Abstract
The formation of ordered amyloid fibrils by proteins and polypeptides is associated with human disorders. A recent extension of the amyloidogenic building block family includes several small metabolites, which form assemblies with structural and functional similarities to well-established amyloids. Here we investigate whether generic amyloid polyphenolic inhibitors can also restrict the formation of metabolite fibrils. We reveal that epigallocatechin gallate and tannic acid inhibit amyloid-like fibrillation of adenine, phenylalanine, and tyrosine. Moreover, the compounds reduce the cytotoxicity triggered by these assemblies. In contrast, acetylsalicylic acid, used as a control does not have an inhibitory effect. The compounds’ differential effects at various time points is consistent with molecular dynamics simulations, providing information about the inhibition mechanisms and inhibitors’ key interactions with the monomeric and subsequent crystalline fibril states. Taken together, we provide additional evidence for the fundamental similarities between protein- and metabolite-based amyloids, the inhibition process and dynamics of association.
Similar content being viewed by others
Introduction
Amyloid aggregates are well-ordered β-sheet-rich supramolecular structures, formed by the self-assembly of a wide class of proteins and polypeptides, which do not exhibit simple structural or functional similarities1,2,3,4,5. This group of proteins and polypeptides is associated with a family of amyloid diseases, related to intra- and extra-cellular protein-misfolding processes, which result in the formation of deposits and plaques and lead to cell death. There are currently > 30 known amyloid-associated disorders, including common neurodegenerative diseases, such as Parkinson’s disease, Alzheimer’s disease and prion disorders4,6,7,8,9,10. Amyloid assemblies possess a common set of biological, chemical, physical, and ultrastructural characteristics, including a twisted and elongated fibrillar morphology with a diameter of 5–20 nm3,11,12. The formed fibrils are known to interact with amyloid-specific dyes, such as thioflavin-T (ThT), owing to their β-sheet-rich secondary structure, resulting in a typical fluorescence signature that can be monitored in a time-dependent manner13,14.
Although several potential therapeutic approaches for ameliorating amyloidogenic diseases currently exist, one of the more promising approaches allows the inhibition of amyloid formation by small-molecule inhibitors. This approach is supported by the propensity of small molecules to specifically interact with amyloid assemblies at the molecular level and, thus, inhibit the self-assembly process15,16,17,18. Polyphenols, an important group of chemical entities found to be effective in the inhibition of amyloid structures formation, are composed of one or more small aromatic phenolic rings that specifically and efficiently inhibit amyloid aggregation. High concentrations of natural polyphenols can be found in a wide variety of plants, where they function in protection against diseases and UV light. This family of molecules, which represent the first generation of amyloid-based potential therapeutic agents, cause a dramatic reduction in amyloidogenic-related cell death, as well as efficient inhibition of amyloid self-assembly in vitro. Moreover, dietary polyphenols have shown beneficial health-promoting effects in chronic and neurodegenerative diseases19,20,21.
By employing a reductionist approach aimed to identify the minimal assembly units required for the formation of ordered amyloid aggregates, it was demonstrated that penta- and tetra- peptides can form well-ordered amyloid assemblies, which were also found to be cytotoxic via an apoptotic cell death pathway22,23. Interestingly, in aqueous solution, the diphenylalanine dipeptide (FF), the core recognition motif of the β-amyloid polypeptide, was also found to self-assemble into ordered nanotubular assemblies24,25, which show similarities to amyloids in terms of their mechanical rigidity, formation of reactive oxygen species and luminescence properties26,27,28,29. Thus, minimal building blocks assemblies, composed of peptides as short as dipeptides, reflect some of the fundamental physical properties of full-length amyloid proteins and polypeptides (Fig. 1a).
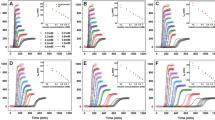
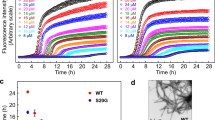
Reductionist approach toward amyloid formation and the inhibition of metabolite fibril self-assembly. a Systematic, reductionist approach toward the identification of minimal amyloidogenic building blocks. Full-length amyloid-β polypeptide (crystal structure PDB ID 2NAO and TEM adapted from ref. 66), KLVFFA hexapeptide (crystal structure PDB ID 2Y2A and TEM adapted from ref. 67), FF dipeptide (crystal structure modified from ref. 68 and TEM adapted from ref. 69) and the single amino-acid phenylalanine (crystal structure CCDC 985094 and TEM micrograph of phenylalanine at 4 mg/ml was taken as described in the Methods). b–g Inhibition of metabolite aggregation by polyphenol-based inhibitors. All metabolites were dissolved at 90 °C in PBS and mixed with the inhibitors, either EGCG or TA, at the stated concentrations, or with PBS as a control, followed by addition of ThT in PBS. ThT emission data at 480 nm (excitation at 450 nm) were measured over time. The results represent three biological repeats. b Adenine 8 mg/ml + EGCG. c Adenine 8 mg/ml + TA. d Phenylalanine 40 mg/ml + EGCG. e Phenylalanine 40 mg/ml + TA. f Tyrosine 2 mg/ml + EGCG. g Tyrosine 2 mg/ml + TA
Further studies, presented by our group and extended by others, have shown that even the single phenylalanine amino acid can self-assemble to form well-ordered supramolecular amyloid-like assemblies (Fig. 1a), possessing the common biophysical, biochemical, and ultrastructural amyloid properties30,31,32,33,34. Moreover, these assemblies display a cytotoxic effect on a neuronal cell model, which can be depleted by antibodies specifically raised against the assemblies30,32. Interestingly, in pathological conditions, a physiological accumulation of phenylalanine can be found in patients of the genetic phenylketonuria (PKU) inborn error of metabolism disorder35. The presence of antibodies raised against the phenylalanine fibrils in a PKU mouse model, as well as the presence of phenylalanine aggregate deposits in PKU patients’ brains post mortem, indicate a possible association between PKU and phenylalanine amyloid-like assemblies30.
To further extend the new paradigm linking genetic metabolic disorders to amyloid diseases, we have recently revealed the ability of other amino acids and small metabolites, which were found to accumulate in the course of different inborn error of metabolism disorders, to self-assemble into amyloid-like fibrillary structures. These metabolite-based fibrils were shown to bind amyloid-specific dyes, further implying on the formation of β-sheet supramolecular structures, and, similar to protein and peptide amyloids, to trigger cytotoxicity via an apoptotic cell death mechanism36,37. Each error of metabolism disorder is derived from a mutation in a single gene encoding a metabolic enzyme, thereby resulting in the accumulation of metabolites corresponding to the substrates of the faulty enzymes. These accumulated substrates can be toxic to cells and tissues, and thus cause severe abnormalities. Although these inherited disorders are relatively rare, collectively they constitute a very significant portion of pediatric genetic diseases38,39. Up until recently, the pathology and course of toxicity have been poorly understood, leading to the lack of a viable course of treatment, relying mainly on very restrictive diets.
