Abstract
Evolution of microbial traits depends on the interaction of a species with its environment as well as with other coinhabiting species. However, our understanding of the evolution of specific microbial traits, such as antibiotic resistance in complex environments is limited. Here, we determine the role of interspecies interactions on the dynamics of nitrofurantoin (NIT) resistance selection among Escherichia coli. We created a synthetic two-species community comprised of two variants of E. coli (NIT susceptible and resistant) and Bacillus subtilis in minimal media with glucose as the sole carbon source. We show that the presence of B. subtilis significantly slows down the selection for the resistant E. coli mutant when NIT is present and that this slowdown is not due to competition for resources. Instead, the dampening of NIT resistance enrichment is largely mediated by extracellular compounds produced by B. subtilis with the peptide YydF playing a significant role. Our results not only demonstrate the impact of interspecies interactions on the evolution of microbial traits but also show the importance of using synthetic microbial systems in unravelling relevant interactions and mechanisms affecting the evolution of antibiotic resistance. This finding implies that interspecies interactions should be considered to better understand and predict resistance evolution in the clinic as well as in nature.
Similar content being viewed by others
Introduction
Antibiotic resistance is a medically important bacterial trait that can arise in a population as a result of de novo mutations or by horizontal gene transfer. Resistant strains typically rapidly outcompete susceptible strains at concentrations above the minimal inhibitory concentration (MIC) leading to dominance or fixation of the trait1,2. However, the emergence and subsequent enrichment of a resistant strain towards dominance or fixation have also been observed at antibiotic concentrations far below the MIC, termed sub-MIC1,3,4,5,6,7,8. Bacteria can be exposed to sub-MIC levels of antibiotics in clinical settings during therapeutic use9,10 as well in the environment (e.g., water, soils, and food) due to anthropogenic pollution9,11,12. Bacterial growth at sub-MIC not only broadens the range of probable mutations leading to antibiotic resistance but the weaker selective pressure by the antibiotic also affects the rate of resistance enrichment3,7.
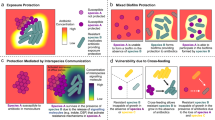
Emergence and enrichment of antibiotic resistance at sub-MIC have been studied quite extensively over the last few decades1,5,8,13,14,15,16,17,18,19,20. However, a majority of these studies have focused on single-species systems, overlooking the fact that bacteria often exist as part of a consortium comprised of many microbial species. The biotic and abiotic interactions a bacterial species is exposed to could influence resistance selection through a range of different mechanisms. For example, competitive interactions between different microbes will reduce the availability of shared resources, which should negatively affect population density, reducing the number of possible doublings of the resistant strain thereby influencing its enrichment in the presence of antibiotics. Alternatively, microbial species can also partake in positive interactions through cross-feeding interactions, where the two species are mutually dependent on each other for essential metabolites21, which will increase the population density and the number of generations that could affect the emergence and enrichment of antibiotic resistance. Also, the presence of another species in the ecosystem might result in sequestration of antibiotics present in the environment, which will reduce the level of available antibiotics affecting bacterial growth, thereby slowing the rate of selection. Finally, some bacterial species are also known to deactivate antibiotics (e.g., degradation of beta-lactams by beta-lactamases22,23,24), which is also expected to reduce the rate of enrichment of the resistant strain.
In the context of antibiotic resistance, the presence of a consortium of species from pig feces has been shown to offset the relative fitness advantage of kanamycin and rifampicin-resistant strains in the presence of a range of antibiotic concentrations20. Also, the presence of Acetobacter species was shown to improve the survival of susceptible Lactobacillus cells in the presence of rifampin25. Other examples have established the effect of interspecies interactions on the modulation of relative fitness of resistant strains in the presence of antibiotics (reviewed in ref. 26), but in a majority of cases, the mechanisms underlying these effects remain undefined, partly due to the difficulties in identifying and manipulating interactions in complex natural communities20,27,28.
Here, we designed a defined and genetically amenable synthetic community to study the impact of species interactions on antibiotic resistance dynamics. To this end, we used Escherichia coli MG1655 and Bacillus subtilis 168 to study the selection for nitrofurantoin (NIT) resistant E. coli at sub-MIC NIT levels. B. subtilis is a Gram-positive bacterium that has found a wide variety of uses as a cell factory29, animal feed additive30, probiotic31, as well as a probable antigen delivery vector to the mucosal layer in the gut32. The use of the bacterium in food as well as a vector to the gut might bring it into direct contact with the gut microflora, where E. coli is generally present. Furthermore, B. subtilis can in immunocompromised people cause several different types of infections33,34, implying that co-infections of E. coli and B. subtilis could occur. NIT is often used for the treatment of E. coli urinary tract infections35,36. NIT is a pro-drug that is activated inside the E. coli cells through nitroreductases, predominantly NfsA and NfsB, forming a free radical37 that is thought to damage DNA38,39 and ribosomes40. Loss of genes encoding these enzymes renders E. coli resistant to NIT41.
In this study, we followed the enrichment of NIT-resistant (NITR) E. coli in a mixed population of susceptible and resistant E. coli in the presence of another species, B. subtilis. We show that the rate of enrichment of the NITR E. coli cells at sub-MIC is impeded by the presence of B. subtilis. Further, our results show that the impediment is due to interference competition between the two species, largely mediated by extracellular molecules excreted by B. subtilis with the peptide, YydF, playing a key role.
