendency as experimental values. Especially, ΦT2 values showed excellent agreement. We also measured ΦT1 at various temperatures (Supplementary Fig. 6), and values were almost the same irrespective of temperature. The insensitivity to temperature indicated that effects of dynamics and disorder are marginal under the conditions employed. These results clearly demonstrated that the energy migration between two perylenes strictly obeys Förster theory. Thus, these results provided the first experimental validation of Förster theory for the calculation of efficiencies and rate constants of energy migration.
Molecular modelling and cylinder model of EQ-E duplexes. a Molecular modelling of EQ-E duplexes with 13- and 2-base-pair separations. Perylene and anthraquinone moieties are shown in CPK model and coloured in green and red, respectively. b Relative orientations of two perylene moieties in EQ-E duplexes. Angles between perylene moieties and neighbouring base pairs are 97 ° and 95 ° for 13-base-pair and 2-base-pair separations, respectively. c Cylinder model of DNA duplex for theoretical calculation of EM efficiencies. Distance and angle increases by 3.3 Å and 33° per base pair in this model, respectively
Dependence of EM efficiency on the number of base pairs between two perylene moieties. Experimentally obtained efficiencies from emission intensities and lifetimes are shown in blue and red circles, respectively. Theoretically calculated values assuming random or fixed orientations are shown in black and red lines, respectively. Error bars of ΦT1 show standard deviation of three independent fluorescence measurements
Analysis of energy migration among four perylenes
We then used a similar system to investigate multi-step energy migration among four perylenes. A schematic of the experimental design is illustrated in Fig. 5a (see Supplementary Table 4 for DNA sequences). Four perylenes were introduced into DNA at 3-nucleotide intervals in strand 4E. The complementary strand, cQ, has one anthraquinone opposite the terminal perylene of 4E. When 4E is hybridized with cQ, emission from the three perylenes located distant from the anthraquinone should be quenched through multi-step energy migration. Therefore, energy migration among four perylenes can be monitored by analysis of emission intensity and lifetime. We also synthesized DNA strands tethering perylene residues at different positions (1E-1 to 1E-4) as controls. The controls showed similar emission intensities when each was hybridized with a complementary strand without a quencher (cN) as shown in Supplementary Fig. 7. 4E/cN showed almost identical intensity to the sum of intensities of 1E-1/cN, 1E-2/cN, 1E-3/cN and 1E-4/cN. The emission intensity of 1E-1/cQ was almost nil, whereas other combinations were the same as 1E duplex without Q (Fig. 5b). The 4E/cQ duplex with four fluorophores had much lower emission than the sum of the emission from each perylene (Fig. 5b, compare red line with dotted line). This large difference between 4E/cQ and the sum is clear evidence of multi-step energy migration among four perylenes. The higher Tm of 4E/cQ than those of control duplexes (1E-1/cQ to 1E-4/cQ) indicated that base pairing of natural base pairs is not severely disturbed upon the introduction of non-natural residues (Supplementary Table 5). We also performed time-resolved fluorescence measurements, and decay curves of 4E/cQ and 4E/cN are shown in Fig. 5c. 4E/cN showed a monoexponential decay with a lifetime of 8.40 ns (Table 2). In contrast, biexponential decay with lifetimes of 1.19 ns and 3.80 ns was observed for 4E/cQ. The decrease in lifetime associated with the incorporation of anthraquinone clearly demonstrates the efficient energy migration among four perylenes.