Given the amyloid-like properties of small metabolite assemblies and the ability of known amyloid inhibitors to inhibit the formation of phenylalanine32,33,34,40 and tyrosine33,41 assemblies, it is of great interest to examine whether chemical agents known to inhibit the assembly of protein and peptide-based amyloid structures can also restrict the formation of these metabolite fibrils. Furthermore, owing to the similarity between the protein and metabolite assemblies, the later can serve as an attractive minimalistic model to probe the mechanism of amyloid inhibition, which is yet to be fully deciphered.
Here, we explore the inhibitory effect exerted on metabolite amyloid formation by two polyphenolic compounds, (-)-Epigallocatechin gallate (EGCG) and tannic acid (TA). Both inhibitors were previously shown to efficiently inhibit the formation of various protein amyloids in vitro and in vivo, and display beneficial preventive and therapeutic effects in neurodegenerative diseases42,43,44,45,46. We show that these two polyphenols successfully inhibit the self-assembly of adenine, phenylalanine, and tyrosine into amyloid-like fibrils, which accumulate in adenine phosphoribosyltransferase deficiency, PKU and tyrosinemia metabolic disorders, respectively. We further present the ability of generic polyphenol amyloid inhibitors to effectively reduce the formation of metabolite amyloid fibrils, as demonstrated by in vitro assays, molecular dynamics simulations and using a neuronal cell culture model. Our recent extension of the generic amyloid hypothesis, linking amyloid formation and metabolite amyloids in inborn error of metabolism disorders, and the inhibition of metabolite amyloids by natural small polyphenolic compounds as presented here, may lead to an innovative course of treatment for these less-explored metabolic disorders.
Results
Spectroscopic Monitoring Of Metabolite Amyloid Assembly Inhibition
We first examined the inhibitory potential of polyphenols in the context of metabolite amyloid aggregation. Two polyphenols, EGCG and TA, which were previously shown to inhibit fibril formation by various amyloidogenic proteins and peptides, were selected and their inhibition of metabolites aggregation was studied. EGCG was found to effectively inhibit fibril formation by the prion PrPsc protein42 and to reduce the toxicity of α-synuclein and Aβ polypeptide aggregates43,44,45. Similarly, TA has been previously described as the most potent polyphenol inhibitor of prion PrPsc aggregation42, and was found to inhibit Aβ fibrillation46. Thus, the effect of the inhibitors on the kinetics of amyloid fibril formation by adenine, phenylalanine, and tyrosine was explored. For this purpose, we used ThT, a common fluorescent, amyloid-specific dye, often used as a reporter of the amyloid self-assembly process, showing a strong increase in its fluorescent intensity upon binding to β-sheet-rich supramolecular structures. Thus, this fluorometric technique allows to determine the kinetics of the amyloid fibril assembly process, as well as to screen for compounds that might inhibit this process, based on changes in the intensity of ThT fluorescence13,14. The preparation of metabolite assemblies was performed as previously described36. To identify the metabolite concentrations that result in similar ThT signals, and therefore presumably in similar levels of amyloid-like fibrils, different metabolite concentrations were examined prior to the addition of the inhibitors. Although resulting in the formation of assemblies with similar morphologies, the required concentrations varied between the metabolites, as previously shown for different proteinaceous amyloids47. Next, these concentrations were used for time-dependent monitoring of ThT fluorescence intensity in the presence of the same increasing concentrations of each inhibitor (Fig. 1b–g).
Both EGCG and TA were found to inhibit adenine (8 mg/ml, ~ 60 mM) fibrils in a dose-dependent manner, as demonstrated by the ThT fluorescence assay, presenting near complete inhibition of adenine aggregate formation at their higher concentrations (Fig. 1b, c). Some adenine aggregates could be observed only at the lowest inhibitor concentration, where EGCG and TA treatment resulted in a reduction of ca. 50% (Fig. 1b) and 70% (Fig. 1c) in ThT intensity, respectively, indicating a stronger inhibitory effect of TA. Similarly, in the case of phenylalanine (40 mg/ml, ~ 242 mM) aggregates, both EGCG and TA inhibited the formation of the assemblies in a dose-dependent manner (Fig. 1d, e). EGCG presented a substantial inhibition at all concentrations (Fig. 1d), as reflected by the great reduction in the ThT fluorescence intensity curves, whereas TA presented a complete inhibition only at its highest concentration, a significant reduction in aggregation when applying the intermediate concentration and almost no effect using the lowest concentration (Fig. 1e). Finally, both inhibitors presented near complete inhibition of tyrosine (2 mg/ml, ~ 10 mM) aggregates formation at their highest concentration and a significant reduction at the lower concentrations used, where nearly no formation of aggregates was detected, as reflected from the ThT fluorescence intensity assay (Fig. 1f, g). It should be noted that ThT was previously indicated to promote amyloid formation48. Thus, the clear inhibitory effect observed is even more notable.
To examine whether the decrease in the ThT fluorescence signal was due to an interaction between the aromatic polyphenol compounds and the ThT dye, rather than inhibition of structure formation, the inhibitory effect of both polyphenols at their highest concentration was also examined using a turbidity assay. Consistent with the ThT fluorescence results, both EGCG and TA indeed hindered the formation of metabolite amyloids (Supplementary Fig. 1a–c). Moreover, the inhibitory effect was confirmed by dynamic light scattering (DLS) measurements (Supplementary Fig. 2). The addition of both inhibitors to the adenine assemblies resulted in a dramatic decline of the autocorrelation function (ACF) amplitude, indicating far fewer aggregates. In addition, there was a substantial reduction in the number of larger particles (longer correlation times) compared with the smaller particles (shorter correlation times) (Supplementary Fig. 2a). In the case of the phenylalanine assemblies, the addition of TA resulted in a dramatic decline of the ACF amplitude and a reduction in size, indicating the occurrence of far fewer aggregates. EGCG had a smaller effect compared with TA, resulting in a smaller amplitude and intermediate size shift (Supplementary Fig. 2b). Tyrosine solution without the inhibitors was too turbid even at low concentrations, and thus could not be characterized by DLS.