Results
Species interactions slow nitrofurantoin resistance enrichment at sub-MIC
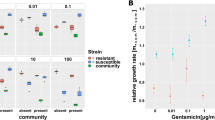
Using the broth microdilution method, we determined the MIC of NIT for both E. coli and B. subtilis to be 4 mg/L (Supplementary Fig. 1) in minimal media with glucose as the sole carbon source. Next, we elucidated the role of interspecies interactions on the selection of NIT resistance at sub-MIC (1/4, 1/6, and 1/8 of MIC of the susceptible strain) by competing susceptible and resistant (ΔnfsAB) fluorescent strains of E. coli. Starting from an initial frequency of ~0.01 (1%), the resistant cells enriched to 0.5–0.75 in frequency within three cycles of 24-h competitions in the absence of interactions with B. subtilis, compared to 0.05–0.15 in its presence (Fig. 1a), demonstrating a significant interaction induced dampening of NITR enrichment at sub-MIC (Fig. 1b). B. subtilis and E. coli both coexisted stably during the course of the experiment (Supplementary Fig. 2).
a Lines depict increases in frequencies of NIT-resistant (NITR) cells over 3 days in the presence of different sub-MIC levels of NIT. b Depiction of log-transformed relative fitness of NITR cells in the presence and absence of B. subtilis after 3 days of competition with NITS cells. Values on each plot represent those after Welch’s one-sided t test between log-transformed relative fitness estimates of resistant strain in the presence and absence of B. subtilis for each antibiotic concentration. The log-transformed relative fitness value of zero (gray horizontal line) represents no fitness advantage to either NITR or NITS strains, while positive values indicate an advantage to NITR strains. Lighter points depict individual replicates, darker points represent means and error bars represent 95% confidence intervals (t-distribution, n = 12 for both panels).
NITR enrichment is unaffected by resource competition
Competitive interaction between two species can take the form of scramble competition (where there is no direct interaction between the two species) or interference competition (where one species directly inhibits the other species)42. E. coli and B. subtilis are both capable of consuming glucose as a carbon source, while lactose and salicin can only be consumed by E. coli and B. subtilis, respectively. To examine the kind of interaction dominating the evolutionary dynamics of NIT resistance in our system, we performed the competitions in the presence of (i) glucose, (ii) lactose and salicin, or (iii) all three carbon sources. If the observed change in dynamics of resistance enrichment is largely due to scramble competition, we expect that to be mitigated when species are not involved in a competition for resources (when glucose is not provided as a carbon source) as compared to regimes when glucose was present during part or the entire growth cycle. In the presence of B. subtilis, the rate of resistance enrichment was similar among the three regimes (Fig. 2a) with no significant difference in mean log-transformed relative fitness after three rounds of competitions (Fig. 2b, ANOVA: F2,45 = 3.215, P = 0.0495; Tukey’s HSD test: LS vs G, P = 0.12; GLS vs G, P = 0.94; GLS vs LS, P = 0.06) suggesting a predominant role of interference competition in driving the observed slowdown of NIT resistance enrichment (initial and final population frequencies were used to determine relative fitness estimates over 3 days as per ref. 43).
a Lines depict increases in frequencies of NITR E. coli cells over 3 days at a quarter of MIC for both panels. Colors depict dynamics at the three available carbon sources in the media (G: 0.2% Glu; LS: 0.2% Lac + 0.3% Salicin; GLS: 0.1% Glu + 0.1% Lac + 0.15% Salicin). b Depiction of log-transformed relative fitness of NITR cells in the presence and absence of B. subtilis after 3 days of competition with NITS cells in the different carbon source regimes. The log-transformed relative fitness value of zero (gray horizontal line) represents no fitness advantage to either NITR or NITS strains, while positive values indicate an advantage to NITR strains. Lighter points depict individual replicates, darker points represent means, and error bars represent 95% confidence intervals (t-distribution, n = 16 for both panels).
Heat-sensitive compounds in the B. subtilis culture supernatant are largely responsible for dampening NITR enrichment
Among bacterial species, interference competition between species is frequently mediated by extracellular compounds44. We examined if that was the case here by elucidating the dynamics of NITR enrichment as well as its relative fitness in the presence of cell-free supernatant from a fully grown B. subtilis culture. We found that the relative fitness estimate of the resistant strain after three rounds of 24-h competitions to be significantly reduced in the presence of the B. subtilis supernatant (Fig. 3b, ANOVA: F2,33 = 128.8, P = 2.57 × 10−16; Tukey’s HSD test: PBS vs Filtered, P = 5.91 × 10−14), which reduces the rate of NITR enrichment (Fig. 3a). In addition, we found that the observed reduction is significantly offset with heat-treated supernatant (Fig. 3, Tukey’s HSD test: PBS vs Filtered+Heated, P = 2.09 × 10−12; Filtered+Heated vs Filtered, P = 0.0005). Thus, B. subtilis makes heat-sensitive compounds that affect the enrichment of NIT resistance in the presence of sub-MIC of NIT. Further, we also found that reducing the number of possible E. coli generations through dilution of growth media in half with phosphate-buffered saline did not affect the enrichment or relative fitness of NITR strain (Supplementary Fig. 3, Welch two-sample t test: t18.081 = 1.2833, P = 0.216) further pointing to the possibility of interference by B. subtilis population in dampening NITR enrichment.
a Lines depict increase in the frequency of NITR cells over 3 days at sub-MIC. Colors depict dynamics in the presence of either added PBS or cell-free supernatant (CFS) or heat-treated cell-free supernatant. b Depiction of log-transformed relative fitness of NITR cells after 3 days of competition with NITS cells under three supernatant treatment regimes. The log-transformed relative fitness value of zero represents no fitness advantage to either NITR or NITS strains, while positive values indicate an advantage to NITR strains. Lighter points depict individual replicates, darker points represent means and error bars represent 95% confidence intervals (t-distribution, n = 12 for both panels).
B. subtilis extracellular peptide YydF dampens E. coli NITR enrichment
Apart from significantly affecting NITR enrichment, B. subtilis culture supernatant also inhibited E. coli growth in minimal media (Fig. 4), so we set out to identify the component in the supernatant that might result in this inhibition with the assumption that the same component might be responsible for the change in E. coli NIT resistance enrichment. The approach we used to identify the component in the supernatant that results in this growth inhibition is summarized in Fig. 5. Briefly, we split the supernatant into two fractions—fraction 1 (with components of MWs >20 KDa) and fraction 2 (with components between MWs 3–20 KDa, see “Methods”). We found that fraction 2 results in significant inhibition of E. coli growth (Supplementary Fig. 4b). Further, we found the growth inhibition of E. coli to be highly variable between replicates (Supplementary Fig. 5a, b), which allowed us to identify three supernatants (among >20 tested) that inhibited E. coli growth to different extents (Supplementary Fig. 5c). Thus, we assumed that the degree of inhibition was related to the amount of the unknown protein components. Comparison of protein components in these supernatants by mass spectrometry revealed YydF and YorD to be proteins of interest (see “Methods” for details of the comparison).