Multi-step energy migration among four perylene moieties. a Sequence design to investigate multi-step energy migration. b Fluorescence emission spectra of 4E/cQ, 1E-1/cQ, 1E-2/cQ, 1E-3/cQ and 1E-4/cQ. Sum of spectra of 1E-1/cQ, 1E-2/cQ, 1E-3/cQ and 1E-4/cQ is also shown. c Fluorescence lifetimes of 4E/cQ and 4E/cN duplexes. d Simulated decay curve of 4E/cQ. Time evolution of excited population of each perylene is also shown. Numbering of perylenes is schematically illustrated in the inset. e Molecular modelling of 4E/cQ duplex. Perylene and anthraquinone moieties are shown in CPK model and are coloured in green and red, respectively
We simulated the fluorescence decay of 4E/cQ from the non-coherent energy hopping mechanism based on Förster theory23,25,40. The decay curve of each perylene can be simulated by solving differential equations using migration and decay rate constants. Rate constants of intrinsic decays of each perylene were determined from lifetimes; the exception was the decay rate of the perylene adjacent to anthraquinone, which was estimated from emission intensity. Energy migration rate constants were calculated using decay rates and migration efficiencies (Supplementary Table 6). Migration efficiencies were calculated from Förster theory by using the distances and orientation factors between perylenes, which were determined from molecular modelling (Fig. 5e). Simulated decay curves of the four perylenes are shown in Fig. 5d. The resultant decay curve of each perylene depended on its position. The perylene located far from anthraquinone had the slowest decay, whereas the perylene located to the 5′ side of anthraquinone had the fastest (Fig. 5d). Interestingly, the average curve of four perylenes, which corresponds to the overall decay curve of 4E/cQ, could be fitted with a biexponential curve with lifetimes of 0.05 and 1.42 ns (Table 2 and Supplementary Fig. 8). A 0.05-ns lifetime is too short to be detected by our apparatus. In contrast, observed lifetime of 1.19 ns is in good agreement with the simulated lifetime (1.42 ns). The longer lifetime (3.80 ns) might be due to excess single strand or to structural disorder; either would result in incomplete migration. These results indicated that even efficiency and rate constants of multi-step energy migration can be estimated using Förster theory.
Discussion
Here we clarified the distance and orientation dependence of energy migration in detail. We demonstrated that energy migration between identical chromophores strictly obeys Förster’s theory. Our system can be used to determine and predict efficiencies and rate constants of energy migration steps of various chromophoric arrays. Recent studies have revealed that coherent coupling plays important roles in efficient energy transport in natural photosynthesis15,41. Since our system is simple but robust, it could be used to analyse other fluorophores such as Cy3 or thiazole orange42. Furthermore, it could be used to investigate energy migration processes in strongly coupling regimes. We also demonstrated that multi-step migration can also be estimated from the theory. Energy migration is used in many applications such as light-harvesting antennae, logic gates, chemical sensing, and photon energy conversion43. In order to achieve the best performance of these devices, it is necessary to tune rate constants of each energy transport step precisely. The present results may provide a framework for design and preparation of photonic devices, circuits, and sensors with desirable properties.
Methods
Oligonucleotides
All conventional phosphoramidite monomers, CPG columns, reagents for DNA synthesis, and Poly-Pak II cartridges were purchased from Glen Research. Other reagents for the synthesis of phosphoramidite monomers were purchased from Tokyo Chemical Industry, Wako, and Aldrich. Native oligodeoxyribonucleotides (ODNs) were purchased from Integrated DNA Technologies. ODNs tethering perylene or anthraquinone were synthesized on an automated DNA synthesizer (H-8-SE, Gene World) as we reported previously44,45. ODNs were purified by reversed-phase HPLC and characterized by MALDI-TOF MS (Autoflex II, Bruker Daltonics) and HPLC.
The MALDI-TOF MS data for the modified DNA were as follows: E1: Obsd. 10257 (Calcd. for [E1+H+]: 10257). E2: Obsd. 10257 (Calcd. for [E2+H+]: 10257). E3: Obsd. 10257 (Calcd. for [E3+H+]: 10257). E4: Obsd. 10256 (Calcd. for [E4+H+]: 10257). cQE1: Obsd. 3703 (Calcd. for [cQE1+H+]: 3702). cQE2: Obsd. 3703 (Calcd. for [cQE2+H+]: 3702). cQE3: Obsd. 3703 (Calcd. for [cQE3+H+]: 3702). cQE4: Obsd. 3703 (Calcd. for [cQE4+H+]: 3702). cE1s: Obsd. 6905 (Calcd. for [cE1s+H+]: 6908). cE2s: Obsd. 6908 (Calcd. for [cE2s+H+]: 6908). cE3s: Obsd. 6908 (Calcd. for [cE3s+H+]: 6908). 4E: Obsd. 12227 (Calcd. for [4E+H+]: 12233). 1E-1: Obsd. 10897 (Calcd. for [1E-1+H+]: 10898). 1E-2: Obsd. 10895 (Calcd. for [1E-2+H+]: 10898). 1E-3: Obsd. 10892 (Calcd. for [1E-3 +H+]: 10898). 1E-4: Obsd. 10896 (Calcd. for [1E-4+H+]: 10898). cQ: Obsd. 10822 (Calcd. for [cQ+H+]: 10823).