Ultrastructural Analysis Of The Effect Of Polyphenols On Metabolite Amyloid Formation
To gain insight into the morphological changes of the amyloid assemblies in the presence of the inhibitors, we employed transmission electron microscopy (TEM) and extreme high-resolution scanning electron microscopy (XHR-SEM), using the same concentrations of both metabolites and inhibitors as described above. As a control, TEM and XHR-SEM images of the polyphenolic inhibitors, at the highest concentrations used, were acquired (Supplementary Fig. 3). In the absence of the inhibitors, adenine and tyrosine presented the typical fibrillar morphology of amyloid assemblies (Fig. 2a, o and Supplementary Fig. 4a, g), whereas in the presence of either EGCG or TA, inhibition of fibrils formation was observed (Supplementary Fig. 4b, c, h, i) in a concentration-dependent manner (Fig. 2b–g, p–u). In the case of phenylalanine, lower metabolite concentrations resulted in the formation of a typical fibrillar amyloid morphology (Fig. 1a), yet at the higher concentration of 40 mg/ml, the concentration used in the ThT fluorescence assay, a different morphology was observed (Fig. 2h and Supplementary Fig. 4d). Both EGCG and TA hindered these structures (Supplementary Fig. 4e, f) in a concentration-dependent manner (Fig. 2i–n). Notably, the TA concentrations employed in both ThT fluorescence assay and TEM imaging were at least an order of magnitude lower than those of EGCG, thus indicating a more potent inhibitory effect of TA. This is consistent with the relative inhibitory effect of the two compounds toward the formation of amyloids by protein and peptide building blocks21. In addition, for all metabolites, the lack of the formation of assemblies, as observed using TEM, correlates well with the results of the ThT fluorescence assay.
Inhibition of metabolite amyloid fibril formation by EGCG and TA. a–u Adenine, phenylalanine and tyrosine were dissolved at 90 °C in PBS to a final concentration of 8 mg/ml, 40 mg/ml, and 2 mg/ml, respectively, mixed with PBS alone a, h, o or with the inhibitors, EGCG b–d, i–k, p–r, or TA e–f, l–n, s–u, at the concentrations stated below, and examined by TEM imaging. Scale bars 500 nm. The results represent three biological repeats. a Adenine. b Adenine + EGCG 1 mM. c Adenine + EGCG 0.5 mM. d Adenine + EGCG 0.05 mM. e Adenine + TA 0.1 mM. f Adenine + TA 0.01 mM. g Adenine + TA 0.0025 mM. h Phenylalanine. i Phenylalanine + EGCG 1 mM. j Phenylalanine + EGCG 0.5 mM. k Phenylalanine + EGCG 0.05 mM. l Phenylalanine + TA 0.1 mM. m Phenylalanine + TA 0.01 mM. n Phenylalanine + TA 0.0025 mM. o Tyrosine. p Tyrosine + EGCG 1 mM. q Tyrosine + EGCG 0.5 mM. r Tyrosine + EGCG 0.05 mM. s Tyrosine + TA 0.1 mM. t Tyrosine + TA 0.01 mM. u Tyrosine + TA 0.0025 mM
Inhibition Of Fibrils Cytotoxicity By Polyphenols
As mentioned above, the metabolite assemblies display a dose-dependent cytotoxic effect on a neuronal cell model within six hours36. Therefore, we examined the correlation between the inhibition of metabolite assembly formation by the polyphenol inhibitors and the resulting cytotoxicity. The adenine and phenylalanine concentrations used for this assay have been previously shown to reduce cell viability by at least 50%36, whereas tyrosine concentration was lower than the one presenting this effect owing to its limited solubility. SH-SY5Y cells were incubated with metabolite amyloid fibrils for 24 h, in the presence or absence of the inhibitors, a longer incubation period than was previously examined, thus resulting in a more significant reduction of cell viability. The concentration of the inhibitors was lower by an order of magnitude than the one used in the in vitro experiments, as higher concentrations were toxic to the cells. In the case of adenine, the inhibitors significantly increased cell viability in a dose-dependent manner (Fig. 3a, b), showing a threefold and twofold viability increase in the presence of the highest concentration of EGCG (Fig. 3a) and TA (Fig. 3b), respectively. In the case of phenylalanine, cell viability was almost completely restored in the presence of both inhibitors at their highest concentration, presenting a dose-dependent inhibition (Fig. 3c, d). Finally, in the case of tyrosine, for all the tested concentrations of both inhibitors, cell viability was significantly increased to ~ 70% (Fig. 3e, f). To examine whether the improvement in cell viability was owing to the addition of the polyphenolic inhibitors, which can alter cell viability by themselves, the effect of both inhibitors, when added to untreated cells or to cells following prior six hours treatment, with as the control or with the cytotoxic metabolite assemblies, was measured (Supplementary Fig. 5a–e). In all cases, later addition of the inhibitors did not improve cell viability. This further indicates that the restoration of cell viability was a result of lack of metabolites assembly formation owing to the addition of EGCG or TA.
Inhibition of the cytotoxicity of metabolite assemblies by EGCG and TA. Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in DMEM/Nutrient Mixture F12 (Ham’s) (1:1) without fetal bovine serum (FBS), to a final concentration of 2 mg/ml (~ 15 mM), 4 mg/ml (24 mM), and 2 mg/ml (10 mM), respectively, and mixed with or without the inhibitors at the stated concentrations. The control reflects medium with no metabolites, which was treated in the same manner. SH-SY5Y cells were incubated with the metabolites in the absence or presence of the inhibitors for 24 h, followed by addition of MTT reagents. Following a 4 h incubation, extraction buffer was added for an additional 0.5 h incubation, absorbance was determined at 570 nm. The data are presented as mean ± SD. The results represent three biological repeats (*p < 0.05 **p < 0.01) a–b Adenine (Ade) in the presence or absence of a EGCG and b TA. c–d Phenylalanine (Phe) in the presence or absence of c EGCG and d TA. e–f Tyrosine (Tyr) in the presence or absence of e EGCG and f TA
Acetylsalicylic Acid As A Control
In order to examine whether the inhibition presented by the polyphenol compounds was a result of their interaction with the metabolite amyloid structures, it was crucial to examine the possible activity of a compound that is known to be ineffective as an amyloid formation inhibitor. For this purpose, we used acetylsalicylic acid (ASA), a small-molecule applied to treat pain, fever, and inflammation, which has been previously used as a negative control for amyloid inhibitors of the IAPP amyloid polypeptide, and did not affect the amyloid formation process16,49. When tested as an inhibitor of metabolite amyloid formation, ASA did not affect aggregation, even at the highest concentration used for the other inhibitors, as was observed using ThT fluorescence assay, TEM imaging and the SH-SY5Y neuronal cell culture (Supplementary Fig. 6–8).