Growth of NITS and NITR E. coli depicted as change in OD600 over 24 h in minimal media diluted with either sterile water or cell-free supernatant from B. subtilis. Each line is an average from three independent biological replicates. Error bars, shown as ribbons, represent 95% confidence intervals (t-distribution, n = 3).
To determine if the production of these proteins by B. subtilis did dampen NITR enrichment, we competed resistant and susceptible E. coli populations in the presence of B. subtilis strains knocked out for the genes encoding these proteins. The relative fitness of NITR E. coli was found to be significantly higher in the presence of B. subtilis ΔyydF compared to that in the presence of wild-type strain (Fig. 6b, ANOVA: F3,24 = 10.9, P = 0.0001; Tukey’s HSD test: ΔyydF vs WT, P = 0.017), exhibiting significant role of the protein in the dampening of NIT resistance enrichment. However, though the average relative fitness of the resistant E. coli population was found to be higher in the presence of B. subtilis ΔyorD strain (compared to that in the presence of wild-type strain), the difference was found to be statistically insignificant (Fig. 6b, Tukey’s HSD test: ΔyorD vs WT, P = 0.119), indicating a possible minor role of the protein in dampening NITR enrichment (Fig. 6).
a Lines depict increases in frequencies of NITR cells over 3 days at sub-MIC. Colors depict dynamics during interspecies interactions with different B. subtilis strains. The wild-type strain used here is the parent strain of ΔyydF and ΔyorD ordered from BGSC (see “Methods”). b Depiction of differences in log-transformed relative fitness of NITR cells after 3 days of competition with NITS cells in the presence of B. subtilis strains deleted for or carrying functional copies of either yydF or yorD genes. The log-transformed relative fitness value of zero (gray horizontal line) represents no fitness advantage to either NITR or NITS strains, while positive values indicate an advantage to NITR strains. All competitions were conducted in M9-Glucose with or without NIT, as mentioned in the legends (a) or on the x axis (b). Lighter points depict individual replicates, darker points represent means and error bars represent 95% confidence intervals (t-distribution, n = 8 for both panels).
Effects of YydF pre-pro-peptide and YydF* epipeptide on E. coli growth
Previous studies have shown the peptide YydF to be a 49 amino acid-long pre-pro-peptide that undergoes multiple modifications before being secreted by B. subtilis cells as a 17-mer epipeptide YydF*, which is capable of killing B. subtilis by dissipating its membrane potential45. Since the molecular weight of YydF*, at 2.1 KDa, is lower than the components in fraction 2, we first sought to determine if the 49-mer pre-pro-YydF was capable of affecting E. coli growth in minimal media thereby resulting in previously observed dampening of NITR enrichment. In the presence of NIT, log-transformed relative OD600 (ratio of E. coli OD600 in the presence over that in the absence of pre-pro-YydF after 24 h of growth) was significantly lower in the NITR strain as compared to that of NIT susceptible (NITS) strain at three of the four protein concentrations we tested (Supplementary Fig. 6, Welch’s one-sided t test: t9.87 = −2.199, P = 0.026 (8.75 nM); t13.158 = −4.451, P = 0.0003 (17.5 nM); t8.633 = −5.245, P = 0.0003 (35 nM) and t8.689 = −3.252, P = 0.005 (70 nM)). In the presence of NIT, the NITS strain showed 3.1–7.8% higher absolute OD600, while the NITR strain showed a 1.6–5.0% decline in the final OD600 values at different concentrations of the protein (Supplementary Fig. 6) when compared to control treatment without YydF. The protein did not exhibit any significant effect on NITR and NITS growth in the absence of the antibiotic. Overall, these findings are compatible with the notion that YydF dampens NITR enrichment.
A previous study has shown that the pre-pro-YydF has to be modified to the epipeptide YydF* by other proteins of the yyd operon before it starts affecting B. subtilis growth46. Accordingly, we sought to determine if the epipeptide YydF* could also differentially affect the growth of NITS and NITR E. coli. However, unlike that seen with pre-pro-YydF, we did not find any difference in log-transformed relative OD600 between NITS and NITR E. coli both in the presence and absence of NIT at all three YydF* concentrations (Supplementary Fig. 7, Welch’s one-sided t test: t2.134 = −2.337, P = 0.54 (5 μM); t2.592 = 0.105, P = 0.38 (2.5 μM) and t3.516 = −2.034, P = 0.8 (1.25 μM)]. No growth was observed in the presence of NIT in both control and test populations at the highest tested YydF* concentration of 10 μM (average OD600 after 24 h reached 0.04 and 0.12 in the presence and absence of YydF*, respectively).
Further, YydF* was shown to induce envelope stress in B. subtilis and since we found pre-pro-YydF to slightly affect NITR E. coli growth we looked for evidence of pre-pro-YydF induced differential transcription of envelope stress-related genes among NITR and NITS E. coli. However, a qPCR analysis did not reveal peptide-induced differential transcription of envelope stress-related genes cpxA and rpoE47 among NITR and NITS E. coli strains (Supplementary Fig. 8), suggesting that YydF* has different physiological effects in B. subtilis and E. coli.
Discussion
We show that interspecific interaction with B. subtilis can significantly dampen sub-MIC selection for NIT resistance in E. coli (Fig. 1). Moreover, the slowdown of resistance selection is not linked with resource competition between the two interacting species (Fig. 2). Instead, the significant reduction is probably mediated by extracellular compounds (Fig. 3), most notably the extracellular peptide YydF (Fig. 6).