Absorption measurements
Absorption spectra were measured on a JASCO model V-530, V-550 or V-560. Sample solution contained 5 μM each strand, 100 mM NaCl, 10 mM phosphate buffer (pH 7.0). The melting curves were measured with a UV-1800 (Shimadzu) by monitoring 260 nm absorbance versus temperature. The melting temperature (Tm) was determined from the maximum in the first derivative of the melting curve. Both the heating and the cooling curves were measured, and the calculated Tms agreed to within 1.0 °C. The temperature ramp was 0.5 °C min−1. The sample solutions for melting analyses from absorbance contained 100 mM NaCl, 10 mM phosphate buffer, pH 7, 1.0 µM (for data shown in Supplementary Table 2) or 2.0 μM (for data shown in Supplementary Table 5) each strand.
Fluorescence measurements
Fluorescence spectra were measured on JASCO models FP-6500 and FP-8500. The excitation wavelength was 427 nm. Band widths were 3 nm (FP-6500) or 2.5 nm (FP-8500) for excitation and emission. Before measurements, sample solutions containing DNA duplex were heated at 80 °C, then slowly cooled down to 0 °C at a rate of 4 °C min−1. Fluorescence spectra were measured over the range from 80 to 0 °C at 10 °C intervals with 4-min incubations after each temperature change. Emission spectra shown in this paper were measured at 0 °C unless otherwise noted. Sample solutions contained 100 mM NaCl, 10 mM phosphate buffer, pH 7.0. Concentrations of oligonucleotides were 1.0 µM for those tethering perylene (E1 to E4, cE1s to cE3s, 4E, 1E-1 to 1E-4) and 1.5 µM quencher strands (cQE1 to cQE4, cQ). For control measurements, concentrations of native strands were 1.0 µM for NE and cNEs and 1.5 µM for cNE and cN.
Fluorescence lifetime measurements
A pulse at 780 nm was generated by a Ti:sapphire laser system (Spectra-Physics, Tsunami; 3950-L2S, fwhm 150 fs, 82 MHz). The repetition rate was reduced to 4 MHz by a pulse selector (Spectra-Physics Model 3980). The exciting source was a laser with wavelength converted to 390 nm by passage through SHG crystals. Fluorescence emission was captured by a streak camera (Hamamatsu C4334) operating in photon counting mode. Measurements were performed at room temperature.
Decay curves were obtained from the integration of the photon counts in the spectral region from 470 to 530 nm in the streak image. The fluorescence decay curve was analysed using the U8167-01 programme (Hamamatsu). The decay curves were fitted with the biexponential function α1exp(−t/τ1) + α2exp(−t/τ2).
Determination of EM efficiency
Energy transfer efficiency was calculated from steady-state fluorescence (ΦT1) or fluorescent lifetimes (ΦT2). ΦT1 was calculated from the following equation:
where I, IQ and ID are emission intensities of EQ-E duplex, EQ duplex and E duplex, respectively. When energy transfer between two perylenes does not occur, I should be equal to ID plus IQ so that ΦT1 becomes zero. In contrast, when excitation energy of perylene at distant position perfectly transfers to perylene next to anthraquinone, the intensity of the perylene should be the same as that of the quenched one. Therefore, I should be equal to 2IQ. In this case, ΦT1 becomes unity (Supplementary Figure 9).
The energy migration efficiency, ΦT2, was determined from fluorescence lifetime measurements. For duplexes containing one fluorophore with no quencher (E duplex), the fluorescent lifetime (τD) can be represented as follows:
where kf and kd1 are an emissive rate constant and a non-radiative decay rate, respectively (Supplementary Figure 10). τ, which is a lifetime of EQ-E duplex, can be represented by using the energy migration rate constant (kt):
Lifetimes of EQ duplexes could not be determined in time-resolved fluorescence measurements with our apparatus. Besides, emission intensity of EQ duplexes were much lower than those of E duplexes, indicating quenching by anthraquinone occurs very rapidly. Therefore, we ignored the decay rate of quenched perylene (kd2) because it is much larger than other rate constants. By using equations (2) and (3), energy transfer efficiency from fluorescent lifetimes (ΦT1) can be simply calculated as follows:
Energy minimization based on molecular mechanics
The energy minimization for the molecular structure model of DNA duplexes tethering perylene and anthraquinone was carried out using MacroModel (MacroModel, version 11.6; Schrödinger) applying the AMBER force field. Duplexes were constructed from canonical B-form DNA duplex by using a graphical programme. The water solvent effects were simulated using the analytical Generalized-Born/Surface-Area (GB/SA) model. Convergence threshold was set to 0.05 kJ Å−1 mol−1.