Analysis Of The Mechanism Underlying The Inhibition Of Metabolite Amyloid Formation
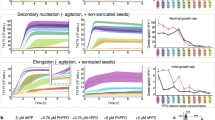
In order to better understand the mechanism by which the polyphenol inhibitors affect the metabolite fibril formation, EGCG and TA were added to the system at different time points of the metabolite fibrillation process (after 0, 0.5, 1, and 2 h), and the kinetics of metabolite fibril formation was monitored using the ThT fluorescence assay. Both endpoint after overnight (Fig. 4a–c) and kinetics (Supplementary Fig. 9) data were recorded. Overall, EGCG inhibited metabolite fibril formation even when added at later stages of fibrillation. TA inhibited the formation when added at an earlier time point, while showing a much lower impact when added at the later stages of fibrillation.
The effect of polyphenol inhibitors added at different time points on fibril self-assembly inhibition. All metabolites were dissolved at 90 °C in PBS and ThT in PBS was added. The inhibitors, EGCG (1 mM, orange) or TA (0.1 mM, gray), or PBS (blue) as a control, were added at different time points (0, 0.5, 1, and 2 h). Following excitation at 450 nm, ThT emission data at 480 nm were measured over time for an overnight and endpoint fluorescence readings are presented. a Adenine 8 mg/ml. b Phenylalanine 40 mg/ml. c Tyrosine 2 mg/ml. d Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in DMEM/Nutrient Mixture F12 (Ham’s) (1:1) without FBS, to a final concentration of 2, 4, and 2 mg/ml, respectively, and mixed after 2 hours with EGCG (orange, 0.1 mM) or TA (gray, 0.01 mM) following an overnight incubation. The samples to which no inhibitor was added were similarly mixed after 2 h with medium without FBS (blue). Then, SH-SY5Y cells were incubated with the metabolites in the absence or presence of the inhibitors for 24 h, followed by addition of MTT reagents. Following a 4 h incubation, extraction buffer was added and after an additional 0.5 h incubation, absorbance was determined at 570 nm. The data are presented as mean ± SD. The results represent three biological repeats
Next, we examined whether the addition of inhibitors at later time points of metabolite fibrillation had an effect on their resulting cytotoxicity (Fig. 4d). When EGCG was added after 2 hours, cell viability was significantly restored. However, the addition of TA did not restore cell viability, which decreased to similar levels as the control without the inhibitor. These results are in agreement with the ThT fluorescence assay (Fig. 4a–c), suggesting that TA mostly inhibits fibrilization at the early stages of nucleation, whereas EGCG acts both at the early and later stages of fibril formation. Taken together, the combination of results obtained via both the in vitro and neuronal cell model systems provides important insights into the differential mode of action of each inhibitor.
Molecular Dynamics Mechanistic Studies
To gain further insight into the distinct inhibition mechanisms of the two inhibitors, we performed molecular dynamics (MD) simulations of the potential inhibition activity of EGCG, TA, and ASA during the formation of metabolites fibrillar assemblies. In particular, we examined the interactions between inhibitor molecules and both free solvated metabolite molecules and small metabolite fibril nuclei. Figure 5a–d presents the MD-simulation of phenylalanine coupled to TA and EGCG (see Supplementary Fig. 10–13 for additional data). The interaction energy was determined by selecting an inhibitor and calculating the average binding energy per metabolite within 4.5 Å of that inhibitor molecule. We partitioned the trajectory into two sections and averaged the binding energy of each section. The midpoint between the averages was chosen as the final value. Error bars were determined from the deviation of the averages of each partition and the midpoint.
Simulations of metabolites and inhibitors coupling. a–d Snapshots of TA a, c and EGCG b, d coupling with Phe metabolites in free a, b and crystal c, d forms. Free metabolites are shown in blue; metabolite molecules in crystal are shown in red, inhibitor molecules are shown in green. Scale bars in a–b, c–d represent 1 Å. e The average total coupling energies between inhibitors and either free (blue) or crystalline (red) metabolites, within 3.5 Å of the inhibitor, normalized per inhibitor and metabolite. f The average number of free metabolites around inhibitors, normalized by the concentration of free metabolites and inhibitors in the system. g The average number of inhibitors adsorbed to metabolite crystals, normalized by the concentration of inhibitors and the surface areas of crystals. Column heights were determined by blocking the trajectory into two halves and calculating the midpoint between the averages of each block. Error bars are the deviation of the average of each half of the trajectory and the midpoint
As revealed by the simulations, the average binding energies of the metabolites to both TA and EGCG are similar, whereas the binding energies to ASA are 2–3 times lower. Moreover, there is a small variation of these energies between the studied metabolites, with the binding energy of adenine found to be ~ 50% of those of phenylalanine and tyrosine (Fig. 5e).
Next, we calculated the number of free metabolites coupled to inhibitors, normalized by the concentrations of metabolites and inhibitors (Fig. 5f). We determined averages and uncertainties using the same procedure we used in determining the interaction energy. This analysis shows that a larger number of free metabolites can adsorb to TA as compared with EGCG or ASA, despite similar binding energies of these inhibitors to individual metabolites (Fig. 5e). This difference results from the significantly larger surface area of TA, as compared with EGCG or ASA. These observations correlate well with the experimental results showing that TA efficiently binds free metabolites, whereas EGCG reaches the same efficiency only at about an order of magnitude higher concentration.
In addition, the number of inhibitors bound to the metabolites in their crystalline state and normalized to the concentrations of inhibitors and the surface areas of the metabolite crystals was calculated (Fig. 5g). The data reveal that at the same concentrations, TA and EGCG have a similar propensity for binding to the ordered lattice, whereas ASA has a very low binding propensity. This explains the experimental observations showing that EGCG can block the growth of fibrils even when added at later stages, as it binds to the crystal at about an order of magnitude higher concentration. Given the relative dimensions of TA and EGCG, EGCG can cover the crystals to a larger extent at the higher concentrations used. Moreover, there is a twofold decrease of inhibitors bound to the tyrosine crystal, probably owing to the fact that the preferable binding sites are not fully exposed in this form as its facets are jagged, unlike adenine and phenylalanine surfaces, preventing the formation of π–π stacking interactions (Fig. 5g).
Discussion
One of the interesting observations regarding the formation of amyloid fibrils is their generic inhibition by various polyphenol molecules. Different polyphenols can serve as universal inhibitors of the process of amyloid fibril formation regardless of the amino-acid composition of the protein and polypeptide assemblies. Nevertheless, the mechanism of inhibition and the specificity and dynamics of inhibitor interaction are not fully understood, giving rise to a genuine need for minimalistic models on which experimental and computational approaches can be applied.