The social nature of microbial existence is now well established, although most interspecies interactions in these microcosms are proposed to be antagonistic48. The extent to which these interactions can determine the outcomes of antibiotic treatment is still a black box awaiting deciphering. This study clearly shows that interspecies interactions can profoundly change the relative fitness of competing resistant and susceptible bacteria in a mixed population26. Extracellular molecules produced by bacterial species can modify its environment thus modulating the relative fitness of antibiotic-resistant strains. For example, acidification25, modification of antibiotics22,49 and bacteriocin production50 have been shown to modulate the relative fitness of antibiotic-resistant cells in a population. Previous studies have shown that the cell-free supernatant of a grown B. subtilis culture possesses a wide variety of growth inhibitory compounds (such as surfactins, iturin A, fengycin, etc.51) as well as a wide variety of proteins and enzymes52. Recently, cell-free supernatant of B. subtilis was shown to inhibit Staphylococcus aureus growth as well as make them susceptible to penicillin and gentamicin by compromising the integrity of its cell wall53. Here, we not only find that cell-free supernatant of B. subtilis to be inhibitory to E. coli growth (Fig. 4) but also show that an excreted extracellular peptide can be an important driver in reducing the relative fitness of NITR E. coli (Fig. 6).
The epipeptide YydF* (recently renamed to EpeX*54) has been shown to carry antibacterial activity towards B. subtilis itself46 by depolarizing the cell membrane through membrane permeabilization45. Moreover, the peptide is synthesized by the yyd operon (recently renamed to epe operon54), the expression of which is driven by a σA-dependent promoter55. The peptide is produced at the onset of stationary phase and elicits cell envelope stress in B. subtilis cells through the LiaRS two-component system45. Here, we show that pre-pro-YydF seems to have a small differential effect on the growth of NITR E. coli in the presence of NIT (Supplementary Fig. 6), which could partly explain its role in the dampening of NITR enrichment at sub-MIC (Fig. 6). However, the epipeptide YydF* does not seem to have any differential effect on NITR E. coli at the concentrations we tested (Supplementary Fig. 7). In addition, the lack of growth in control populations at the highest concentration of the peptide restricts our ability to conclude anything about the role of YydF* in dampening NITR enrichment, if any. It remains unclear how the peptide affects E. coli, it might be acting on all E. coli cells in the population to reduce growth, but it could also be specifically targeting cells that lack nitroreductases NfsA and NfsB, especially since the general reduction in the number of E. coli generations (through dilution of growth media, Supplementary Fig. 2) still lead to a dampening of NITR enrichment. If the specific activity of the peptide is directed at only NITR cells this could be a novel way of targeting NIT resistance. However, it should be noted that the activity of YydF does not fully explain the change in dynamics of NIT resistance and interactions with other molecular components (such as YorD) could also play a vital role in the overall dampening of NITR enrichment by B. subtilis (Fig. 6). Future work will determine the nature of E. coli interactions with pre-pro-YydF as well as YydF* epipeptide and whether these interactions can largely explain the dampening of NITR enrichment by B. subtilis.
Studies employing synthetic multispecies microbial systems are rare, especially those involving the study of evolutionary dynamics. For example, utilization of synthetic microbial biosystems has revealed modulation in secondary metabolite responses56, anti-predator strategies among bacterial prey57, traits which are important for the resilience of microbial communities58 and factors driving the diversification of CRISPR immunity59, emphasizing the importance of such systems in understanding the mechanisms of interspecies microbial interactions that play a role in modulation of bacterial traits, including antibiotic resistance. Our finding that the interaction between two bacterial species, powered by an extracellular peptide, could impact on the evolution of antibiotic resistance at sub-MIC should help in the endeavor of predicting the trajectory of a resistance trait once it has emerged in a population. Further, identifying the molecular drivers of interspecies interactions affecting the evolutionary dynamics of resistant traits should aid us in better controlling resistance emergence and evolution in bacterial populations.
Methods
Strains and media
Escherichia coli MG1655 strains (DA28100 and DA28102, both derived from parent strain DA4201) previously reported to carry chromosomal copies of fluorescent protein genes bfp and yfp were used for all experiments6. Fluorescent nitrofurantoin-resistant strains were constructed by moving the fluorescent protein genes from DA28100 and DA28102 into nitrofurantoin-resistant DA65117 (ΔnfsAB)60 using the P1 transduction protocol with chloramphenicol marker for selection61. The cultures were streaked onto LB agar (5 g yeast extract, 10 g Tryptone, 10 g NaCl, and 15 g agar per liter) plates and individual colonies were then inoculated into sterile liquid minimal media (1× M9 salts, 1 mM MgSO4, 0.1 mM CaCl2, 50 mg/L Tryptophan, 10 μM MnSO4, 1 μM FeSO4) with 0.2% glucose for overnight growth. B. subtilis subsp. subtilis 168 (also known as B. subtilis 168, a kind donation from Andreas Porse from Danmarks Tekniske Universitet) was used for most interspecies interaction experiments. B. subtilis Δyydf (trpC2 ΔyydF:kan; BGSCID: BKE40180) and ΔyorD (trpC2 ΔyorD:kan; BGSCID: BKE20420) strains were ordered from Bacillus Genetic Stock Center (BGSC), Columbus, Ohio, the US. For competitions involving knockout strains from BGSC, the BGSC parent strain (BGSCID: 1A1) was used as wild-type.