Theoretical calculation of energy transfer efficiencies based on Förster theory
Energy transfer efficiency was calculated from the following equations:
where R is the distance between donor and acceptor, R0 is a Förster radius (the distance where ΦT equals 0.5), J is the integral of spectral overlap between absorption and emission of perylene, n is a refractive index (which is typically assumed to be 1.4 for biomolecules), and ΦD is a quantum yield of perylene. When dyes are located in parallel planes, the orientation factor, κ2, can be calculated by using the angle between transition dipoles of dyes (θT):
Theoretical transfer efficiency can be calculated from the distance and angle between two chromophores. Rise and typical twist angle of B-form duplex, 3.3 Å per base pair and 33° per base pair, respectively, were used to calculate the distance and the angle between dyes. These parameters were determined by fitting theoretically calculated efficiencies with experimental data, and the resultant values are consistent with reported parameters46,47. In our design, anthraquinone is located between two perylenes. Therefore, we hypothesized that the distance between perylenes and anthraquinone was also 3.3 Å. We assumed an angle of 52° between dyes angles based on an estimate from molecular modelling and curve fitting. When anthraquinone was introduced into the counter position of perylene, slight hypochromic effect was observed in UV-vis spectrum (Supplementary Fig. 5), which were used to calculate spectral overlap integral.
Simulation of multi-step energy migration
Energy migration among four perylenes was simulated by solving the following differential equations.
where Pn is probability of nth perylene being excited, kmn is rate constant of energy transfer from mth perylene to nth perylene (see Fig. 5d for the number of perylenes), and kdn is rate constant of intrinsic decay of nth perylene. knms were calculated from equations (5) to (7). Distances and orientation factors were calculated based on molecular modelling shown in Fig. 5e. For simplicity, it was assumed that constants of forward transfers were equal to those of reverse transfer (i.e., kmn = knm). kd2, kd3 and kd4 were calculated from fluorescence lifetimes of 1E-2/cQ, 1E-3/cQ and 1E-4/cQ, respectively. Since kd1 cannot be determined directly due to fast decay of 1E-1/cQ, we estimated kd1 from its fluorescence intensity (Fig. 5b, green line). Since intensity of 1E-1/cQ is about one thousand times lower than that of 1E-1/cN, we used 1.00 × 1011 s−1 as kd1. We also confirmed that small difference of kd1 does not significantly affect the results of the simulation. Rate constants used for the simulation are listed in Supplementary Table 6. Differential equations were solved by using lsode function in GNU Octave 4.2.1. The simulated fluorescence lifetime of 4E/cQ was calculated by fitting the biexponential curve to the averaged decay curve (Supplementary Fig. 8).
Data availability
All relevant data supporting the findings of this study are available within the article and its Supporting information files, and from the corresponding author upon reasonable request.
References
McDermott, G. et al. Crystal structure of an integral membrane light-harvesting complex from photosynthetic bacteria. Nature 374, 517–521 (1995).
Wasielewski, M. R. Self-assembly strategies for integrating light harvesting and charge separation in artificial photosynthetic systems. Acc. Chem. Res. 42, 1910–1921 (2009).
Aratani, N., Kim, D. & Osuka, A. Discrete cyclic porphyrin arrays as artificial light-harvesting antenna. Acc. Chem. Res. 42, 1922–1934 (2009).
Bader, A. N. et al. Homo-FRET imaging as a tool to quantify protein and lipid clustering. ChemPhysChem 12, 475–483 (2011).
Jones, M. R., Seeman, N. C. & Mirkin, C. A. Programmable materials and the nature of the DNA bond. Science 347, 1260901 (2015).
Komiyama, M., Yoshimoto, K., Sisido, M. & Ariga, K. Chemistry can make strict and fuzzy controls for bio-systems: DNA nanoarchitectonics and cell-macromolecular nanoarchitectonics. Bull. Chem. Soc. Jpn. 90, 967–1004 (2017).