Here, we reveal that two aromatic polyphenolic compounds, EGCG and TA, which have been previously shown to generically inhibit the formation of protein and peptide-based amyloid structures, can also inhibit the formation of metabolite amyloid fibrils, even when applied at a very low molar ratio. The examined concentrations, resulting in the same end-effects, varied between the metabolites, as previously shown for different protein and polypeptide amyloid aggregates47. We demonstrated the activity of these inhibitors both in vitro, using ThT fluorescence assay and TEM analysis, and in vivo, using the 3-(4,5-dimethylthiazolyl-2)-2,5-diphenyltetrazolium bromide (MTT) cell viability assay preformed on a neuronal cell model. The addition of the polyphenols at different time points provided a key mechanistic information about the process of inhibition by EGCG and TA and its correlation to reduced cytotoxicity. We demonstrated that in spite of the generic inhibition, the compounds function via two different mechanisms. Although EGCG affects both early and later stages of fibrillation, TA is only effective in the early state. To gain further insights on these different inhibition mechanisms, we preformed MD simulations. The simple chemical composition of the metabolites and the information about their molecular packing allowed to decipher the mechanism of amyloid formation, making them a potential model for understanding and controlling protein amyloid formation. The simulation provided molecular details about the nature of the binding of the two inhibitors to the studied metabolites in both monomeric and crystalline forms. It appears that each metabolite binds to both inhibitors with similar energy. At the molarity experimentally used here, the two inhibitors also bind a comparable number of free metabolites. However, at this molarity, the number of EGCG molecules binding to the crystalline form is several fold higher than the TA molecules, providing a theoretical framework to understand the distinct effect of the inhibitors when added at different stages of amyloid self-assembly.
The metabolites studied in the scope of this work are known to accumulate in different inborn error of metabolism disorders and, as we have previously shown, can self-assemble and form supramolecular β-sheet-like amyloid structures. In protein and peptide-based amyloids aggregation, the core β-sheet structure seems to be primarily stabilized by hydrogen bonds50,51. Based on their structural resemblance and previous findings regarding phenylalanine and tyrosine assemblies’ formation41, we can speculate that the amyloid metabolite supramolecular structures are also stabilized via hydrogen bonds, which may support the findings derived from the simulations presented here regarding the nature of inhibitor-metabolite interactions. This was indeed the case for phenylalanine, for which the newly determined crystal structure of the zwitterionic state, that promotes fibrils formation, shows a clear hydrogen-bonding network as observed in peptides and proteins52. Moreover, the overlay of computer-generated putative poly-phenylalanine β-strands on the crystal structures resulted in remarkable superposition36. The use of known protein and peptide amyloid inhibitors in order to affect metabolite amyloid structure formation further supports the notion of an extended family of amyloid structures that includes protein, peptides, and metabolites. Finally, the less-explored concept of error of metabolism disorders as amyloid diseases may thus lead to the development of new therapeutic strategies, in addition to the current dietary restrictions.
Methods
Materials
Metabolites (phenylalanine and tyrosine purity ≥ 98%; adenine purity ≥ 99%), thiazolyl blue tetrazolium bromide 98% and thioflavine T were purchased from Sigma-Aldrich (Rehovot, Israel). Dulbecco’s modified Eagle’s medium (DMEM)/Nutrient Mixture F12 (Ham’s) (1:1) was purchased from Biological Industries (BI) (Kibbutz Beit Haemek, Israel). Dimethylformamide (DMF) was purchased from Bio-lab (Jerusalem, Israel). Sodium n-dodecyl sulfate (SDS) 99% was purchased from Tzamal D-Chem (Petah Tikva, Israel).
ThT Kinetics Fluorescence Assay
Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in PBS to a final concentration of 8, 40, and 2 mg/ml, respectively, and monomeric solutions of the metabolites were obtained. Next, these solutions were mixed with the inhibitors, EGCG or TA, at the stated concentrations in a black 96-well, clear, and flat bottom microplate (Greiner). As a control, metabolites were diluted with PBS alone to the same final concentrations. For the time points experiments, the inhibitors, EGCG (1 mM) and TA (0.1 mM), or PBS as a control, were added at different time points (0, 0.5, 1, and 2 h). ThT in PBS was added to a final concentration of 40 µM (for phenylalanine) or 20 µM (for adenine and tyrosine). Following excitation at 450 nm, ThT emission data at 480 nm were measured over time and recorded using a TECAN Infinite 200 PRO plate reader. Data processing was preformed using OriginLab software.
Turbidity Measurements
Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in PBS to a final concentration of 8, 40, and 2 mg/ml, respectively, and monomeric solutions of the metabolites were obtained. Next, these solutions were mixed with the inhibitors, EGCG (0.1 mM) or TA (0.01 mM), in a 96-well microplate (Corning). As a control, metabolites were diluted with PBS alone to the same final concentrations. Time-dependent changes in solution turbidity were determined overnight by optical density measurements (600 nm) using a Biotek Synergy HT microplate reader at 25 °C.
DLS
For DLS experiments, adenine (2 mg/ml) and phenylalanine (4 mg/ml) were dissolved at 90 °C in PBS and mixed with the inhibitors, either EGCG (0.1 mM) or TA (0.01 mM), or with PBS as a control. The PBS and inhibitors solutions were both filtered using a 0.02 µm syringe filter prior to the addition of metabolites. After an overnight incubation, 200 µl of the solutions were loaded in quadruplicates into a black 96-well, clear and flat bottom microplate (Greiner). The autocorrelation amplitude of the solutions was measured applying the DynaPro II Plate Reader system (Wyatt Technology). For adenine samples, 5 × 5 s per well was acquired, and for phenylalanine, 4 × 5 acquisitions × 5 s per well was acquired. The results represent an average of each solution quadruplicate.
Transmission Electron Microscopy
Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in PBS to a final concentration of 8, 40, and 2 mg/ml, respectively, to obtain monomeric solutions of the metabolites. Next, these solutions were mixed with the inhibitors, EGCG or TA, at the stated concentrations. As controls, metabolites were diluted with PBS alone to the same final concentrations and the inhibitors alone without the metabolites were examined as well. In Fig. 1a, phenylalanine was dissolved at 90 °C in PBS to a final concentration of 4 mg/ml. Samples of the metabolites, in the presence or absence of inhibitors, or samples of inhibitors without the metabolites, were incubated overnight at room temperature. Subsequently, 10 µl samples were placed on 400-mesh copper grids. After 1 min, excess fluids were removed. Samples were viewed using a JEOL 1200EX electron microscope operating at 80 kV.
Extreme High-resolution Scanning Electron Microscopy
Adenine, phenylalanine, and tyrosine were dissolved at 90 °C in PBS to a final concentration of 8, 40, and 2 mg/ml, respectively, and monomeric solutions of the metabolites were obtained. Next, these solutions were mixed with the inhibitors, EGCG (1 mM) or TA (0.1 mM). As controls, metabolites were diluted with PBS alone to the same final concentrations and the inhibitors alone without the metabolites were examined as well. Samples of the metabolites, in the presence or absence of inhibitors, or samples of inhibitors without the metabolites, were incubated overnight at room temperature. Subsequently, 10 µl samples of these solutions were placed on a glass slide and left to dry at room temperature. Samples were then coated with 3 nm chrome and viewed using Verios XHR 460 L SEM operating at 3 kV. Micrographs displayed are representative of three independent experiments conducted.