Competitions
Competition experiments to determine nitrofurantoin resistance enrichment as well as to estimate relative fitness were performed using the genetically tagged strains with chromosomal copies of bfp and yfp as described above. Six to eight colonies of each bacterial strain were grown in minimal media with 0.2% glucose. Then the susceptible strains tagged with one of the two fluorescent markers were mixed at 99:1 with the nitrofurantoin-resistant strain carrying the other marker in the same media with sub-MIC levels of nitrofurantoin (a quarter of MIC or unless otherwise mentioned). One microliter of the E. coli culture mix was then mixed with an equal volume of sterile water or fully grown B. subtilis culture (grown the same way as E. coli cultures) before being added to a well in a 96-well plate containing 198 μL of sterile minimal media to start the competitions. The competitions were performed under shaking at 32 °C. The following day, the competing strains were passaged by 100-fold dilution into fresh medium and were allowed to grow for another 24 h and passaged once more the next day. The ratio of resistance to susceptible cells was measured by counting at least 105 cells using a fluorescence-activated cell sorter (BD FACS Aria) before the passage and every 24-h thereafter for 4 days in total. The relative fitness of the resistant strain was estimated using the formula of Ross-Gillespie et al.43:
where \(x1\) and \(x2\) are the initial and final frequencies (after three rounds of competition) of the NITR cells, respectively.
Thus, the relative fitness estimate uses the initial and final frequencies of the NITR cells and does not generate per generation or per unit time estimate. The frequency of B. subtilis is estimated as the ratio of the total number of non-fluorescent cells over the total number of fluorescent cells in each sample where B. subtilis was added on day 0. The frequency of resistant cells (within E. coli populations), B. subtilis cells (in total population) as well as relative fitness was averaged across all 12–16 independent biological replicates across the two marker pairs.
Growth measurements
MIC using broth microdilution methods as well as growth of parental and fluorescent E. coli populations were determined in minimal medium using a Bioscreen C reader (Oy Growth Curves Ab Ltd). Three independent cultures for each strain were grown overnight and diluted 1:100 in fresh minimal medium. Three hundred microlitres of this suspension was added to the wells of the honeycomb plates. The plates were incubated in the Bioscreen C analyzer at 32 °C with shaking for 24–40 h as appropriate. The OD at 600 nm wavelength (OD600) value was measured every 4 min.
Experiments with supernatant
B. subtilis frozen stocks were streaked onto fresh LB agar plates and incubated overnight at 32 °C. Individual colonies were then inoculated into 25 mL minimal media with 0.2% glucose in 50 mL flasks and grown for 25 h under shaking at 32 °C. Fully grown cultures were then centrifuged at 4500 rpm for 15 min, and the supernatant was then filtered through a 0.2-micron filter to get rid of any cells. Sterile minimal media was then mixed with an equal volume of either the supernatant or with PBS (for the initial experiment) or water (for subsequent experiments) and used as growth media for competitions at sub-MIC of NIT. To test the effect of supernatants on E. coli growth, sterile minimal media with 0.2% glucose were mixed with an equal volume of supernatant or water and OD600 was measured every 4 min using Bioscreen under constant shaking at 32 °C. Non-fluorescent parental strains (DA4201 and DA65117) were used to test for the effect of the supernatant on growth. Heat-treated supernatant was made by heating the supernatant in an autoclave at 121 °C for 20 min.
Supernatant fractionation
Supernatant from B. subtilis were fractionated using PierceTM Protein Concentrators from ThermoFisher with filters at desired molecular weight cut-offs (MWCO). The supernatant was first passed using a centrifuge at 4000 × g through a column with MWCO of 10 KDa for 60 min, the retentate formed fraction 1 with components >10 KDa. The filtrate from the previous filtration was then filtered through a column with MWCO of 3 KDa at 4000 × g for 60 min, the retentate of which formed fraction 2 with components between 3 and 20 KDa. Fractions 1 and 2 were reconstituted to a final volume of 25 mL with fresh sterile minimal media and filtered through the same column again to rid of acids or other small molecular compounds. The resulting fractions were then reconstituted to a final volume of 25 mL with fresh sterile minimal media before being used to test their effect on E. coli growth the same way as with supernatant. The supernatant and the fractions were generated fresh for each experiment and were stored at 4 °C between use.
Proteomics
An aliquot of frozen B. subtilis 168 stock was thawed and spread onto multiple fresh LB agar plated with a sterile loop and incubated overnight at 32 °C. More than 20 equally sized colonies were then resuspended in 1 mL of minimal media and 900 μL of which was inoculated into 25 mL minimal media in 50-mL flasks and grown for exactly 25 h under shaking at 32 °C and used to harvest supernatant as before and tested for their effect on E. coli growth. Supernatant from replicate B. subtilis cultures had a varied non-repeatable effect on E. coli growth, so from the more than 20 cultures that were tested, we chose three cultures that showed high, intermediate and low effect on E. coli growth. Fresh supernatants were harvested from the three cultures (two replicates of each culture) as before, but the effect of the supernatant on growth was not repeatable, however, the three cultures still have varied effect on E. coli growth (Supplementary Fig. 3). Supernatants from both replicates of the three cultures were then concentrated using a PierceTM Protein Concentrators with MWCO of 3 KDa, protein bands in the supernatant were checked using SDS-PAGE (Supplementary Fig. 9) and were then stored at −80 °C until proteomic analysis. The concentrated supernatant samples were sent to Proteomics Core Facility at the University of Gothenburg for mass spectrometry-based analysis. Mass spectrometry-based relative quantification of protein across the samples were done using tandem mass tag (TMT) technology62. The analysis detected 346 unique protein or peptide sequences (Supplementary Data 1) across the six samples from which the peptides of interest were identified by first filtering for proteins or peptides that were relatively abundant in high-activity supernatant (1 S) as compared to the one with lowest activity (9 S) [(9 S)/(1 S) abundance ratio <0.1]. Next, we filtered the remaining proteins for intermediate presence in 1 S relative to the supernatant of intermediate activity (4 S) [(4 S)/(1 S) abundance ratio between 0.2 and 0.7]. Finally, we filtered out the proteins and peptides with MW higher than 20 KDa to arrive at five peptides (Supplementary Fig. 10) from which two (YydF and YorD) were characterized to have activities of interest as per the protein database Uniprot and were hence considered for further analysis.