Malinovskii, V. L., Wenger, D. & Häner, R. Nucleic acid-guided assembly of aromatic chromophores. Chem. Soc. Rev. 39, 410–422 (2010).
Teo, Y. N. & Kool, E. T. DNA-multichromophore systems. Chem. Rev. 112, 4221–4245 (2012).
Kashida, H. & Asanuma, H. Preparation of supramolecular chromophoric assemblies using a DNA duplex. Phys. Chem. Chem. Phys. 14, 7196–7204 (2012).
Ensslen, P. & Wagenknecht, H.-A. One-dimensional multichromophor arrays based on DNA: from self-assembly to light-harvesting. Acc. Chem. Res. 48, 2724–2733 (2015).
Stulz, E. Nanoarchitectonics with porphyrin functionalized DNA. Acc. Chem. Res. 50, 823–831 (2017).
Stein, I. H., Steinhauer, C. & Tinnefeld, P. Single-molecule four-color FRET visualizes energy-transfer paths on DNA origami. J. Am. Chem. Soc. 133, 4193–4195 (2011).
Graugnard, E. et al. DNA-controlled excitonic switches. Nano Lett. 12, 2117–2122 (2012).
Buckhout-White, S. et al. Assembling programmable FRET-based photonic networks using designer DNA scaffolds. Nat. Commun. 5, 5615 (2014).
Boulais, É. et al. Programmed coherent coupling in a synthetic DNA-based excitonic circuit. Nat. Mater. 17, 159–166 (2017).
Ohya, Y., Yabuki, K., Hashimoto, M., Nakajima, A. & Ouchi, T. Multistep fluorescence resonance energy transfer in sequential chromophore array constructed on oligo-DNA assemblies. Bioconjugate Chem. 14, 1057–1066 (2003).
Vyawahare, S., Eyal, S., Mathews, K. D. & Quake, S. R. Nanometer-scale fluorescence resonance optical waveguides. Nano Lett. 4, 1035–1039 (2004).
Benvin, A. L. et al. Fluorescent DNA nanotags: supramolecular fluorescent labels based on intercalating dye arrays assembled on nanostructured DNA templates. J. Am. Chem. Soc. 129, 2025–2034 (2007).
Hannestad, J. K., Sandin, P. & Albinsson, B. Self-assembled DNA photonic wire for long-range energy transfer. J. Am. Chem. Soc. 130, 15889–15895 (2008).
Kumar, C. V. & Duff, M. R. DNA-based supramolecular artificial light harvesting complexes. J. Am. Chem. Soc. 131, 16024–16026 (2009).
Dutta, P. K. et al. DNA-directed artificial light-harvesting antenna. J. Am. Chem. Soc. 133, 11985–11993 (2011).
Garo, F. & Häner, R. A DNA‐based light‐harvesting antenna. Angew. Chem. Int. Ed. 51, 916–919 (2012).
Pan, K., Boulais, E., Yang, L. & Bathe, M. Structure-based model for light-harvesting properties of nucleic acid nanostructures. Nucleic Acids Res 42, 2159–2170 (2014).
Hemmig, E. A. et al. Programming light-harvesting efficiency using DNA origami. Nano Lett. 16, 2369–2374 (2016).
Melinger, J. S. et al. FRET from multiple pathways in fluorophore-labeled DNA. ACS Photonics 3, 659–669 (2016).
Nicoli, F. et al. Directional photonic wire mediated by homo-Förster resonance energy transfer on a DNA origami platform. ACS Nano 11, 11264–11272 (2017).
Lewis, F. D., Zhang, L. & Zuo, X. Orientation control of fluorescence resonance energy transfer using DNA as a helical scaffold. J. Am. Chem. Soc. 127, 10002–10003 (2005).
Iqbal, A. et al. Orientation dependence in fluorescent energy transfer between Cy3 and Cy5 terminally attached to double-stranded nucleic acids. Proc. Natl Acad. Sci. USA 105, 11176–11181 (2008).
Börjesson, K. et al. Nucleic acid base analog FRET-pair facilitating detailed structural measurements in nucleic acid containing systems. J. Am. Chem. Soc. 131, 4288–4293 (2009).
Preus, S., Kilså, K., Miannay, F.-A., Albinsson, B. & Wilhelmsson, L. M. FRETmatrix: a general methodology for the simulation and analysis of FRET in nucleic acids. Nucleic Acids Res. 41, e18 (2013).