Cell Cytotoxicity Experiments
SH-SY5Y cells (2 × 105 cells/ml) were cultured in 96-well tissue microplates (100 µl per well) and allowed to adhere overnight at 37 °C. Each plate was divided and only half of it was plated with cells, with the other half later serving as a control for the solutions alone. The treatment solutions were prepared as follows: adenine, phenylalanine, and tyrosine were dissolved at 90 °C in DMEM/Nutrient Mixture F12 (Ham’s) (1:1) (Biological Industries) without fetal bovine serum (FBS), to a final concentration of 2, 4, and 2 mg/ml, respectively, to obtain monomeric solutions of the metabolites. Next, these solutions were mixed with the inhibitors, EGCG or TA, at the stated concentrations. As a control, metabolites were diluted with DMEM/Nutrient Mixture F12 without FBS to the same final concentrations. The inhibitors without the metabolites were examined as well. For the inhibition mechanism experiments, EGCG (100 µM), TA (10 µM), or DMEM/Nutrient Mixture F12 (Ham’s) without FBS as a control, were added to the metabolites samples after 2 hours. The samples were incubated overnight at room temperature. Then, cells were washed and treated with the solutions (100 µl per well), followed by overnight incubation at 37 °C. In order to explore the effect of a later addition of the inhibitors on cells viability, EGCG (100 µM), and TA (10 µM), were added after 6 h to untreated cells or to cells that were pre-treated with metabolite fibrils, followed by overnight incubation at 37 °C. Cell viability was evaluated using the MTT assay. In brief, 10 µl of 5 mg/ml MTT dissolved in PBS were added to each well. After a 4 h incubation at 37 °C, 100 µl extraction buffer (20% SDS dissolved in a mixture of 50% DMF and 50% DDW (pH 4.7)) were added to each well, and the plates were incubated again at 37 °C for 30 min. Finally, color intensity was measured using an ELISA reader at 570 nm. The data are presented as mean ± SEM.
Statistical Analysis
Two-tailed Student’s t-test was performed when two groups were compared.
Force Field Fitting
The force field parameters for inhibitor molecules were calculated using Gaussian0953. Owing to the large size of TA, this molecule was divided to two sections in the calculations: (1) an inner core, which is a beta-d glucose derivative, with each hydroxyl group being methylated; (2) ligands, with the ester group that is bonded to the core methylated. In the calculations of the EGCG core, the two main fused rings and the ester group were considered; the aromatic groups of EGCG were replaced with methyl groups. Force field parameters for aromatic groups used the results from TA calculations or were approximated with methyl groups. Geometries and force constants of inhibitors were determined using MP2/6-31 g(d)//MP2/6-31 g(d) level for the EGCG core and TA core. Owing to the extent of ligand calculations, MP2/6-31 g(d)//MP2/6-31 g methods were used instead. Force field fitting was performed using VMD Force Field Toolkit Plugin54. To determine dihedral parameters, MP2 level of calculations were also used to derive a quantum target, with the angle scanned six steps in both positive and negative directions at a step size of 15˚. Charges were determined using the ChelpG algorithim55.
Solvated Metabolite Simulations
Systems formed by 288 ADN, PHE, and TYR freely solvated metabolite molecules were simulated in the presence of a small nucleation crystal of 125, 108, and 100 metabolite molecules, respectively. Each system consisted of 7 TA, 10 EGCG, or 15 ASA molecules. Systems with TA or EGCG were simulated for 20 ns, and those with ASA for 30 ns, using NAMD256 and the CHARMM force field. The systems were simulated in (NaCl) = 0.15 M aqueous solution, in order to imitate physiological conditions. The Langevin dynamics with a damping coefficient of 1 ps−1 and a time step of 2 fs was used to describe systems in a NPT ensemble at a pressure of 1 atm and a temperature of 310 K. During minimization, pre-equilibration, and equilibration, Particle Mesh Ewald57 was used with a grid spacing of 1.0. The non-bonded interactions used the SHAKE switching algorithm with a switch on/off distance of 10/12 Å. Non-bonded pairs lists were 13.5 Å, with the list updated every 20 steps; 1–4 non-bonded interactions were not scaled. There were 50,000 steps of minimization followed by 2 ns of equilibration, after which the simulations were performed. The same approach was used in the simulations of inhibitors binding to metabolite crystals.
Crystalline Metabolite Simulations
Simulated bulk-like crystals of ADN, PHE, and TYR had 564, 768, and 800 molecules, respectively, which were cut from their bulk crystal structures52,58,59. The solutions of simulated bulk crystals contained 25 TA, 50 EGCG, or 50 ASA molecules, and no free metabolite molecules. No specific fibril structures were considered.
Solvated Metabolite Interaction Energies
Average interaction energies between inhibitors and free metabolites or crystals were calculated from the simulated systems. The CHARMM force field60,61,62,63,64 and NAMDenergy plugin in VMD65 were used to determine the strength of total interaction energy for each inhibitor by calculating vdW and electrostatic interaction energies. Interaction energy between each inhibitor and any metabolite molecule, which had at least one atom within 3.5 Å of any atom of the selected inhibitor, was considered for the calculation. For each snapshot, the total energy and number of metabolites interacting with the inhibitor was recorded. Interaction energy per metabolite and inhibitor was determined at each snapshot. When averaging the interaction energy, only cases with at least one metabolite interacting with the inhibitor were included. The average number of free metabolites was computed for the entire trajectory. The average number of interacting metabolites was normalized with respect to the concentrations of both inhibitors and solvated metabolites (Supplementary Table 1).
Crystalline Metabolites Interaction Energies
Similar to solvated metabolites, we chose one inhibitor for the crystal cases and evaluated its interaction energies with metabolites on the crystal. Metabolites on crystal, which were within 3.5 Å of the inhibitor, were considered. The average interaction energy was computed using the same procedure as for the solvated metabolites. The number of inhibitors interacting with the crystal was determined by calculating any inhibitor with at least one atom within 3.5 Å of the crystal. This number was averaged throughout the trajectory. Then, it was normalized with respect to the concentration of inhibitors and surface area of the crystal, obtained at the end of the trajectory (Table S1), because some metabolite molecules dissolved from the crystals during the simulation.
Data Availability
The authors declare that all of the data supporting the findings of this study are available within the article and its Supplementary Information files or from the author upon reasonable request.
References
Knowles, T. P., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Ross, C. A. & Poirier, M. A. Protein aggregation and neurodegenerative disease. Nat. Med. 10, S10–S17 (2004).
Rambaran, R. N. & Serpell, L. C. Amyloid fibrils: abnormal protein assembly. Prion 2, 112–117 (2008).