Experiments with pre-pro-YydF and YydF*
YydF peptide or pre-pro-YydF was synthesized and purified by the Protein Science Facility of the Karolinska Institutet in Stockholm, Sweden. Briefly, based on the available genetic sequence of yydF, the peptide was transformed in to E. coli BL21 with a SUMO tag. The cells were cultivated in Terrific Broth (TB) medium. Protein expression was induced with isopropyl-β-D-1- thiogalactopyranoside (IPTG) and the protein was purified by immobilized metal-ion chromatography (IMAC), followed by size exclusion chromatography (SEC). The SUMO tag of the peptide was cleaved with Ubiquitin-like-specific protease 1 (ULP1). The cleaved SUMO tag was removed by reverse IMAC purification. The peptide was delivered in buffer at a concentration of 0.8 mg/mL (or 140 μM) in 20 mM HEPES, 300 mM NaCl, 10% glycerol, 2 mM TCEP, pH 7.5. This was diluted in sterile minimal media to appropriate concentrations before being checked for effect on E. coli growth.
The epipeptide YydF*: WYFVDKSKENRWIDLGSGH (where “D” denotes d-amino acid residues) was synthesized by Red Glead Discovery AB based at Lund, Sweden, as per the sequence reported in ref. 45. The lyophilized peptide was dissolved in 25% acetic acid and then added to minimal media to check for its effect on E. coli growth. Dissolved peptide was stored at −80 °C between uses, peptide was aliquoted into a number of tubes to avoid repeated thawing and freezing. Growth in the presence of peptide were compared with control treatments that contained equivalent amount of acetic acid in minimal media.
Measurement of changes in expression of cpxA and rpoE
Overnight cultures of NITS and NITR E. coli were diluted 1:100 in 20 mL minimal media and were grown at 32 °C to OD600 value of 0.5, upon which pre-pro-YydF were added to achieve a final concentration of 70 nM and grown for further 30 min. The entire culture was then spun down and resuspended in 1 mL of RNA protect reagent, vortexed, kept on ice for 5 min, spun down again and the pellets were stored at −80 °C until RNA extraction. RNA extraction and cDNA production was performed as per ref. 63 and described herein. RNA extraction was performed using the RNeasy Mini Kit (Qiagen) as per the manufacturer’s protocol. The extracted RNA was then treated with DNase using the Turbo DNA-free kit (Ambion) as per the manufacturer’s protocol. About 500 ng of RNA (quantified using the Qubit RNA BR assay kit) was used for cDNA preparation using the High Capacity Reverse Transcription Kit (Applied Biosystems). PerfecTa Sybr Green SuperMix (Quanta Biosciences) was used to perform the RT-qPCRs. The transcript abundance measured for the different genes was normalized to the geometrical mean of the levels obtained for housekeeping genes cysG, rssA, and hcaT. Ratio of expression in the presence of the peptide over that in the absence was used to generate plots. Two biological and three technical replicates were used in each case.
Statistics and reproducibility
Relative fitness values in all cases were log-transformed to achieve normal distribution. The difference in average relative fitness values post transformation in the presence and absence of B. subtilis was tested using one-sided t tests (Fig. 1b). The effect of available carbon source (Fig. 2b), supernatant treatment (Fig. 3b), and different B. subtilis strains (Fig. 6b) on relative fitness was tested using ANOVA and when found significant was further tested with Tukey’s HSD posthoc test to determine if there are significant differences in average log-transformed relative fitness between the treatments. Statistical significance of the effect of pre-pro-YydF and YydF* on growth of susceptible and resistant strains in the presence and absence of NIT was tested using one-sided t tests with alpha set at 0.0125 following Bonferroni’s multiple-significance-test correction (Supplementary Figs. 6 and 7). The number of biological replicates for each experiment is reported as n in each of the respective figure legends.
Statistical analyses were performed in R64 using RStudio 2022.02.3 + 492. All graphs were made using the R package ggplot265 and combined using the package patchwork66. The flow diagram (Fig. 5) was made using draw.io (www.diagrams.net).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data used for all the figures in the manuscript is available on Figshare (https://doi.org/10.6084/m9.figshare.21977045.v1). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE67 partner repository with the dataset identifier PXD040869.
References
Sandegren, L. Selection of antibiotic resistance at very low antibiotic concentrations. Ups. J. Med. Sci. 119, 103 (2014).
Drlica, K. The mutant selection window and antimicrobial resistance. J. Antimicrob. Chemother. 52, 11–17 (2003).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
Liu, J., Gefen, O., Ronin, I., Bar-Meir, M. & Balaban, N. Effect of tolerance on the evolution of antibiotic resistance under drug combinations. Science 367, 200–204 (2020).
Kraupner, N. et al. Selective concentration for ciprofloxacin resistance in Escherichia coli grown in complex aquatic bacterial biofilms. Environ. Int. 116, 255–268 (2018).
Gullberg, E., Albrecht, L. M., Karlsson, C., Sandegren, L. & Andersson, D. I. Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. mBio. 5; https://doi.org/10.1128/MBIO.01918-14 (2014).
Wistrand-Yuen, E. et al. Evolution of high-level resistance during low-level antibiotic exposure. Nat. Commun. 9, 1599 (2018).
Lundström, S. V. et al. Minimal selective concentrations of tetracycline in complex aquatic bacterial biofilms. Sci. Total. Environ. 553, 587–595 (2016).
Andersson, D. I. & Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 12, 465–478 (2014).
Baquero, F. & Negri, M. C. Challenges: selective compartments for resistant microorganisms in antibiotic gradients. BioEssays 19, 731–736 (1997).
Wallinga, D. & Burch, D. G. S. Does adding routine antibiotics to animal feed pose a serious risk to human health? BMJ 347, f4214–f4214 (2013).
Love, D. C., Davis, M. F., Bassett, A., Gunther, A. & Nachman, K. E. Dose imprecision and resistance: free-choice medicated feeds in industrial food animal production in the United States. Environ. Health Perspect. 119, 279–283 (2011).
Andersson, D. I. The biological cost of mutational antibiotic resistance: any practical conclusions? Curr. Opin. Microbiol. 9, 461–465 (2006).