Fessl, T., Lilley & David, M. J. Measurement of the change in twist at a helical junction in RNA using the orientation dependence of FRET. Biophys. J. 105, 2175–2181 (2013).
Kato, T. et al. Development of a robust model system of FRET using base surrogates tethering fluorophores for strict control of their position and orientation within DNA duplex. J. Am. Chem. Soc. 135, 741–750 (2013).
Wranne, M. S. et al. Toward complete sequence flexibility of nucleic acid base analogue FRET. J. Am. Chem. Soc. 139, 9271–9280 (2017).
Bradforth, S. E., Jimenez, R., van Mourik, F., van Grondelle, R. & Fleming, G. R. Excitation transfer in the core light-harvesting complex (LH-1) of Rhodobacter sphaeroides: an ultrafast fluorescence depolarization and annihilation study. J. Phys. Chem. 99, 16179–16191 (1995).
Kashida, H. et al. In-stem molecular beacon containing a pseudo base pair of threoninol nucleotides for the removal of background emission. Angew. Chem. Int. Ed. 48, 7044–7047 (2009).
Asanuma, H., Murayama, K., Kamiya, Y. & Kashida, H. Design of photofunctional oligonucleotides by copolymerization of natural nucleobases with base surrogates prepared from acyclic scaffolds. Polym. J. 49, 279–289 (2016).
Fujii, T., Kashida, H. & Asanuma, H. Analysis of coherent heteroclustering of different dyes by use of threoninol nucleotides for comparison with the molecular exciton theory. Chem. Eur. J. 15, 10092–10102 (2009).
Ishii, A. & Miyasaka, T. A high voltage organic–inorganic hybrid photovoltaic cell sensitized with metal–ligand interfacial complexes. Chem. Commun. 48, 9900–9902 (2012).
Kashida, H., Kurihara, A., Kawai, H. & Asanuma, H. Orientation-dependent FRET system reveals differences in structures and flexibilities of nicked and gapped DNA duplexes. Nucleic Acids Res. 45, e105 (2017).
Cho, H. S. et al. Excitation energy transport processes of porphyrin monomer, dimer, cyclic trimer, and hexamer probed by ultrafast fluorescence anisotropy decay. J. Am. Chem. Soc. 125, 5849–5860 (2003).
Chenu, A. & Scholes, G. D. Coherence in energy transfer and photosynthesis. Annu. Rev. Phys. Chem. 66, 69–96 (2015).
Kashida, H., Morimoto, K. & Asanuma, H. A stem-less probe using spontaneous pairing between Cy3 and quencher for RNA detection. Sci. Technol. Adv. Mater. 17, 267–273 (2016).
Peng, H.-Q. et al. Biological applications of supramolecular assemblies designed for excitation energy transfer. Chem. Rev. 115, 7502–7542 (2015).
Asanuma, H. et al. Quencher-free linear probe with multiple fluorophores on an acyclic scaffold. Chem. Sci. 3, 3165–3169 (2012).
Doi, T. et al. Hetero-selective DNA-like duplex stabilized by donor–acceptor interactions. Chem. Eur. J. 21, 15974–15980 (2015).
Dickerson, R. E. & Drew, H. R. Structure of a B-DNA dodecamer. J. Mol. Biol. 149, 761–786 (1981).
Dickerson, R. et al. The anatomy of A-, B-, and Z-DNA. Science 216, 475–485 (1982).
Acknowledgements
This work was supported by JSPS KAKENHI grant numbers JP18H03933 (H.A.), JP16H05925 (H.K.), JP17H05150 (H.K.) and JP16H04127 (Y.A.), and by PRESTO grant number JPMJPR14F7 (H.K.) and A-STEP (H.A.) from Japan Science and Technology Agency. Supports from the JSPS A3 Fore-sight Programme and the Asahi Glass Foundation (H.A.) are also acknowledged.
Author information
Authors and Affiliations
Contributions
H. Kashida and H.A. designed experiments. H. Kawai, R.M. and Y.K. performed spectroscopic analyses. Y.A. and T.W. performed fluorescence lifetime measurements. H. Kashida and H.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kashida, H., Kawai, H., Maruyama, R. et al. Quantitative evaluation of energy migration between identical chromophores enabled by breaking symmetry. Commun Chem 1, 91 (2018). https://doi.org/10.1038/s42004-018-0093-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0093-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.