Kapurniotu, A. Shedding light on Alzheimer’s β‐amyloid aggregation with chemical tools. Chembiochem 13, 27–29 (2012).
Gazit, E. The “correctly folded” state of proteins: is it a metastable state? Angew. Chem. Int. Ed. 41, 257–259 (2002).
Eichner, T. & Radford, S. E. A diversity of assembly mechanisms of a generic amyloid fold. Mol. Cell 43, 8–18 (2011).
Gruden, M. A. et al. Intranasal administration of alpha-synuclein aggregates: a Parkinson’s disease model with behavioral and neurochemical correlates. Behav. Brain Res. 263, 158–168 (2014).
Milanesi, L. et al. Direct three-dimensional visualization of membrane disruption by amyloid fibrils. Proc. Natl. Acad. Sci. USA 109, 20455–20460 (2012).
Grudzielanek, S. et al. Cytotoxicity of insulin within its self-assembly and amyloidogenic pathways. J. Mol. Biol. 370, 372–384 (2007).
Aguzzi, A. & O’connor, T. Protein aggregation diseases: pathogenicity and therapeutic perspectives. Nat. Rev. Drug. Discov. 9, 237–248 (2010).
Adamcik, J. et al. Understanding amyloid aggregation by statistical analysis of atomic force microscopy images. Nat. Nanotechnol. 5, 423–428 (2010).
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006).
Biancalana, M. & Koide, S. Molecular mechanism of Thioflavin-T binding to amyloid fibrils. BBA-Proteins Proteom. 1804, 1405–1412 (2010).
LeVine, H. Thioflavine T interaction with synthetic Alzheimer’s disease beta-amyloid peptides: detection of amyloid aggregation in solution. Protein Sci. 2, 404–410 (1993).
DaSilva, K. A., Shaw, J. E. & McLaurin, J. Amyloid-β fibrillogenesis: structural insight and therapeutic intervention. Exp. Neurol. 223, 311–321 (2010).
Young, L. M. et al. Screening and classifying small-molecule inhibitors of amyloid formation using ion mobility spectrometry–mass spectrometry. Nat. Chem. 7, 73–81 (2015).
Feng, B. Y. et al. Small-molecule aggregates inhibit amyloid polymerization. Nat. Chem. Biol. 4, 197–199 (2008).
Doig, A. J. & Derreumaux, P. Inhibition of protein aggregation and amyloid formation by small molecules. Curr. Opin. Struct. Biol. 30, 50–56 (2015).
Bhullar, K. S. & Rupasinghe, H. P. Polyphenols: multipotent therapeutic agents in neurodegenerative diseases. Oxid. Med. Cell Longev. 2013, 891748 (2013).
Ebrahimi, A. & Schluesener, H. Natural polyphenols against neurodegenerative disorders: potentials and pitfalls. Ageing Res. Rev. 11, 329–345 (2012).
Porat, Y., Abramowitz, A. & Gazit, E. Inhibition of amyloid fibril formation by polyphenols: structural similarity and aromatic interactions as a common inhibition mechanism. Chem. Biol. Drug. Des. 67, 27–37 (2006).
Tenidis, K. et al. Identification of a penta-and hexapeptide of islet amyloid polypeptide (IAPP) with amyloidogenic and cytotoxic properties. J. Mol. Biol. 295, 1055–1071 (2000).
Reches, M., Porat, Y. & Gazit, E. Amyloid fibril formation by pentapeptide and tetrapeptide fragments of human calcitonin. J. Biol. Chem. 277, 35475–35480 (2002).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Adler-Abramovich, L. et al. Self-assembled arrays of peptide nanotubes by vapour deposition. Nat. Nanotechnol. 4, 849–854 (2009).
Yan, X., Zhu, P. & Li, J. Self-assembly and application of diphenylalanine-based nanostructures. Chem. Soc. Rev. 39, 1877–1890 (2010).
Adler-Abramovich, L. & Gazit, E. The physical properties of supramolecular peptide assemblies: from building block association to technological applications. Chem. Soc. Rev. 43, 6881–6893 (2014).
Souza, M. I. et al. The role of water and structure on the generation of reactive oxygen species in peptide/hypericin complexes. J. Pept. Sci. 20, 554–562 (2014).
Amdursky, N. et al. Blue luminescence based on quantum confinement at peptide nanotubes. Nano. Lett. 9, 3111–3115 (2009).
Adler-Abramovich, L. et al. Phenylalanine assembly into toxic fibrils suggests amyloid etiology in phenylketonuria. Nat. Chem. Biol. 8, 701–706 (2012).
Singh, V., Rai, R. K., Arora, A., Sinha, N. & Thakur, A. K. Therapeutic implication of L-phenylalanine aggregation mechanism and its modulation by D-phenylalanine in phenylketonuria. Sci. Rep. 4, 3875 (2014).
De Luigi, A. et al. Doxycycline hinders phenylalanine fibril assemblies revealing a potential novel therapeutic approach in phenylketonuria. Sci. Rep. 5, 15902 (2015).
Banik, D., Banerjee, P., Sabeehuddin, G. & Sarkar, N. Effects of a common worldwide drink (Beer) on L-Phenylalanine and L-Tyrosine fibrillar assemblies. Chem. Phys. Lett. 687, 44–53 (2017).
Banik, D., Dutta, R., Banerjee, P., Kundu, S. & Sarkar, N. Inhibition of fibrillar assemblies of l-phenylalanine by crown ethers: a potential approach toward phenylketonuria. J. Phys. Chem. B. 120, 7662–7670 (2016).
Hanley, W. B. Adult phenylketonuria. Am. J. Med. 117, 590–595 (2004).
Shaham-Niv, S., Adler-Abramovich, L., Schnaider, L. & Gazit, E. Extension of the generic amyloid hypothesis to nonproteinaceous metabolite assemblies. Sci. Adv. 1, e1500137 (2015).
Shaham‐Niv, S. et al. Formation of apoptosis‐inducing amyloid fibrils by tryptophan. Isr. J. Chem. 57, 729–737 (2016).
Ferrier, D. & Harvey, R. Lippincott’s Ilustrated Reviews: Biochemistry. (Lippincott Williams & Wilkins, Philadelphia, 2014).
Valle, D. et al. The Online Metabolic and Molecular Bases of Inherited Disease. (McGraw-Hill, New York, 2014).
Tian, Y. et al. Crown ethers attenuate aggregation of amyloid beta of Alzheimer’s disease. Chem. Commun. 50, 15792–15795 (2014).
Banik, D., Kundu, S., Banerjee, P., Dutta, R. & Sarkar, N. Investigation of fibril forming mechanisms of l-phenylalanine and l-tyrosine: microscopic insight toward phenylketonuria and tyrosinemia type II. J. Phys. Chem. B. 121, 1533–1543 (2017).