Hughes, D. & Andersson, D. I. Evolutionary trajectories to antibiotic resistance. Annu. Rev. Microbiol. 71, 579–596 (2017).
Andersson, D. I. & Levin, B. R. The biological cost of antibiotic resistance. Curr. Opin. Microbiol. 2, 489–493 (1999).
Baquero, F., Alvarez-Ortega, C. & Martinez, J. L. Ecology and evolution of antibiotic resistance. Environ. Microbiol. Rep. 1, 469–476 (2009).
Khan, S., Beattie, T. K. & Knapp, C. W. The use of minimum selectable concentrations (MSCs) for determining the selection of antimicrobial resistant bacteria. Ecotoxicology 26, 283–292 (2017).
Stanton, I. C., Murray, A. K., Zhang, L., Snape, J. & Gaze, W. H. Evolution of antibiotic resistance at low antibiotic concentrations including selection below the minimal selective concentration. Comm. Biol. 3, 1–11 (2020).
Murray, A. K. et al. The ‘SELection End points in Communities of bacTeria’ (SELECT) Method: a novel experimental assay to facilitate risk assessment of selection for antimicrobial resistance in the environment. Environ. Health Perspect. 128, 107007-1–107007–10 (2020).
Klümper, U. et al. Selection for antimicrobial resistance is reduced when embedded in a natural microbial community. ISME J. 13, 2927–2937 (2019).
D’Souza, G. et al. Ecology and evolution of metabolic cross-feeding interactions in bacteria. Nat. Prod. Rep. 35, 455–488 (2018).
Perlin, M. H. et al. Protection of Salmonella by ampicillin-resistant Escherichia coli in the presence of otherwise lethal drug concentrations. Proc. R. Soc. B: Biol. 276, 3759–3768 (2009).
Bottery, M. J. et al. Inter-species interactions alter antibiotic efficacy in bacterial communities. ISME J. 16, 812–821 (2022).
Nicoloff, H. H. & Andersson, D. I. Indirect resistance to several classes of antibiotics in cocultures with resistant bacteria expressing antibiotic-modifying or -degrading enzymes. J. Antimicrob. Chemother. 71, 100–110 (2015).
Aranda-Díaz, A. et al. Bacterial interspecies interactions modulate pH-mediated antibiotic tolerance. eLife 9, e51493 (2020).
de Wit, G., Svet, L., Lories, B. & Steenackers, H. P. Microbial interspecies interactions and their impact on the emergence and spread of antimicrobial resistance. Annu. Rev. Microbiol. 76, 179–192 (2022).
Cairns, J. et al. Construction and characterization of synthetic bacterial community for experimental ecology and evolution. Front. Genet. 9, 312 (2018).
Cairns, J., Jokela, R., Becks, L., Mustonen, V. & Hiltunen, T. Repeatable ecological dynamics govern the response of experimental communities to antibiotic pulse perturbation. Nat. Ecol. Evol. 4, 1385–1394 (2020).
Westers, L., Westers, H. & Quax, W. J. Bacillus subtilis as cell factory for pharmaceutical proteins: a biotechnological approach to optimize the host organism. BBA-Mol. Cell. Res. 1694, 299–310 (2004).
Dänicke, S. & Döll, S. A probiotic feed additive containing spores of Bacillus subtilis and B. licheniformis does not prevent absorption and toxic effects of the Fusarium toxin deoxynivalenol in piglets. Food Chem. Toxicol. 48, 152–158 (2010).
Joerger, R. D. & Ganguly, A. Current status of the preharvest application of pro- and prebiotics to farm animals to enhance the microbial safety of animal products. Microbiol. Spectr. 5, 5.1.19 https://doi.org/10.1128/microbiolspec.PFS-0012-2016 (2017).
Batista, M. T. et al. Gut adhesive Bacillus subtilis spores as a platform for mucosal delivery of antigens. Infect. Immun. 82, 1414–1423 (2014).
Oggioni, M. R., Pozzi, G., Galieni, P., Valensin, P. E. & Bigazzi, C. Recurrent septicemia in an immunocompromised patient due to probiotic strains of Bacillus subtilis. J. Clin. Microbiol. 36, 325–326 (1998).
Richard, V., van der Auwera, P., Snoeck, R., Daneau, D. & Meunier, F. Nosocomial bacteremia caused by Bacillus species. Eur. J. Clin. Microbiol. Infect. Dis. 7, 783–785 (1988).
Huttner, A. et al. Nitrofurantoin revisited: a systematic review and meta-analysis of controlled trials. J. Antimicrob. Chemother. 70, 2456–2464 (2015).
Wijma, R. A., Huttner, A., Koch, B. C. P., Mouton, J. W. & Muller, A. E. Review of the pharmacokinetic properties of nitrofurantoin and nitroxoline. J. Antimicrob. Chemother. 73, 2916–2926 (2018).
Asnis, R. E. The reduction of Furacin by cell-free extracts of Furacin-resistant and parent-susceptible strains of Escherichia coli. Arch. Biochem. Biophys. 66, 208–216 (1957).
Jenkins, S. T. & Bennett, P. M. Effect of mutations in deoxyribonucleic acid repair pathways on the sensitivity of Escherichia coli K-12 strains to nitrofurantoin. J. Bacteriol. 125, 1214–1216 (1976).
Woody-Karrer, P. & Greenberg, J. Resistance and cross resistance of Escherichia coli S mutants to the radiomimetic agent nitrofurazone. J. Bacteriol. 85, 1208–1216 (1963).
McOsker, C. C. & Fitzpatrick, P. M. Nitrofurantoin: mechanism of action and implications for resistance development in common uropathogens. J. Antimicrob. Chemother. 33, 23–30 (1994).
McCalla, D. R., Kaiser, C. & Green, M. H. L. Genetics of nitrofurazone resistance in Escherichia coli. J. Bacteriol. 133, 10–16 (1978).
Łomnicki, A. Competition and behavior. in Encyclopedia of Ecology. Behavioral Ecology (eds Jorgensen, S. E. & Fath, B.) 695–700 (Elsevier, 2008).
Ross-Gillespie, A., Gardner, A., West, S. A. & Griffin, A. S. Frequency dependence and cooperation: theory and a test with bacteria. Am. Nat. 170, 331–342 (2007).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2009).
Popp, P. F., Benjdia, A., Strahl, H., Berteau, O. & Mascher, T. The epipeptide YydF intrinsically triggers the cell envelope stress response of Bacillus subtilis and causes severe membrane perturbations. Front. Microbiol. 11, 151 (2020).
Benjdia, A., Guillot, A., Ruffié, P., Leprince, J. & Berteau, O. Post-translational modification of ribosomally synthesized peptides by a radical SAM epimerase in Bacillus subtilis. Nat. Chem. 9, 698–707 (2017).
Raivio, T. L. MicroReview: envelope stress responses and Gram-negative bacterial pathogenesis. Mol. Microbiol. 56, 1119–1128 (2005).
Foster, K. R. & Bell, T. Competition, not cooperation, dominates interactions among culturable microbial species. Curr. Biol. 22, 1845–1850 (2012).
Egorov, A. M., Ulyashova, M. M. & Rubtsova, M. Y. Bacterial enzymes and antibiotic resistance. Acta Nat. 10, 33–48 (2018).
Koch, G. et al. Evolution of resistance to a last-resort antibiotic in Staphylococcus aureus via bacterial competition. Cell 158, 1060–1071 (2014).
Bonmatin, J.-M., Laprevote, O. & Peypoux, F. Diversity among microbial cyclic lipopeptides: iturins and surfactins. Activity-structure relationships to design new bioactive agents. Comb. Chem. High. Throughput Screen 6, 541–556 (2003).
Kaspar, F., Neubauer, P. & Gimpel, M. Bioactive secondary metabolites from Bacillus subtilis: a comprehensive review. J. Nat. Prod. 82, 2038–2053 (2019).
Zhang, F. et al. Bacillus subtilis revives conventional antibiotics against Staphylococcus aureus osteomyelitis. Microb. Cell. Fact. 20, 102 (2021).
Popp, P. F. et al. The epipeptide biosynthesis locus epeXEPAB is widely distributed in firmicutes and triggers intrinsic cell envelope stress. Microb. Physiol. 31, 306–318 (2021).
Butcher, B. G., Lin, Y. P. & Helmann, J. D. The yydFGHIJ operon of Bacillus subtilis encodes a peptide that induces the LiaRS two-component system. J. Bacteriol. 189, 8616–8625 (2007).
Traxler, M. F., Watrous, J. D., Alexandrov, T., Dorrestein, P. C. & Kolter, R. Interspecies interactions stimulate diversification of the Streptomyces coelicolor secreted metabolome. mBio 4, e00459–13 (2013).
Nair, R. R. et al. Bacterial predator-prey coevolution accelerates genome evolution and selects on virulence-associated prey defences. Nat. Commun. 10, 4301 (2019).
Amor, D. R. & Gore, J. Fast growth can counteract antibiotic susceptibility in shaping microbial community resilience to antibiotics. Proc. Natl Acad. Sci. USA 119, e2116954119 (2022).
Guillemet, M. et al. Competition and coevolution drive the evolution and the diversification of CRISPR immunity. Nat. Ecol. Evol. 6, 1480–1488 (2022).
Roemhild, R., Linkevicius, M. & Andersson, D. I. Molecular mechanisms of collateral sensitivity to the antibiotic nitrofurantoin. PLoS Biol. 18, e3000612 (2020).
Ikeda, H. & Tomizawa, J.-i Transducing fragments in generalized transduction by phage P1: I. molecular origin of the fragments. J. Mol. Biol. 14, 85–109 (1965).
Rauniyar, N. & Yates, J. R. Isobaric labeling-based relative quantification in shotgun proteomics. J. Proteome Res. 13, 5293–5309 (2014).
Warsi, O., Knopp, M., Surkov, S., Jerlström Hultqvist, J. & Andersson, D. I. Evolution of a new function by fusion between phage DNA and a bacterial gene. Mol. Biol. Evol. 37, 1329–1341 (2020).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria https://www.R-project.org/ (2022).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. Media (Springer-Verlag New York, 2016).
Thomas Lin Pedersen. patchwork: the composer of plots. https://CRAN.R-project.org/package=patchwork (2022).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Acknowledgements
This work was supported by grant UPD2020-0072 from the Wenner-Gren Foundation (RRN), grant KAW 2018.0168 from the Wallenberg Foundation (DIA) and grant 2021-02091 from the Swedish Research Council, Medicine and Health (DIA). Additionally, we thank all current and former members of the Andersson lab as well as other scientists in D7:3 corridor and Per Jemth for helpful discussions. We thank Omar Mahmud Warsi for the discussions as well as for his help in the lab. Marie Wrande and Po-Cheng Tang for help with flow cytometry. Ulrika Lustig for help in the lab. Members of Samay Pande lab at IISC Bangalore and Deepa Agashe lab at NCBS Bangalore for inputs. Emilia Strandback and Tomas Nyman at Protein Science Facility, KI, Stockholm, for help with YydF peptide, Egor Vorontsov and Carina Sihlbom at Proteomics Core Facility, University of Gothenburg, for help with supernatant analysis and Samara Mamidi at Red Glead Discovery for help with YydF* epipeptide.
Funding
Open access funding provided by Uppsala University.
Author information
Authors and Affiliations
Contributions
R.R.N. and D.I.A. planned the project and co-wrote the final manuscript. R.R.N. performed all the experiments, collected and analyzed the data, and wrote the initial draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Michael J Bottery and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Zhijuan Qiu. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nair, R.R., Andersson, D.I. Interspecies interaction reduces selection for antibiotic resistance in Escherichia coli. Commun Biol 6, 331 (2023). https://doi.org/10.1038/s42003-023-04716-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04716-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.