Kocisko, D. A. et al. New inhibitors of scrapie-associated prion protein formation in a library of 2000 drugs and natural products. J. Virol. 77, 10288–10294 (2003).
Ehrnhoefer, D. E. et al. EGCG redirects amyloidogenic polypeptides into unstructured, off-pathway oligomers. Nat. Struct. Mol. Biol. 15, 558–566 (2008).
Bieschke, J. et al. EGCG remodels mature α-synuclein and amyloid-β fibrils and reduces cellular toxicity. Proc. Natl. Acad. Sci. USA 107, 7710–7715 (2010).
Rezai-Zadeh, K. et al. Green tea epigallocatechin-3-gallate (EGCG) modulates amyloid precursor protein cleavage and reduces cerebral amyloidosis in Alzheimer transgenic mice. J. Neurosci. 25, 8807–8814 (2005).
Ono, K., Hasegawa, K., Naiki, H. & Yamada, M. Anti-amyloidogenic activity of tannic acid and its activity to destabilize Alzheimer’s β-amyloid fibrils in vitro. Biochim. Biophys. Acta 1690, 193–202 (2004).
Tartaglia, G. G., Pechmann, S., Dobson, C. M. & Vendruscolo, M. Life on the edge: a link between gene expression levels and aggregation rates of human proteins. Trends Biochem. Sci. 32, 204–206 (2007).
D’Amico, M. et al. Thioflavin T promotes Aβ (1–40) amyloid fibrils formation. J. Phys. Chem. Lett. 3, 1596–1601 (2012).
Tu, L. H., Noor, H., Cao, P. & Raleigh, D. P. Aspirin, diabetes, and amyloid: re-examination of the inhibition of amyloid formation by aspirin and ketoprofen. Acs. Chem. Biol. 9, 1632–1637 (2014).
Dobson, C. M. Protein misfolding, evolution and disease. Trends Biochem. Sci. 24, 329–332 (1999).
Tsemekhman, K., Goldschmidt, L., Eisenberg, D. & Baker, D. Cooperative hydrogen bonding in amyloid formation. Protein Sci. 16, 761–764 (2007).
Mossou, E. et al. The self-assembling zwitterionic form of L-phenylalanine at neutral pH. Acta Crystallogr. C. Struct. Chem. 70, 326–331 (2014).
Frisch, M. J. et al. Gaussian 09 (Gaussian, Inc., Wallingford CT, 2016).
Mayne, C. G., Saam, J., Schulten, K., Tajkhorshid, E. & Gumbart, J. C. Rapid parameterization of small molecules using the force field toolkit. J. Comput. Chem. 34, 2757–2770 (2013).
Breneman, C. M. & Wiberg, K. B. Determining atom-centered monopoles from molecular electrostatic potentials. The need for high sampling density in formamide conformational analysis. J. Comput. Chem. 11, 361–373 (1990).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10100 (1993).
Mahapatra, S., Nayak, S. K., Prathapa, S. J. & Guru Row, T. N. Anhydrous adenine: crystallization, structure, and correlation with other nucleobases. Cryst. Growth Des. 8, 1223–1225 (2008).
Mostad, A., Nissen, H. M. & Romming, C. L-Tyrosine_structure.pdf. Acta Chem. Scand. 26, 3819–3833 (1972).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Mackerell, A. D., Feig, M. & Brooks, C. L. Extending the treatment of backbone energetics in protein force fields: Limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulation. J. Comput. Chem. 25, 1400–1415 (2004).
Vanommeslaeghe, K., Raman, E. P. & M. J., A. Automation of the CHARMM general force field (CGenFF) II: Assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52, 3155–3168 (2012).
Vanommeslaeghe, K. & MacKerell, A. D. Automation of the CHARMM general force field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model 52, 3144–3154 (2012).
Yu, W., He, X., Vanommeslaeghe, K. & MacKerell, A. D. Extension of the CHARMM general force field to sulfonyl-containing compounds and its utility in biomolecular simulations. J. Comput. Chem. 33, 2451–2468 (2012).
Humphry, W., Dalke, A. & Schulten, K. VMD - visual MolecularDynamics. J. Mol. Graph. 14, 33–38 (1996).
Ahmed, M. et al. Structural conversion of neurotoxic amyloid-β1–42 oligomers to fibrils. Nat. Struct. Mol. Biol. 17, 561–567 (2010).
Colletier, J. P. et al. Molecular basis for amyloid-β polymorphism. Proc. Natl. Acad. Sci. USA 108, 16938–16943 (2011).
Görbitz, C. H. The structure of nanotubes formed by diphenylalanine, the core recognition motif of Alzheimer’s β-amyloid polypeptide. Chem. Commun. 22, 2332–2334 (2006).
Conte, M. P., Singh, N., Sasselli, I. R., Escuder, B. & Ulijn, R. V. Metastable hydrogels from aromatic dipeptides. Chem. Commun. 52, 13889–13892 (2016).
Acknowledgements
This work was supported by the Israel Science Foundation (grant no. 802/15; E.G.), The FEBS Long-Term Fellowship (A.L.), The Strauss Institute (fellowship; S.S.-N.), The Adelis Forever Foundation (E.G.), and the NSF Division of Materials Research (grant no. 1506886; P.K.). We thank D. Some of Wyatt Technology Corp. and A. Tsadok of Danyel Biotech for performing the dynamic light scattering analysis and the members of the Gazit, Adler-Abramovich, Vuković, and Král groups for helpful discussions.
Author information
Authors and Affiliations
Contributions
S.S.-N., L.A.-A., A. L., and E.G. conceived and designed the experiments. S.S.-N. and D.Z planned and performed the experiments. P.R., L.V., and P. K. planned and performed the MD simulations. S.S.-N., L.A.-A., P.R, L. V., P.K, and E.G. wrote the paper. All authors discussed the results, provided intellectual input and critical feedback, and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shaham-Niv, S., Rehak, P., Zaguri, D. et al. Differential inhibition of metabolite amyloid formation by generic fibrillation-modifying polyphenols. Commun Chem 1, 25 (2018). https://doi.org/10.1038/s42004-018-0025-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0025-z
This article is cited by
-
Biomolecule-mimetic nanomaterials for photothermal and photodynamic therapy of cancers: Bridging nanobiotechnology and biomedicine
Journal of Nanobiotechnology (2022)
-
Metabolite medicine offers a path beyond lists of metabolites
Communications Chemistry (2021)
-
Induction of retinopathy by fibrillar oxalate assemblies
Communications Chemistry (2020)
-
Fibril formation and therapeutic targeting of amyloid-like structures in a yeast model of adenine accumulation
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.